Affiliation:
Ankara University Biotechnology Institute, Ankara 06135, TÜRKİYE
ORCID: https://orcid.org/0000-0001-7751-1405
Affiliation:
Ankara University Biotechnology Institute, Ankara 06135, TÜRKİYE
ORCID: https://orcid.org/0000-0003-2550-1605
Affiliation:
Ankara University Biotechnology Institute, Ankara 06135, TÜRKİYE
Email: evrimgunes@gmail.com; egunes@ankara.edu.tr
ORCID: https://orcid.org/0000-0003-4897-9388
Explor Foods Foodomics. 2026;4:1010137 DOI: https://doi.org/10.37349/eff.2026.1010137
Received: January 06, 2026 Accepted: April 02, 2026 Published: April 21, 2026
Academic Editor: Javad Sharifi Rad, Universidad Espíritu Santo, Ecuador
The article belongs to the special issue Natural Bioactive Compounds in Functional Foods: From Foodomics Insights to Clinical Relevance
Aim: This study aimed to investigate how the presence or absence of disulfide bonds affects the antimicrobial activity and thermal stability of pediocin PA-1.
Methods: To achieve this, the native pediocin peptide and a Cys → Ser mutant lacking the disulfide bridge were evaluated using both in vitro assays and molecular dynamics simulations. Antimicrobial activities of pediocin PA-1 and the mutant peptide were tested at varying temperatures (25–100°C) against selected indicator microorganisms. In parallel, molecular dynamics simulations were performed for both peptides, and RMSD, RMSF, and DSSP analyses were conducted to evaluate structural stability and secondary structure profiles.
Results: The Cys → Ser mutant peptide exhibited a substantial loss of antimicrobial activity, especially at elevated temperatures, demonstrating the necessity of the disulfide bridge for functional stability. In contrast, pediocin PA-1 retained approximately 96% of its activity even after exposure to 100°C. In silico analyses revealed that while the mutant partially preserved α-helix and β-sheet elements, it displayed pronounced disruption in its three-dimensional conformation.
Conclusions: The results highlight the critical structural role of Cys residues and disulfide bonds in ensuring both antimicrobial functionality and thermal resilience of pediocin PA-1. These findings provide valuable insights for the rational design of thermally stable antimicrobial peptides for food industry applications.
Bacteriocins are ribosomally synthesized antimicrobial peptides produced by a wide range of bacteria and are increasingly regarded as promising alternatives to conventional antimicrobials due to their ability to inhibit pathogenic and spoilage microorganisms [1–4]. Among them, pediocins and pediocin-like bacteriocins constitute a major subgroup of Class IIa bacteriocins and are well known for their potent anti-Listeria activity, structural stability, and robustness under food processing conditions [5, 6]. Pediocin is predominantly produced by Pediococcus species, members of the lactic acid bacteria (LAB), and has been widely applied as a natural preservative in dairy, meat, and minimally processed foods [7, 8]. Recent research further highlights the therapeutic potential of pediocin, including its cytotoxic effects against cancer cells and immunomodulatory functions [9–11].
Pediocin PA-1 (PA-1)/AcH, one of the best-characterized Class IIa bacteriocins, exhibits exceptional activity against Listeria monocytogenes and retains its antimicrobial performance even after exposure to high temperatures [12, 13]. Studies demonstrating the effect of thermal treatment with the pediocin peptide on bioactivity have shown that this bacteriocin remains stable for up to 30 min at temperatures between 60 and 100°C and even retains its antimicrobial peptide activity under autoclave conditions at 121°C for 15 min [14, 15]. This thermostability is largely attributed to the presence of two disulfide bonds, particularly the conserved C-terminal disulfide bond, which plays a critical role in maintaining structural integrity under thermal stress. Mutants lacking this C-terminal disulfide bond show dramatic reductions in antimicrobial activity, especially at elevated temperatures, whereas PA-1 typically maintains consistent activity across a broad temperature spectrum [12, 16].
