Affiliation:
1Departamento de Química y Bioquímica, Bromatología, Facultad de Ciencias Exactas y Naturales, Universidad Nacional de Mar del Plata, Mar del Plata 7600, Argentina
2Instituto de Investigaciones Marinas y Costeras (IIMyC - CONICET), Facultad de Ciencias Exactas y Naturales, Universidad Nacional de Mar del Plata, Mar del Plata 7600, Argentina
Affiliation:
1Departamento de Química y Bioquímica, Bromatología, Facultad de Ciencias Exactas y Naturales, Universidad Nacional de Mar del Plata, Mar del Plata 7600, Argentina
Affiliation:
1Departamento de Química y Bioquímica, Bromatología, Facultad de Ciencias Exactas y Naturales, Universidad Nacional de Mar del Plata, Mar del Plata 7600, Argentina
Affiliation:
1Departamento de Química y Bioquímica, Bromatología, Facultad de Ciencias Exactas y Naturales, Universidad Nacional de Mar del Plata, Mar del Plata 7600, Argentina
Email: aliciadrobles@gmail.com; aliciadrobles@mdp.edu.ar
ORCID: https://orcid.org/0000-0002-9763-1380
Affiliation:
1Departamento de Química y Bioquímica, Bromatología, Facultad de Ciencias Exactas y Naturales, Universidad Nacional de Mar del Plata, Mar del Plata 7600, Argentina
Email: aisaiz@mdp.edu.ar
ORCID: https://orcid.org/0009-0000-1652-6523
Explor Foods Foodomics. 2026;4:1010127 DOI: https://doi.org/10.37349/eff.2026.1010127
Received: September 11, 2025 Accepted: January 15, 2026 Published: March 20, 2026
Academic Editor: Zhaowei Zhang, Oil Crops Research Institute of the Chinese Academy of Agricultural Sciences, China
The article belongs to the special issue Organic and Inorganic Compounds in Foods and Plants from Latin America
Aim: Organophosphorus pesticides (OPPs) have massively polluted ecosystems worldwide. Bioremediation by lactic acid bacteria (LAB) has been demonstrated to be an effective method to degrade them. This study aimed to evaluate the degradation capacity of four LAB strains on OPPs, using chlorpyrifos (CF) as the target pesticide. In addition, the interaction mechanism between CF and phosphatase enzyme was approached.
Methods: The degradation of CF by LAB strains was assessed over 24 h, and the remaining CF, along with its degradation products, were detected by gas chromatography-mass spectrometry (GC-MS). Molecular docking analysis was performed to determine the binding affinity between CF and phosphatase and to visualize the interaction within the binding pocket.
Results: The biodegradation of CF by L. mesenteroides, L. paramesenteroides, P. pentosaceus, and L. fermentum followed first-order kinetics, with degradation rate constants of 0.1318, 0.0279, 0.0241, and 0.0178 h–1, respectively. In accordance with the higher k value, L. mesenteroides isolated from vegetables exhibited the highest CF degradation rate (97%). Supporting this observation, CF showed significant binding affinity toward phosphatase from L. mesenteroides, with free energy values ranging from –5.79 to –5.77 kcal mol–1.
Conclusions: A positive correlation (P < 0.05) was observed between L. mesenteroides degradation behavior, phosphatase activity, and the degradation rate constant, indicating a metabolism better adapted to OPP stress conditions. The active site of the phosphatase, containing the Gly127-Glu128-Ser129-Ser130-Gly131 motif, was identified in pocket 1, suggesting that catalysis likely occurs at this site.
The excessive use of organophosphorus pesticides (OPPs) produces residues in agricultural products, the atmosphere, water bodies, and soil, leading to potential safety problems [1]. OPPs are a group of chemicals used in agricultural activities to control insect pests and thus protect plants in order to increase productivity [2], providing huge economic benefits to farmers. The application of OPPs in agriculture to control pests during pre-harvest and post-harvest practices results in the presence of their residues in food, which eventually reach consumers [3].
A joint report from the World Health Organization (WHO) and the United Nations Environment Programme (UNEP) estimates that pesticide exposure causes around 200,000 deaths and three million poisoning incidents annually worldwide, with nearly 95% of these cases occurring in developing countries [2]. OPPs are neurotoxic, and their lethality depends on their dose and level. They can cause seizures, strokes, respiratory failure, and even death [4]. Recent studies have reported that pesticide exposure exerts oxidative stress and produces damage to RNA, DNA, and proteins [5]. However, as the complete ban on pesticide use would put food production at risk, the elimination of pesticide residues from food has become a fundamental task [1, 6]. The compound diethoxy-sulfanylidene-(3,5,6-trichloropyridin-2-yl)oxy-lambda5-phosphane (IUPAC) [7], commonly referred to as chlorpyrifos (CF), was one of the OPPs most widely used in the United States (US), and it represented 11% of the total pesticides employed [8].
