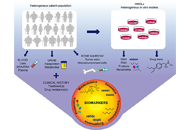
Off-Label Drugs and -Omics Data in Cancer Treatment
Guest Editors
Dr. Luca Agnelli E-Mail
Department of Oncology and Hemato-oncology, University of Milan, Milan, Italy
Research Keywords: lymphoid malignancies; bioinformatics and biostatistics in cancer; -omics analyses; NGS; non-coding RNA
Dr. Giancarlo Pruneri E-Mail
Fondazione IRCCS Istituto Nazionale dei Tumori, Milan, Italy
About the Special lssue
The knowledge about cancer has improved on the basis of -omics studies. The integration of genetic information about cancers with data on how the cancers respond to target based-therapy help to define optimum cancer treatment. As a consequence, in the last decades, targeted therapies led to unprecedented clinical benefit in first-line therapies, as well as in patients with aggressive or advanced neoplasms bearing specific genomic alterations (gene mutations, amplifications, translocations, microsatellite instability). Although it would be desirable to increase the number of clinical trials and standardized treatments, several factors influence so far the choice of therapies, including the non-trivial aspect of the cost-effective manageability by the Healthcare National Institutions. For this reason, several non-standard treatments are commonly considered, and off-label drug usage is a common practice in treating cancer, in most cases based on physician choice. This common clinical practice suffers from a lack of knowledge base for proper cancer drug selections. The present issue is aimed at focusing on all the aspects (clinical evidence, procedures, scientific hypothesis, and investigations) involving off-label drugs for cancer treatment in the context of -omics data.
Keywords: Off-label drugs; omics data; real-world cancer treatments; (targeted) NGS; biomarkers
Published Articles