Experimental characterization of bacteriocins, including thermostability and structural-functional relationships, often requires extensive laboratory work, advanced analytical instrumentation, and significant time and financial resources. Consequently, there is a growing shift toward computational approaches that enable the simulation of biological behavior, structure-activity relationships, and environmental stability of peptides [17–19]. In silico modeling plays an essential role in modern peptide engineering and drug discovery, allowing researchers to predict molecular interactions, explore sequence modifications, and optimize peptide performance prior to experimental validation [20, 21]. However, computational predictions must be supported by experimental data to ensure reliability, especially for parameters such as temperature stability, folding behavior, and activity against target pathogens.
Although several studies have examined pediocin and its derivatives using in vitro techniques, in silico investigations addressing the thermal behavior and structural dynamics of pediocin under high-temperature conditions remain limited [20, 22, 23]. Considering that bacteriocins intended for use in heat-treated foods must withstand processing conditions such as pasteurization or sterilization, understanding temperature-dependent structural stability is crucial. Therefore, further computational studies are needed to elucidate the thermal resilience of pediocins and to evaluate how sequence or structural modifications influence their antimicrobial function.
The proposed study aims to investigate the behavior and structure of both PA-1 and the mutant pediocin PA-1 (mPA-1; lacking disulfide bonds) at different temperatures using in vitro and in silico approaches.
The sequence of mature PA-1 is KYYGNGVTCGKHSCSVDWGKATTCIINNGAMAWATGGHQGNHKC according to the UniProt database (P29430.PPA1_PEDAC). In addition to the original PA-1 peptide, a mutant peptide lacking the cysteine (Cys) residues [replaced by serine (Ser) residues, sequenced as KYYGNGVTSGKHSSSVDWGKATTSIINNGAMAWATGGHQGNHKS] was synthesized by Elabscience Biotechnology Inc. (Houston, USA). The synthesized peptides, PA-1 and mPA-1, were obtained in lyophilized powder form. PA-1 has a purity of 94.1% and a molecular weight of 4,628.18 Da, while mPA-1 has a purity of 97.2% and a molecular weight of 4,563.91 Da (Mass Spectra and HPLC Chromatograms of the peptides are provided in Supplementary materials).
The antimicrobial activity of PA-1 and mPA-1 was evaluated using a modified well diffusion assay similar to that of Schillinger and Lücke [24]. Listeria monocytogenes ATCC 7644 served as the indicator strain. Tryptic Soy Agar (TSA) plates were prepared with 0.75% agar (soft TSA) and inoculated with 0.5% bacterial culture. Wells of 8 mm diameter were created, and 1 mg of each peptide was dissolved in 5 mL sterile distilled water and added to the wells (approximately 20 µg of peptide per well was used in the assay). Plates were incubated at 30°C for 24 hours, and inhibition zones were measured in millimeters. The inhibition zones were measured at their widest diameter using a digital/standard metric ruler. For each sample, measurements were performed in triplicate from different angles to ensure accuracy, and the mean values were recorded to the nearest millimeter. The diameter of the disc/well was included in the total zone measurement, consistent with standard CLSI (Clinical and Laboratory Standards Institute) guidelines.
Thermal stability was investigated by exposing 1 mg/mL peptide solutions to a temperature gradient (25–100°C) for 15 min in a thermal block device (Techne Dri-Blok DB-2D), consistent with previous studies [6, 25]. After heating, samples were rapidly cooled on ice, and their antimicrobial activity was reassessed using the well diffusion assay against Listeria monocytogenes.
MD simulations were performed using GROMACS 2022.3 [17, 26]. The OPLS-AA force field [27, 28] and TIP3P water model [29] were used for system preparation. MD simulations were conducted for 300 ns in explicit water. PA-1’s crystal structure (PDB ID: 5UKZ) served as the reference model. Simulations were carried out at 298, 310, 313, 323, 333, 343, 348, 353, 363, and 373 K (25–100°C) to evaluate thermal effects on peptide stability.