The Programme for Chemical Security from the WHO classified CF as Class II, which means moderate toxicity for humans [9]. The United States Environmental Protection Agency (USEPA) forbade the domestic use of CF and finally, in 2021, revoked any tolerance for the pesticide in foods [10]. In addition, from 2020, the Standing Committee on Plants, Animals, Food and Feed (PAFF Committee) of the European Union has established the removal of any product containing CF and CF-methyl. Thus, methods to eliminate pesticide residues from food are mandatory.
Different techniques have been employed for this purpose, such as oxidation technologies that use ozone, ultraviolet light, and ultrasound [11]. Another approach to OPP degradation is bioremediation, defined as the use of microorganisms, either naturally occurring or deliberately introduced, to reduce and decompose environmental toxicants. Fungal and bacterial strains isolated from soils [1, 12] and even cyanobacterial strains [13] have been evaluated. This type of non-toxic and harmless method has shown advantages due to its low cost, safety, and effectiveness [14]. In food matrices, OPP degradation by lactic acid bacteria (LAB) has been described mainly in fermentation processes [1], such as kimchi fermentation [15], wheat fermentation [16], both of which demand several days. In this sense, Lactobacillus plantarum [17], Lactobacillus brevis [18], Lactobacillus sakei and Leuconostoc mesenteroides [15] have been the main reported strains.
Considering the enzyme’s stereoselectivity and substrate preference is crucial in order to predict the affinity between them. Therefore, determining the structure of specific complexes formed between LAB phosphatase and CF is essential for a better understanding of the biodegradation function. AutoDock, a well-known docking protocol, has been widely used to dock small molecules with target enzymes [19, 20]. The docking procedure can predict the best binding pose of a specific ligand in terms of energy and conformation, thereby allowing the investigation of interactions between an enzyme and its ligands.
To the best of our knowledge, the isolation of LAB naturally found in food, such as raw fruits or vegetables, applied to OPP degradation over short incubation periods, has not been reported.
In this study, endogenous LAB strains isolated from vegetable samples (L. mesenteroides and Leuconostoc paramesenteroides) were evaluated for their ability to degrade OPPs within 24 h, using CF as the target. In addition, reference strains, such as Lactobacillus fermentum and Pediococcus pentosaceus, were used to compare degradation effects on OPPs. Finally, degradation rates and phosphatase activity allowed the determination of the efficiency of the bioremediation method. In addition, molecular docking was used to investigate the most probable binding sites, determine their positions, binding energies, and inhibition constants between CF and phosphatase from L. mesenteroides.
The CF was acquired from Gleba (La Plata, Buenos Aires, Argentina) as a commercial formulation of 48% w/v and stored at 16°C before use. DeMan-Rogosa-Sharpe (MRS) broth and agar were purchased from Britania (Buenos Aires, Argentina). The kit for the identification of API CH-50 strains (REF 50300, API systems, BioMerieux®, France) was stored at 4°C. The chemicals and organic solvents used in the different tests were of analytical grade and were acquired from Sigma-Aldrich (Merck - USA). The water used was deionized by membrane filtration (OSMOION 5, APEMA water purifier equipment). Fresh white cabbage (Brassica oleracea) was purchased in the local market in Mar del Plata, Argentina.
Two LAB strains, L. mesenteroides and L. paramesenteroides, were isolated from white cabbage (Brassica oleracea), and two reference strains of LAB, L. fermentum (CRL 220) and P. pentosaceus (CRL 922), provided by Reference Center for Lactobacilli (CERELA, Tucumán, Argentina), were used in this study.
Raw materials were washed with tap water and sliced. Ten grams of vegetable were suspended in 90 mL Butterfield’s phosphate sterile buffered dilution water (pH 7.2 ± 0.1). The standard pour plate technique, using MRS agar, was employed to isolate LAB strains, and plates were incubated for 48 h at 35°C under conditions of anaerobiosis [21]. The pure colonies were subjected to morphological and preliminary tests (catalase and oxidase test, nitrate reduction, indole production, gelatin and casein metabolization). Afterwards, LAB strains were re-isolated with MRS agar and broth, and finally identified as L. mesenteroides and L. paramesenteroides, using the kit API CH-50.
The reference strains and isolates from cabbage were grown overnight in 9 mL of MRS broth. Incubation was carried out at 35°C for 19 h. The cultures were centrifuged at 6,000 g for 10 min, and suspended in sterile saline solution (0.85% w/v). Then, the inoculum was adjusted to a McFarland standard of 2, which corresponds to a concentration of 6 × 108 CFU mL–1. One milliliter of this solution was re-suspended in 9.0 mL of 0.9% w/v sterilized saline solution, thus obtaining a 6 × 107 CFU mL–1 inoculum.
CF sterile solution was added to 20 mL MRS broth for a final concentration of 1.2 mg L–1, and then vortexed homogenized for 30 s. The bacterial suspension was inoculated into the flask containing MRS medium supplemented with CF to achieve a concentration of 105 CFU mL–1. The reaction mixture was vortexed for 2 min to ensure a homogeneous distribution of LAB strain cells and CF, and then incubated for 24 h at 35°C. Aliquots of 1 mL were taken at 0, 6, 12, and 24 h in order to quantify the residual CF. At the same time, LAB counts were performed in order to evaluate the population over time. The standard pour plate technique using MRS agar was carried out, and the plates were incubated at 35°C for 48 h. A control containing MRS and CF was carried out without bacterial suspension.
Extraction and purification were performed according to Pegoraro et al. [22] (2016) with some modifications. One milliliter from the OPP degradation assay was centrifuged at 10,000 g for 5 min, and the supernatant (300 μL) was cleaned using a glass mini-column filled with 1 g sodium sulfate (Na2SO4) and 0.6 g of activated silica. n-Hexane (270504-1L, Sigma-Aldrich, PRO - ACS analysis) was used as elution solvent (5 mL final volume), and a 1 mL prewash was performed. The clean extract was evaporated to dryness using N2 stream and finally, reconstituted with 1.0 mL of n-hexane for final analysis.
The CF was quantified by gas chromatography-mass spectrometry (GC-MS; Shimadzu CGMSQP2010 ULTRA-AOC20I) according to Zhang et al. [17] (2016) with light modifications. The column was an HP-5MS capillary column (15 m × 250 μm, 0.25 μm). The flow rate of the carrier gas (99.99% helium gas) was maintained at 35.2 cm s–1. The injection mode was split less. The injector temperature was set at 300°C. The interface and the ionization source were maintained at 280 and 230°C, respectively. Ionization was carried out by electron impact at 70 eV. Purified extracts (2.0 μL) were analyzed by a temperature ramp programmed as follows: held at 80°C for 0.5 min, ramped up to 230°C at 25°C/min, and then up to 300°C at a rate of 35°C/min and kept for 5 min, with a total run time of 13.5 min. Using a CF standard solution on the GC-MS system allowed the selection of three ion mass-to-charge ratio (m/z) in the selected ion monitoring (SIM) mode. The ion m/z = 314 was used for quantification and, m/z = 199 and m/z = 197 for identification. The instrument detection limit (IDL) was 5 μg/L, defined as a signal to noise ratio of 3:1 of the amount of pesticide, and was determined by calculating the signal to noise ratio for the CF in the lowest calibration standard. The method detection limit (MDL) was determined in accordance with the USEPA. To achieve this, we prepared seven solutions, which were spiked with CF at 15 μg/L. The resulting data were used to calculate the standard deviation of the measured concentrations. The MDL was then calculated by multiplying this standard deviation by the appropriate one-sided Student’s t-value for six degrees of freedom and a 99% confidence level (t = 3.14), resulting in 7.2 μg/L.
The products of CF degradation were also detected. GC-MS scan mode was employed to qualitatively distinguish these products, 3,5,6 trichloro-2-pyridinol (TCP) and diethyl thiophosphate (DETP), and they were identified by comparing the MS spectrum with data reported at the National Institute of Standards and Technology (NIST) Library. The GC-MS conditions used were according to the method of Ahir et al. [23] (2020) with slight modifications. The injection mode was split less. The oven temperature was held initially at 130°C for 1 min, programmed from 130 to 290°C at 15°C/min, and held at 290°C for 4 min. Run time was 15.67 min. Subsequently, the main m/z peaks were corroborated by SIM mode.
Cells were counted for growth evaluation at each time by the spread-plate method in MRS agar. Serial dilutions were spread on culture dishes and then incubated at 35°C for 48 h. Results were expressed as log10 CFU mL−1.
A solution of MRS broth, inoculated with 5% v/v of LAB strains overnight culture and 1.2 mg L–1 of CF, was incubated for 20 h at 35°C. Five milliliters of carbonate/bicarbonate buffer (10.0 mmol L–1, pH 10.5) was added and stirred for 1 min. The reaction mixture was treated by sonication 10 times (10 s with 15 s intervals) to break the bacterial cell wall and release the enzyme, then the suspension was centrifuged for 15 min at 10,000 g according to Sasikala et al. [24] (2012). The supernatant (crude enzyme solution) was used for the phosphatase activity assay.
Phosphatase activity was evaluated using the colorless reagent p-nitrophenyl phosphate (p-NPP) that produces p-nitrophenol (p-NP) as a yellow product [25]. The methodology was based on Zhang et al. [18] (2014) with minor modifications. The reaction mixture consisted of 1.0 mL of 5.0 mmol L–1 p-NPP [prepared in carbonate buffer with 1.0 mmol L–1 MgCl2·6H2O, Mg (II) acting as a cofactor], 1.0 mL of the carbonate buffer, 1.0 mL of the crude enzyme solution and distilled water, with a fixed final volume of 5.0 mL. After a reaction time of 60 min at 35°C, 3.0 mL of buffer termination (0.1 mol L–1 NaOH with 5.0 mmol L–1 EDTA) was added to stop the reaction. The generated p-NP was quantified by UV-VIS spectroscopy (Agilent 8435) at 400 nm. One unit (U) of phosphatase activity was defined as the amount of enzyme liberating 1 μmol of p-NP per minute at 35°C [18]. The reaction control was the assay without incubation.