Analyses included root mean square deviation (RMSD), root mean square fluctuation (RMSF), secondary structure assessment via dictionary of secondary structure in proteins (DSSP), and solvent-accessible surface area (SASA) calculations. These metrics were interpreted to assess structural changes, flexibility, and thermal resilience of both PA-1 and mPA-1 peptides [20, 22, 23, 30].
In the current study, all in vitro experiments were performed in duplicate, and the results were expressed as mean ± standard deviation (SD). Statistical analyses were performed using one-way analysis of variance (ANOVA), and differences between groups were considered statistically significant at p < 0.05. Unlike in vitro analyses, considering the computational cost and the exploratory nature of the study, in silico analyses were performed as a single replicate. Long simulation times were used to reduce stochastic effects, and convergence was confirmed by monitoring the stability of RMSD and related parameters over time.
The antimicrobial activity of PA-1 against Listeria monocytogenes ATCC 7644 was evaluated across a range of temperatures (25–100°C). PA-1, synthesized with a purity of 94.1%, retained activity at all tested temperatures. Average inhibition zone diameters were 28.5, 28.0, 28.8, 29.0, 29.0, 28.0, 28.0, 28.0, 28.0, 27.8, and 27.8 mm from 25 to 100°C, respectively, compared to 29.0 mm for the unheated control. Statistical analysis (t-test) revealed a significant reduction in activity at 60–100°C (p < 0.05), yet even at 100°C, the activity decreased only by ~4% (Figure 1).
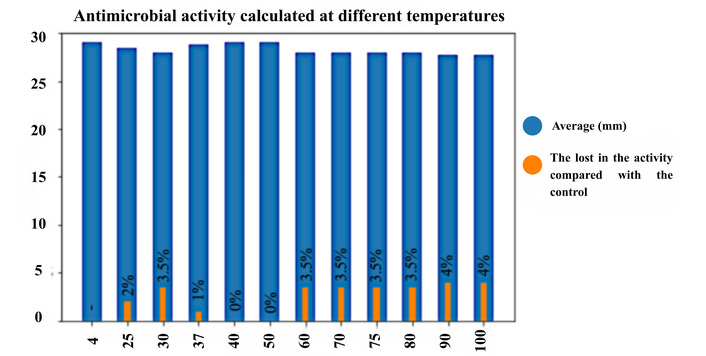
The antimicrobial activity of the pediocin PA-1 peptide on Listeria monocytogenes ATCC 7644 was examined at different temperatures. Blue bars show the antimicrobial activity zone diameters obtained at different temperatures. Orange bars show the loss of percent antimicrobial activity compared to the control sample for each temperature value.
The mPA-1 variant, in which all Cys residues were replaced by Ser, completely lost antimicrobial activity at all tested temperatures (25–100°C) (Figure 2). This observation underscores the critical role of disulfide bonds in maintaining the antimicrobial function of PA-1.
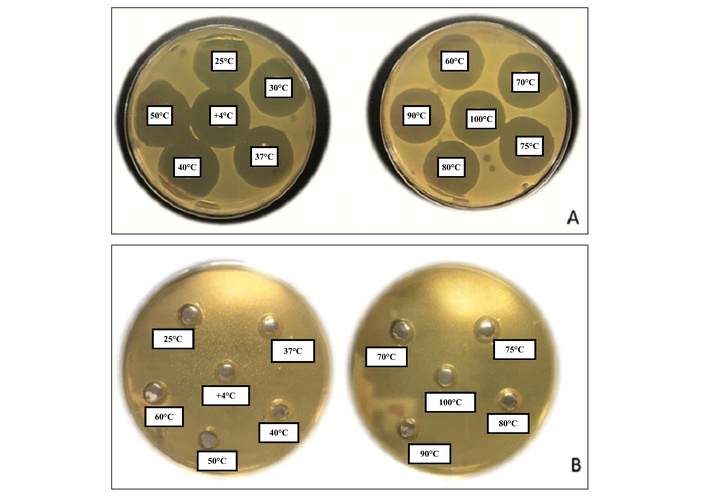
Petri images obtained in antimicrobial activity assays with the pediocin PA-1 peptide (A) and Cys-Ser mutant peptide (B).
MD simulations, including RMSD, RMSF, DSSP, and SASA analyses, were performed to investigate the structural stability of PA-1 (natural variant of pediocin) and mPA-1 (mutant variant of pediocin structure) at various temperatures. In a previous study conducted by our group [19], the structural changes of PA-1 were monitored at temperatures of 37, 40, 50, 60, 70, and 75°C using the GROMACS program (Version 5.1.4). The interactions between the bonds were analyzed, and it was observed that the disulfide bonds were closer together at 40°C. The structural deviations were found to be the least at 313 K (40°C), with a deviation of 0.6 nm, while the highest deviation of 1.2 nm was observed at 343 K (70°C). Notably, the structure exhibited significant conformational changes during the simulation performed at 348 K (75°C) compared to other temperatures. However, such drastic shifts were not observed at 313 K (40°C) and 343 K (60°C). DSSP analysis revealed a major structural change between 343 K (70°C) and 348 K (75°C). The peptide showed significant bending and rotational conformation, which was largely preserved during the simulation at 343 K (70°C). Additionally, during the initial 10 ns of the simulation, residues 24–34 maintained an α-helix structure at 343 K (70°C).
PA-1 stability: At moderate temperatures in this study (310–323 K; 37–50°C), PA-1 (the native variant of pediocin) generally retained its α-helical and β-sheet regions, despite temperature-dependent differences in backbone deviation (Figure 3A: RMSD plot of PA-1; Figure 3B: RMSD plot of mPA-1). RMSD reached approximately 0.7 nm at 310 K after 100 ns, whereas it remained around 0.3–0.4 nm at 313 and 323 K, suggesting that the peptide preserved its secondary structure throughout this range but exhibited comparatively greater conformational mobility at 310 K. At elevated temperatures (353–373 K; 80–100°C), RMSD and RMSF analyses revealed increasing fluctuations, partial unfolding, and α-helix transitions to 310-helices and random coils (Figures 4, 5, and 6). Despite these structural perturbations, the peptide retained some helical content, consistent with the retention of antimicrobial activity observed experimentally.
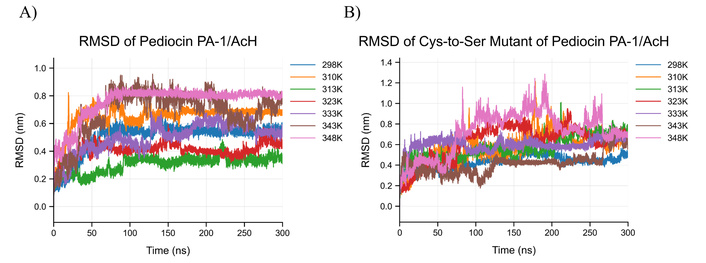
RMSD analysis of reference and mutant peptides at different temperatures based on MD simulations. A) RMSD plot calculated with Cα atoms as a result of MD simulations performed with reference peptide at different temperatures (298 K, 310, 313, 323, 333, 343, 348), B) performed with mutant peptide at different temperatures (298 K, 310, 313, 323, 333, 343, 348) RMSD graph calculated with Cα atoms as a result of MD simulations. MD: molecular dynamics; RMSD: root mean square deviation.
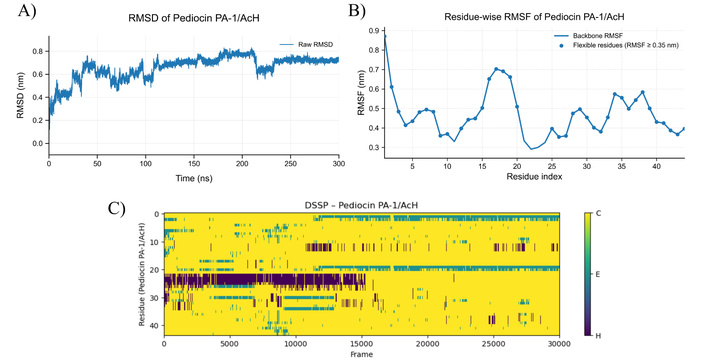
MD simulation results for pediocin PA-1/AcH at 353 K. A) RMSD plot of pediocin PA-1/AcH structure at 353 K, B) RMSF graph showing residual-based fluctuations of pediocin PA-1/AcH structure at 353 K, C) DSSP plot showing secondary structure changes of pediocin PA-1/AcH molecule at 353 K (C: Random coil, E: Beta sheet, H: Helices). MD: molecular dynamics; RMSD: root mean square deviation; RMSF: root mean square fluctuation; DSSP: dictionary of secondary structure in proteins.