The phosphatase activity calculation was made from Equation 1 [26]. The activity coefficient of p-NP at 400 nm was calculated from a calibration curve with a p-NP standard (Sigma-Aldrich Lot.: 13621 PD-166) in carbonate buffer 10.0 mmol L–1 pH 10.5.
where Ap-NP = absorbance of the incubation reaction mixture; Acontrol = absorbance of the reaction mixture without incubation; E = molar extinction coefficient of p-NP; L = optical path of the cell; Vtotal = total reaction volume; 106 = correction factor; and time = incubation time.
The interaction between the CF and a phosphatase from L. mesenteroides was analyzed through molecular docking investigation. The study aimed to identify the most probable binding sites, determining their position, binding energy, and inhibition constant. A structural prediction of L. mesenteroides hydrolase was used to search for potential CF binding sites.
A model of the target protein in .pdb format was obtained from the “AlphaFold” protein database, with the ID A0A896W8H3 in UniProt. A search for potential interaction sites was conducted on the LigQ website using a related protein, 4ZRS, a feruloyl esterase, as a model. This choice was based on its similarity as an esterase and its alignment with the study protein, as reported by Haque et al. [27] (2020). Alignment of both structures was performed using the Chimera program (UCSF Chimera 1.15/1.16) to visualize their similarity and locate potential interaction sites.
The structure of CF was obtained from PubChem, and optimization was conducted using Gaussian 09 with the semi-empirical PM6 method, followed by geometry convergence in the Molden program (Molden 7.0). A conformational space search of CF was carried out via molecular dynamics using the Gabedit program (Gabedit 2.5.0). The first four conformations with the lowest energy were selected for further optimization.
Molecular docking investigations were performed using the AutoDock4 program (AutoDock 4.2), with the aligned protein structure A0A896W8H3 from L. mesenteroides used as a rigid structure. Hydrogens were added to the starting .pdb file, and the atom model was configured as AD4 with Gasteiger charges to generate a “.pdbqt” file. Three different grids were generated for the structure, centered on the three potential binding sites obtained from LigQ, with a spacing of 0.303 Å.
The trials were performed in triplicate and subjected to statistical tests to evaluate variability. Kinetic parameters were studied by linear regression. Data analysis was performed using Statistica 8.0 software. Statistical significance between groups was assessed by comparing the difference in their mean values relative to the pooled standard error using independent samples t-test. The resulting test statistic was used to derive the corresponding P-value, which determined whether the observed differences were statistically significant (P < 0.05). All analyses were performed using standard parametric procedures unless otherwise indicated.
Several pesticide residue testing methods use dichloromethane in a liquid-liquid extraction [17]. However, chlorinated hydrocarbons should be replaced with less harmful alternatives for ecological and toxicological reasons [18, 28]. In this work, a solid-liquid partition method was used, employing a homemade mini-column assembled with a Pasteur pipette. Unlike liquid-liquid extraction, only a few milliliters of n-hexane were required as the eluent. The stationary phase consisted of activated silica, which acted as a purifying agent for organic and biological residues [28], and sodium sulfate acted as a desiccant. For quantification, 2 μL of the purified sample was injected into the GC-MS. The concentration of CF at each time point was calculated from a standard calibration curve.
Figure 1 shows the degradation rate of CF by four LAB strains. All strains exhibited the ability to degrade CF, achieving degradation rates ranging from 37 ± 2% to 97 ± 5% after 24 h of incubation. L. mesenteroides reached 80% degradation after just 12 h, while P. pentosaceus, L. paramesenteroides, and L. fermentum reached 34%, 27%, and 23%, respectively.
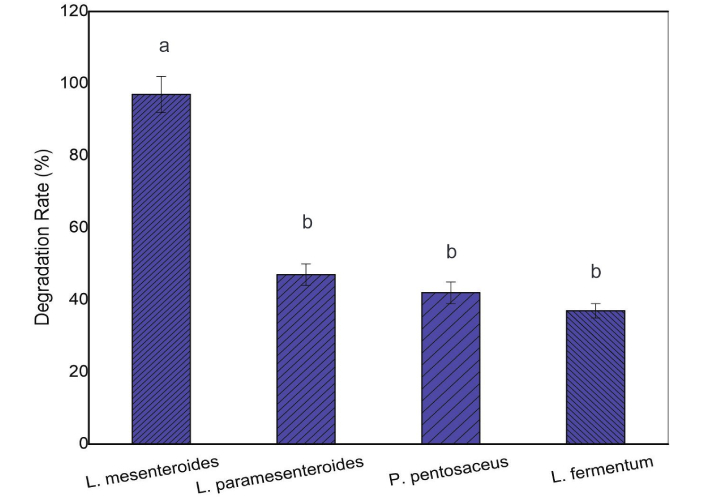
Degradation rate (%) of chlorpyrifos by LAB strains after 24 h of incubation at 35°C. Different letters above the bars indicate a significant difference (P < 0.05). LAB: lactic acid bacteria.