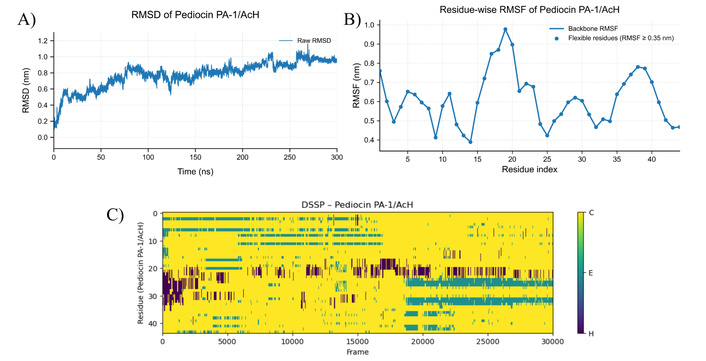
MD simulation results for pediocin PA-1/AcH at 363 K. A) RMSD plot of pediocin PA-1/AcH structure at 363 K, B) RMSF graph showing residual-based fluctuations of pediocin PA-1/AcH structure at 363 K, C) DSSP plot showing secondary structure changes of pediocin PA-1/AcH molecule at 363 K (C: Random coil, E: Beta sheet, H: Helices). MD: molecular dynamics; RMSD: root mean square deviation; RMSF: root mean square fluctuation; DSSP: dictionary of secondary structure in proteins.
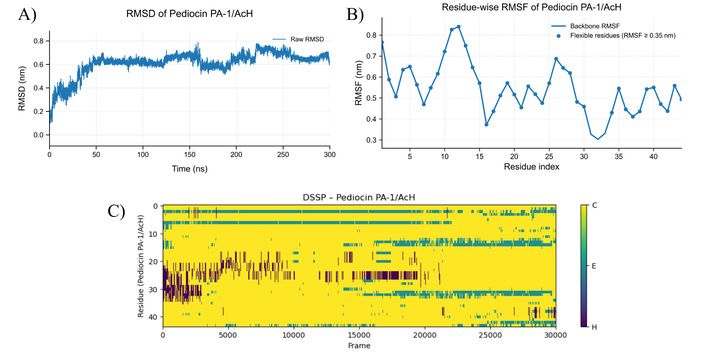
MD simulation results for pediocin PA-1/AcH at 373 K. A) RMSD plot of pediocin PA-1/AcH structure at 373 K, B) RMSF graph showing residual-based fluctuations of pediocin PA-1/AcH structure at 373 K, C) DSSP plot showing secondary structure changes of pediocin PA-1/AcH molecule at 373 K (C: Random coil, E: Beta sheet, H: Helices). MD: molecular dynamics; RMSD: root mean square deviation; RMSF: root mean square fluctuation; DSSP: dictionary of secondary structure in proteins.
mPA-1 instability: In contrast, the mPA-1 exhibited high RMSD fluctuations even at 298 K (25°C), and the α-helix and β-sheet regions were unstable across all temperatures. The PA-1/AcH peptide consists of an antiparallel β-sheet and an α-helix structure. Two disulfide bonds were present in close proximity to the N- and C-terminal regions of the peptide (Cys9-Cys14 and Cys24-Cys44) (Figure 6A). At the temperature of 310 K (37°C), the length of the disulfide bond between Cys9-Cys14 had an average value of 2.1 Å between the S-S atoms, while the average length of the disulfide bond between Cys24-Cys44 was 2.0 Å between the S-S atoms. Based on the DSSP results, it is evident that the secondary structure of the mPA-1 peptide exhibited high fluctuations at every temperature.