The CF degradation products were consistent with previous reports, in which TCP and DETP were described as the major metabolites formed during CF biodegradation [29–31]. Based on a comparison of GC-MS spectral data with literature reports on CF hydrolysis, the peaks at m/z 199, 197, 171, 169, 134, and 107 confirmed the formation of TCP, detected at 7.2 min (Figure 2). DETP, the second major metabolite, was detected at 9.2 min through peaks at m/z 95, 141, and 78. Regarding their toxicity, TCP has been found in samples of different origins (milk, urine, and blood) [32]. Unfortunately, TCP, like other products formed through the transformation of pesticides, has no established biological exposure limits, making it difficult to interpret the results in terms of human health risk [33, 34].
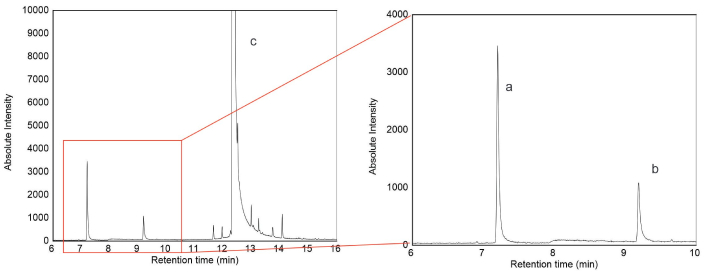
GC-MS chromatogram of chlorpyrifos degraded by LAB. Peak a (RT = 7.2 min) = 3,5,6 trichloro-2-pyridinol (TCP). Peak b (RT = 9.2 min) = diethyl thiophosphate (DETP). Peak c (RT = 12.35 min) = chlorpyrifos. GC-MS: gas chromatography-mass spectrometry; LAB: lactic acid bacteria; RT: retention time.
Degradation kinetics followed a first-order model in all strains (Figure 3, Table 1). The degradation rate constants for L. paramesenteroides, L. fermentum, and P. pentosaceus were comparable to those reported for other LAB strains [18]. Zhang et al. [18] (2014) found similar values for L. plantarum and S. thermophilus in milk, which showed degradation constants of 0.0186 and 0.0197 h–1 and degradation rates of 64 and 70%, respectively. In contrast, L. mesenteroides exhibited the highest rate constant and degradation efficiency, reaching 75% degradation after only 6 h of incubation.
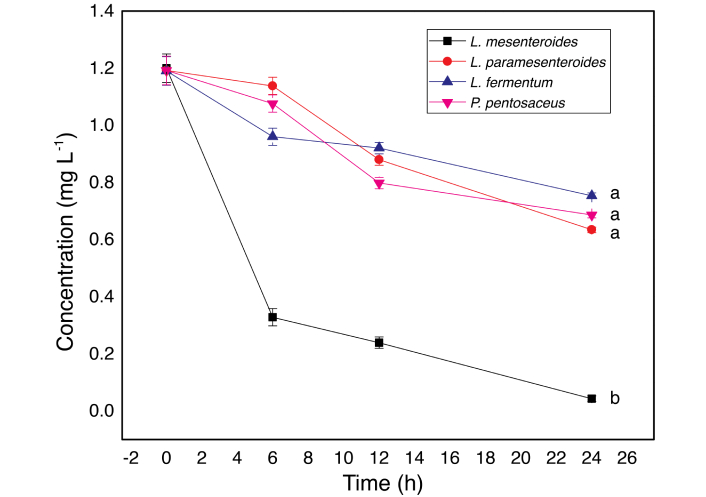
Degradation kinetics for CF by L. mesenteroides, L. paramesenteroides, P. pentosaceus, and L. fermentum. Data presented are means of three replicates ± standard deviation (n = 3). Different letters indicate a significant difference (P < 0.05). CF: chlorpyrifos.
Kinetic parameters of degradation and half-life times for CF by four LAB strains.
| LAB strains | Regression equation | R2 | k (h–1) | T½ (h) |
|---|---|---|---|---|
| L. mesenteroides | ln C = –0.1318 t + 0.0065 | 0.9571 | 0.1318 | 5.3 |
| L. paramesenteroides | ln C = –0.0279 t + 0.2229 | 0.9626 | 0.0279 | 24.9 |
| P. pentosaceus | ln C = –0.0241 t + 0.1652 | 0.8904 | 0.0241 | 28.7 |
| L. fermentum | ln C = –0.0178 t + 0.1282 | 0.9118 | 0.0178 | 39.9 |
CF: chlorpyrifos; LAB: lactic acid bacteria.