Figure 3A and Figure 3B present the RMSD changes with temperatures for the reference peptide and the mutant peptide, respectively. The RMSD fluctuations of the reference peptide indicated that it was highly stable at different temperatures (Figure 3A). At 298 K (25°C), the fluctuations ranged from 0.5 to 0.6 nm, and the structure stabilized after the initial 50 ns. The lowest fluctuations were observed at 313 K (40°C) and 323 K (50°C), indicating a higher degree of stability. In contrast, the RMSD plots of the mutant peptide without disulfide bonds (Figure 3B) showed significant fluctuations, particularly after 50 ns, as the structure lost its stability at higher temperatures.
The findings of the antimicrobial assay confirm the well-known heat resistance of pediocins. Similar observations have been reported in previous studies: Todorov et al. [31] demonstrated that pediocin-like peptides from Lactobacillus plantarum maintained activity after 120 min at 100°C, while Mathys et al. [32] reported that Pediococcus acidilactici UVA1 bacteriocin remained active up to 100°C. Recombinant forms of pediocin also preserve activity at high temperatures [20, 22, 23], supporting the relevance of these findings. According to Yu et al. [20], a bacteriocin produced by Pediococcus pentosaceus Z-1 demonstrated heat resistance between 50–100°C; in another study by Bungenstock et al. [22], the cell-free supernatant of Pediococcus spp. found active for up to 100°C for 30 min. Anh Thu et al. [23] conducted research on the recombinant form of PA-1 produced by the Escherichia coli BL21(DE3), fused with a NusA tag using a vector. The recombinant protein retained its activity even after heat treatment at 100°C for 15 min.
Although the amino acid sequence suggests the presence of a helical structure, it is not completely stable. The helical structure, which is known to exhibit antimicrobial activity by forming pores in the cell membrane and containing the second disulfide bond, frequently fluctuates between the α-helix and 310-helix structures. This observation indicates that the structure can not maintain stability without the presence of disulfide bonds. While the secondary structures without disulfide bonds individually have the potential to maintain their conformation (β-sheet region and α-helix region), the absence of disulfide bonds results in the loss of overall stability and complete disruption of the structure at all temperatures included in the MD simulations. The loss of helical stability of the C-terminal region may cause PA-1 to be unable to perform its function of opening pores that allow ions and small molecules to leak into the cytoplasm of target pathogenic organisms, which is fundamental to its mechanism of action. In addition, considering that previous studies have identified amino acids Cys14, Cys24, and Cys44 as essential for PA-1 activity, it can also be said that changes affecting the helical region at the C-terminus and the second disulfide bond, which is absent in other bacteriocins, would cause a decrease or loss of antimicrobial activity [33, 34].
The lower fluctuation range of the reference peptide compared to the mutant peptide suggested that the reference peptide could maintain stability, which is crucial for its antimicrobial activity. These findings are consistent with the experimental data. When comparing the structures of the two peptides, it is likely that the mutant peptide may lose antimicrobial activity compared to the reference peptide. This observation supports the importance of disulfide bonds in the activity and stabilization of PA-1/AcH, which is in agreement with the information available in the literature. DSSP analysis confirmed frequent transitions between α-helix and 310-helix, with complete disruption of the secondary structure at higher temperatures. These observations explain the complete loss of antimicrobial activity for mPA-1. The analyses demonstrate that disulfide bonds are essential for stabilizing the tertiary structure of PA-1 and preserving its antimicrobial function. The RMSF profiles further highlight that residues involved in disulfide bond formation are critical stabilization points, with high fluctuations in these regions correlating with loss of function. However, although these results were obtained by analyzing a single trajectory performed over a reliable and valid period of 300 ns, the results of the study are limited due to the nature of MD simulations, which may contain randomness.