As shown in Figures 1 and 3, after 24 h of incubation, the strains had a positive impact on the degradation of CF, with percentages of 97 ± 5%, 47 ± 3%, 42 ± 3%, and 37 ± 2% for L. mesenteroides, L. paramesenteroides, P. pentosaceus, and L. fermentum, respectively. These results support previous findings on the bioremediation potential of LAB strains commonly used in food processing [15, 18, 35].
The bacterial population growth during CF degradation is shown in Figure 4. All four strains exhibited tolerance to CF. L. mesenteroides, L. paramesenteroides, and P. pentosaceus showed significant population increases during the first 6 h, with counts rising by 3 log cycles. After 12 h, increases of 4–6 log cycles were observed. However, a gradual decline was noted during the final 12 h, suggesting that CF imposes stress on bacterial cells [36].
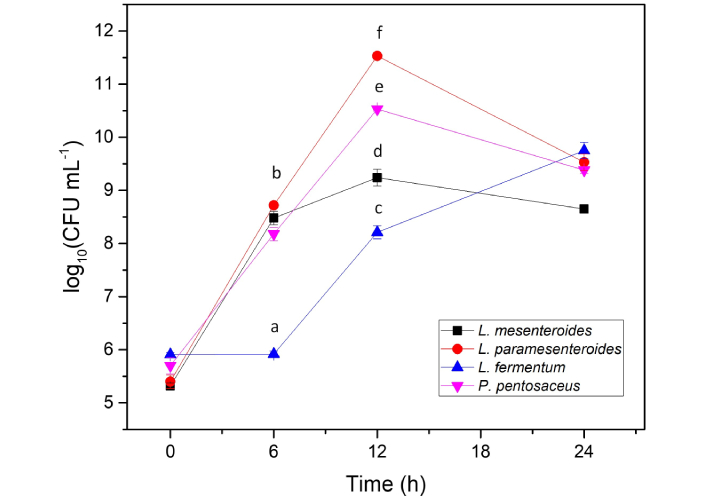
Growth of L. mesenteroides, L. paramesenteroides, L. fermentum, and P. pentosaceus in the presence of CF. Data presented are means of three replicates ± standard deviation (n = 3). Different letters indicate a significant difference (P < 0.05). CF: chlorpyrifos.
L. fermentum displayed a longer lag phase, with little change in population during the first 6 h (Figure 4). After that, its population increased continuously, consistent with delayed adaptation and enzyme synthesis. This behavior also matched its lower CF degradation rate. Notably, no significant correlation was found between bacterial growth and CF degradation in any strain.
Tolerance to CF may be linked to enzymatic activity. LAB strains are known to hydrolyze OPPs using enzymes such as phosphatases [1, 15, 18]. GC-MS analysis supported this, showing a tentative identification of TCP and DETP as hydrolysis products. Table 2 summarizes phosphatase activity and degradation constants for each strain. L. mesenteroides showed significantly higher degradation and phosphatase activity (P < 0.05) compared to the other strains.
Degradation rate, degradation constants (h–1), and phosphatase activity constants (U mL–1) of the strains L. mesenteroides, L. paramesenteroides, P. pentosaceus, and L. fermentum for CF.
| LAB strains | Degradation rate (%) | k (h–1) | Activity (U mL–1) |
|---|---|---|---|
| L. mesenteroides | 97 ± 5a | 0.1318a | 0.889 ± 0.018a |
| L. paramesenteroides | 47 ± 3b | 0.0279b | 0.2130 ± 0.0053b |
| P. pentosaceus | 42 ± 3b | 0.0241b | 0.2640 ± 0.0015b |
| L. fermentum | 37 ± 2b | 0.0178b | 0.2600 ± 0.0097b |
Different letters indicate a significant difference (P < 0.05). CF: chlorpyrifos; LAB: lactic acid bacteria.
Phosphatase activity values were 0.2130 ± 0.0053, 0.2600 ± 0.0097, and 0.2640 ± 0.0015 U mL–1 for L. paramesenteroides, L. fermentum, and P. pentosaceus, which showed CF degradation ranging between 37 and 47%. These results were similar to those of Zhang et al. [18] (2014), who reported phosphatase activities of 0.266–0.357 U mL–1 in LAB. Similarly, Zhou and Zhao [35] (2015) reported 0.081–0.175 U mL–1 in yogurt cultures. In contrast, L. mesenteroides reached 0.889 ± 0.018 U mL–1, suggesting greater metabolic versatility and stress resistance.
Docking analysis was performed using aligned protein structures and an optimized CF ligand. Although the 4ZRS protein had a greater number of residues, the structures overlapped satisfactorily. Binding interactions in pocket 1 involved van der Waals forces and hydrogen bonds with Arg59 and Lys66, while in pocket 3, they involved Lys41 and Ala42. These results differ from Haque et al. [27] (2020), who identified Ser128, Asp256, and His285 as key residues, likely due to methodological differences in pocket identification.