These results align with previous mutagenesis studies, where replacement of Cys residues in the C-terminal region abolished peptide activity [12, 35]. Various aspects of the structure of pediocin have been investigated by numerous researchers. Kaur et al. [12] conducted a study on PA-1, where they replaced methionine with norleucine (Nle), resulting in a mutant peptide called ped[M31Nle]. This mutation was found to enhance the stability of the peptide against aerobic oxidation. In a study by Fimland et al. [35], similar to our study, a site-specific mutation was performed to replace the Cys residues with Ser residues to investigate the effect of the C-terminal disulfide bond on the structure of PA-1. While this study demonstrated the importance of the second disulfide bond in the antimicrobial activity of the peptide, the mutation was limited to the Cys residues involved in forming the second disulfide bond. These studies, along with our study, contribute to a better understanding of the structure-function relationship of pediocin, particularly the role of specific amino acid residues and disulfide bonds in its stability and antimicrobial activity.
Previous studies support the findings presented in the current study. Bédard et al. [36] conducted a study to understand the instability conditions of PA-1 and its analogs by utilizing a combination of solid- and solution-phase strategies. The results were evaluated using circular dichroism (CD) and NMR spectroscopy data. One of the analog peptides synthesized, labeled as 3c, shared a similar structure with PA-1 and exhibited high antimicrobial activity against Listeria, which was expected based on its structural similarity. On the other hand, the peptide labeled as 2c, which contained an oxidized form of Met31, did not display activity against the pathogens. The most surprising finding of the study was that the linear forms of the peptide (analogs 1 and 4) exhibited similar activity to that of PA-1 (analog 3c). This suggests that disulfide bonds can form in the medium even in the absence of chaperone-like proteins. These results shed light on the importance of disulfide bonds in the stability and antimicrobial activity of PA-1 and highlight the potential for alternative forms of the peptide to exhibit similar activity under certain conditions.
The activity of single-site mutants of the heterodimeric Class IIb bacteriocin, plantaricin EF [the bacteriocin consists of two peptides, plantaricin E (PlnE) and plantaricin F (PlnF)] was searched by Kyriakou et al. [37]. Using MD simulation with the NAMD 2.9 simulation engine, the researchers investigated the antimicrobial activity of the single-site mutants of PlnE and PlnF. They observed that the two-peptide system exhibited significantly stronger antimicrobial activity compared to the single-site amino acid substitutions. Furthermore, the researchers noted that the N-terminus of PlnF and the C-terminus of PlnE interacted with the membrane in most systems, emphasizing the importance of the interaction between the two peptides in creating the bactericidal activity of Class IIb bacteriocins. The study highlights the crucial role of the interaction between the two peptides in the antimicrobial activity of plantaricin EF, providing insights into the mechanism of action of Class IIb bacteriocins. In another study [38], a Class IIa bacteriocin called MMFII, which is a single-chain 37-residue polypeptide containing a single intramolecular disulfide bond, was investigated. The researchers synthesized MMFII using the solid-phase method and evaluated its antimicrobial activity against Listeria ivanovii. In their study, the researchers examined N-terminal analogs of MMFII, including Cys9-Trp37, Trp15-Trp37, and Val18-Trp37. They found that synthetic MMFII exhibited activity against Listeria ivanovii, whereas no activity was detected for the N-terminal analogs. This suggests that the presence of the YGNGV motif, which is characteristic of Class IIa bacteriocins, is crucial for the antimicrobial activity of MMFII. The researchers observed that deletion of the YGNGV motif could affect the β-turn structure of the peptide. They also suggested that the N-terminal β-sheet of the peptide might play a role in altering the bioactivity. The study emphasizes the importance of both N-termini and C-termini in the antimicrobial activity of MMFII and provides insights into the structure-activity relationship of Class IIa bacteriocins.
The study conducted by Wang et al. [39] investigated the thermal stability of pediocin, which is produced by Pediococcus acidilactici PA003. The results showed that pediocin remained active at high temperatures, such as 40, 60, 80, 100, and 121°C, for 1 hour. CD spectra analysis revealed that an increase in pediocin α-helix levels was accompanied by a decrease in β-strand and β-turn content, and this correlated with a decrease in pediocin activity. Interestingly, the study found that high temperatures, including a treatment at 100°C for 15 min, did not affect the disulfide bridges in pediocin. This indicated that the β-strand and turn structures were not destroyed, and the activity of pediocin was conserved. These findings suggest that β-strands and turns in the peptide structure are important for maintaining the high activity of pediocin.
The pediocin-like bacteriocins are peptides of the hydrophilic, cationic, and highly conserved N-terminal region that form a three-strand antiparallel β-layer supported by a shielded disulfide bridge. This N-terminal β-leaf region is followed by a central amphiphilic α-helix, followed by a rather long C-terminal tail that folds back into the central α-helix in most- if not all- of these peptides [40, 41]. In the current study, the RMSD, RMSF, and DSSP data emphasized the importance of disulfide bonds in maintaining the stability of pediocin. The mutant peptide without disulfide bonds attempted to maintain its secondary structure, including the β-sheet and α-helix regions, but exhibited an unstable structure with high fluctuations. These results suggest that the presence of disulfide bonds is crucial for the stability and antimicrobial activity of pediocin. The study highlights that the stability of the structure is directly related to the antimicrobial effect of pediocin, and a molecule lacking disulfide bonds is unlikely to play an active role in the antimicrobial action mechanism.
A study recently performed by our group [42] aimed to create mutant peptides capable of establishing a third disulfide bond through Cys substitutions. Five mutants (Mut 1–5) were systematically created with double Cys substitutions and assessed for thermal stability through MD simulations at 298–394 K (25–121°C) temperatures. In silico analysis indicated that Mut 4 (region of substitutions: Thr35 → Cys35, Gly37 → Cys37) retained its helical structure and exhibited thermal stability similar to PA-1. The study results imply that compared to PA-1, Mut 4 may have high stability and exceptional resistance to high temperatures, which also supports the results of the currently presented study.
In conclusion, this study highlights the critical role of disulfide bonds in maintaining both the antimicrobial efficacy and thermal stability of PA-1/AcH. While the wild-type peptide retained its structural integrity and bioactivity under elevated temperatures, the mutant variant lacking disulfide bridges exhibited significant conformational instability and complete loss of antimicrobial function. These findings underscore the structural advantages conferred by Cys-mediated disulfide linkages and provide valuable insights for the design of heat-stable bacteriocins. Ultimately, understanding such structure-function relationships may facilitate the development of next-generation peptide-based antimicrobials, particularly bacteriocins, which are a key functional compound of probiotics and postbiotic forms, for use in thermally demanding environments such as food safety at industrial applications.
CD: circular dichroism
Cys: cysteine
DSSP: dictionary of secondary structure in proteins
MD: molecular dynamics
mPA-1: mutant pediocin PA-1
PA-1: pediocin PA-1
PlnE: plantaricin E
PlnF: plantaricin F
RMSD: root mean square deviation
RMSF: root mean square fluctuation
SASA: solvent-accessible surface area
Ser: serine
TSA: Tryptic Soy Agar
The supplementary materials for this article are available at: https://www.explorationpub.com/uploads/Article/file/1010137_sup_1.pdf and https://www.explorationpub.com/uploads/Article/file/1010137_sup_2.pdf.
The MD simulations reported in this paper were partially performed at TUBITAK ULAKBIM, High Performance and Grid Computing Center (TRUBA resources).
BS: Conceptualization, Methodology, Investigation, Data curation, Writing—original draft. AAC: Investigation, Data curation. EGA: Conceptualization, Data curation, Formal analysis, Visualization, Writing—original draft, Resources, Writing—review & editing, Supervision, Funding acquisition, Project administration. All authors read and approved the submitted version.
The authors declare no competing interests.
Not applicable.
Not applicable.
Not applicable.
The datasets supporting the findings of this study are available from the corresponding author upon reasonable request.
This work was financially supported by the Scientific and Technological Research Council of Turkey (TUBITAK) [Grant No. TOVAG-119O343]. The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
View: 459
Download: 41
Times Cited: 0
María A. Cipollone ... Valeria A. Tironi