Table 3 summarizes the pocket parameters, binding energies, and inhibition constants. Pockets 1 and 3 showed similar free energies (ΔG ≈ –5.77 to –5.79 kcal mol–1), but pocket 1 contained a canonical hydrolase motif (Gly127-Glu128-Ser129-Ser130-Gly131), suggesting that catalysis likely occurs there. Hydrogen bonding also contributes to ligand stability [37, 38]. Figure 5 shows the loop (in orange) containing the Gly-Glu-Ser-Ser-Gly motif, where the dotted line represents the hydrogen-bond distance between the hydrogen of Ser129 and the oxygen of the thiophosphate group (2.99 Å). Pocket 3 may represent an allosteric site.
Interaction pockets of CF with protein model (4ZRS) with docking energies, inhibition constants, positions and amino acids involved in H-bonding, and H-bond lengths.
| Interaction modes | ΔGbinding (kcal mol–1) | Kinhibition (μM) | H-bond | Spacing (Å) |
|---|---|---|---|---|
| Pocket 1 | –5.79 | 57.14 | Arg59Lys66 | 0.303 |
| Pocket 3 | –5.77 | 58.8 | Lys41Ala42 | 0.303 |
∆Gbinding: Free binding energy between the phosphatase pocket and chlorpyrifos (CF); Kinhibition: inhibition constant reflecting the stability of the enzyme-ligand conformer; H-bond: amino acids that establish hydrogen bonds with CF; Spacing: H-bond length.
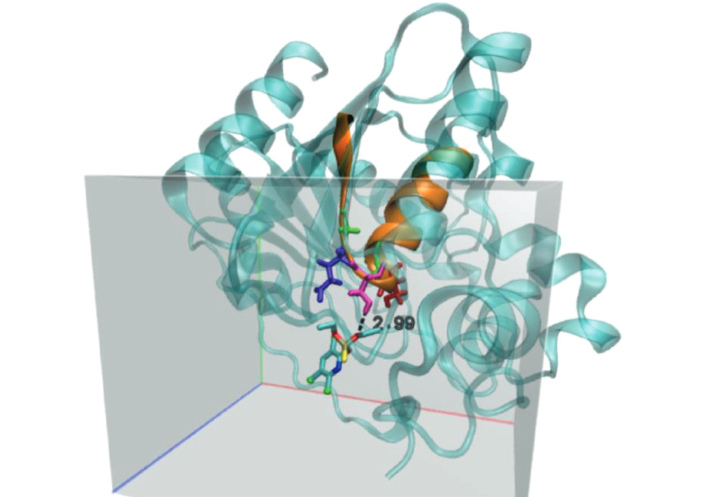
Active site of phosphatase containing the Gly-Glu-Ser-Ser-Gly motif. Amino acids were highlighted as green-blue-pink-red-green, respectively. The dotted line represents an H-bond. Chlorpyrifos molecule was obtained from https://pubchem.ncbi.nlm.nih.gov/compound/2730.
Although L. brevis showed stronger binding energy in previous reports (ΔG = –7.7 kcal mol–1) [37], the inhibition constant for L. mesenteroides was higher (> 50 μM), indicating greater resistance to CF and better degradation performance. This aligns with the 97% CF degradation observed in L. mesenteroides, compared to 76.5% for L. brevis [39].
The degradation of CF by LAB strains has been previously reported for L. plantarum, L. brevis, and S. thermophilus [15, 18, 39]. In this study, we demonstrated that L. mesenteroides, P. pentosaceus, L. paramesenteroides, and L. fermentum are also capable of degrading this pesticide, among which L. mesenteroides showed the highest efficiency. The results indicate that over a 24-h period, these strains positively impacted CF degradation, as evidenced by the degradation rate constants and the percentage of pesticide degradation, which ranged between 37 ± 2% and 97 ± 5%. Zhang et al. [18] (2014) tested CF degradation by L. plantarum and S. thermophilus on milk samples, and reported degradation rates of 64 and 70%, respectively. Cho et al. [15] (2009) studied the degradation of CF by LAB during kimchi fermentation and found that CF could be utilized by the strain as the sole source of carbon and phosphorus, showing a degradation percentage around 83.3%. In addition to the strains, environmental conditions can modify the adherence of OPP to cells. Bacterial cellular behavior may be altered by structural modifications resulting from pesticide adherence to the cell wall, which can cause damage to the cell surface, as reported by Wang et al. [40] (2016).
The biodegradation of CF by LAB used in the present study followed first-order kinetics, which is consistent with prior findings [18, 31]. However, rate constant values can vary depending on the OPP compound. Yuan et al. [1] (2021) reported differences in degradation rate constant values from L. plantarum on four OPPs (dimethoate, parathion methyl, trichlorfon, and CF), which have different molecular structures. Thus, the k value of dimethoate was 1.41 times higher than that of CF; therefore, bioremediation of OPPs is dependent not only on the bacterial species and culture conditions but also on the molecular structure of organophosphorus compounds [41].
The hydrolysis products, TCP and DETP, generated by the phosphatase activity on CF were tentatively identified by GC-MS. In agreement with previous reports, phosphatase activity appeared to be a key factor in the biodegradation process [1, 15, 18]. The strain L. mesenteroides exhibited both the highest phosphatase activity and the greatest CF degradation efficiency, indicating a strong correlation between enzymatic activity and pesticide breakdown.
Although all four strains showed CF tolerance, differences in population dynamics were observed. L. mesenteroides, P. pentosaceus, and L. paramesenteroides exhibited significant growth during the first 12 h, while L. fermentum displayed a longer lag phase. These differences may reflect the varying degrees of metabolic stress caused by CF, which can interfere with enzyme expression and bacterial growth [36].
The molecular docking results provided further insight into the possible mechanism of CF degradation. Docking simulations suggested that CF can bind to pocket 1 of the phosphatase enzyme, which includes a conserved hydrolase motif (Gly127-Glu128-Ser129-Ser130-Gly131). The free binding energy (ΔG ≈ –5.79 kcal mol–1) and the presence of hydrogen bonds with key residues such as Arg59 and Lys66 support this interaction as catalytically relevant. Although pocket 3 showed a similar binding energy, its lower functional significance and the absence of the catalytic motif suggest a possible allosteric or regulatory role.
These findings are consistent with those of Zhang et al. [18] (2014) and Zhou and Zhao [35] (2015), who also reported the involvement of phosphatases in OPP degradation by LAB. However, the binding sites and energy values differ slightly from those reported by Haque et al. [27] (2020), possibly due to variations in the docking protocols, pocket prediction algorithms, or protein-ligand optimization methods used.
From a practical standpoint, the ability of food-grade LAB to degrade CF has important implications for food safety and environmental bioremediation. LAB strains such as L. mesenteroides could be applied in the decontamination of pesticide residues in food matrices or in probiotic formulations designed to mitigate dietary pesticide exposure. Nevertheless, it is crucial to consider the potential toxicity of degradation products such as TCP, which has been associated with neurotoxicity and endocrine disruption [32–34]. Further studies are needed to evaluate the complete metabolic fate and safety profile of CF metabolites in biological systems.
This study confirmed that the ability of LAB strains to remove OPPs is positively correlated with phosphatase production. Indeed, TCP and DETP were tentatively identified by GC-MS as the main degradation metabolites. Significant removal of CF, measured in terms of degradation kinetic parameters, was observed in the four tested LAB strains. Among them, L. mesenteroides isolated from vegetables efficiently degraded CF and reached values above 90%, indicating a metabolism better suited to OPP stress conditions. Notably, the binding energy between CF and phosphatase was exergonic, indicating that the interaction occurs spontaneously. These findings imply that enzyme catalysis plays a crucial role in the degradation of CF by L. mesenteroides. This study is one of the key supporting studies needed to assess the remediation efficiency of bacteria commonly found in food. In this regard, L. mesenteroides could be applied post-harvest to remove OPP residues.
Finally, this study provides experimental and computational evidence supporting the role of LAB phosphatases in CF degradation. The combination of microbial screening, enzymatic assays, and in silico analyses offers a comprehensive approach to understanding pesticide biodegradation by food-grade bacteria.
CF: chlorpyrifos
DETP: diethyl thiophosphate
GC-MS: gas chromatography-mass spectrometry
L. brevis: Lactobacillus brevis
L. fermentum: Lactobacillus fermentum
L. mesenteroides: Leuconostoc mesenteroides
L. paramesenteroides: Leuconostoc paramesenteroides
L. plantarum: Lactobacillus plantarum
LAB: lactic acid bacteria
m/z: mass-to-charge ratio
MDL: method detection limit
MRS: DeMan-Rogosa-Sharpe
OPPs: organophosphorus pesticides
P. pentosaceus: Pediococcus pentosaceus
p-NP: p-nitrophenol
p-NPP: p-nitrophenyl phosphate
SIM: selected ion monitoring
TCP: 3,5,6 trichloro-2-pyridinol
U: unit
USEPA: United States Environmental Protection Agency
WHO: World Health Organization
AAP: Data curation, Formal analysis, Investigation, Methodology, Validation, Writing—original draft, Writing—review & editing. MOI: Conceptualization, Investigation, Methodology, Supervision. ARM: Formal analysis. ADR: Software, Visualization, Writing—original draft, Writing—review & editing. AIS: Conceptualization, Funding acquisition, Investigation, Project administration, Resources, Supervision, Visualization, Writing—original draft, Writing—review & editing. All authors read and approved the submitted version.
The authors declare that they have no conflicts of interest.
Not applicable.
Not applicable.
Not applicable.
The data that support the findings of this study are available from the corresponding authors, ADR and AIS, upon reasonable request.
This work was supported by the National University of Mar del Plata [Project EXA 1013/20]. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
View: 708
Download: 70
Times Cited: 0
Joseph A. Adeyemi ... Fernando Barbosa
Perla A. Magallanes-Cruz ... Nina R. Martínez-Ruiz
Antonio Berardi ... Antonia Tamborrino
