12 results in Exploration of Targeted Anti-tumor Therapy
Latest
Sort by :
- Latest
- Most Viewed
- Most Downloaded
- Most Cited
Open Access
Review
Risk factors for Gleason score upgrade from prostate biopsy to radical prostatectomy
Shayan Smani ... Michael S. Leapman
Published: July 30, 2024 Explor Target Antitumor Ther. 2024;5:981–996

Open Access
Commentary
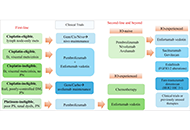
The rapidly changing treatment landscape of first-line advanced urothelial cancer (aUC) or metastatic urothelial cancer (mUC)
Minira Aslanova ... Jeanny B. Aragon-Ching
Published: July 29, 2024 Explor Target Antitumor Ther. 2024;5:971–980
This article belongs to the special issue Emerging Molecular Targets and Therapies of Genitourinary Tumors
Open Access
Review
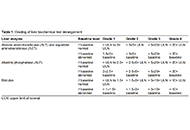
Hepatobiliary complications of immune checkpoint inhibitors in cancer
Donna Zhuang ... Stephen Riordan
Published: July 26, 2024 Explor Target Antitumor Ther. 2024;5:955–970
This article belongs to the special issue Immune Checkpoint Therapy and Biomarkers in Cancer
Open Access
Review
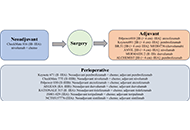
A narrative review on perioperative systemic therapy in non-small cell lung cancer
Robert Hsu ... David J. Benjamin
Published: July 26, 2024 Explor Target Antitumor Ther. 2024;5:931–954
This article belongs to the special issue Integrated Approaches for Non-Small-Cell Lung Cancer
Open Access
Case Report
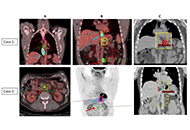
Oligometastatic esophageal cancer cured by systemic therapy combined with radiotherapy to primary tumor and metastasis (metastasis-directed therapy)—small case series
Mohan Hingorani, Hannah Stubley
Published: July 26, 2024 Explor Target Antitumor Ther. 2024;5:921–930
Open Access
Review
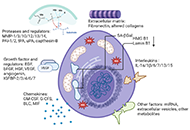
Therapy-induced senescence in breast cancer: an overview
Suraj Narayanan Chembukavu, Andrew J Lindsay
Published: July 25, 2024 Explor Target Antitumor Ther. 2024;5:902–920
This article belongs to the special issue Mechanisms of Targeted Therapy Resistance and Reversal Strategies
Open Access
Review
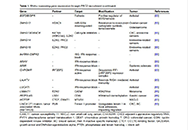
The genomic and molecular landscape of splenic marginal zone lymphoma, biological and clinical implications
Amatta Mirandari ... Jonathan C. Strefford
Published: July 23, 2024 Explor Target Antitumor Ther. 2024;5:877–901

Open Access
Review
Regulatory RNAs: role as scaffolds assembling protein complexes and their epigenetic deregulation
Palmiro Poltronieri
Published: July 22, 2024 Explor Target Antitumor Ther. 2024;5:841–876
This article belongs to the special issue Cancer Epigenetics: Implications for Novel Therapeutic Strategies
Open Access
Review
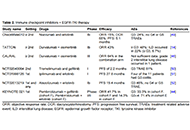
It might be a dead end: immune checkpoint inhibitor therapy in EGFR-mutated NSCLC
Ken Akao ... Kazuyoshi Imaizumi
Published: July 19, 2024 Explor Target Antitumor Ther. 2024;5:826–840
This article belongs to the special issue Novel Strategies and Targets for Immunotherapy of Cancer
Open Access
Case Report
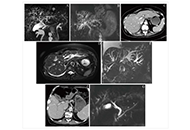
Immune checkpoint inhibitor-induced cholangitis—a three-case series
Simon Gray ... Anna Olsson-Brown
Published: July 19, 2024 Explor Target Antitumor Ther. 2024;5:818–825
Open Access
Review
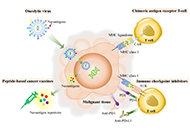
Immunopeptidomics in the cancer immunotherapy era
Sutatip Pongcharoen ... Supachai Topanurak
Published: July 17, 2024 Explor Target Antitumor Ther. 2024;5:801–817
This article belongs to the special issue Novel Strategies and Targets for Immunotherapy of Cancer
Open Access
Correction
Correction: Deep learning based automated epidermal growth factor receptor and anaplastic lymphoma kinase status prediction of brain metastasis in non-small cell lung cancer
Abhishek Mahajan ... Kumar Prabhash
Published: July 08, 2024 Explor Target Antitumor Ther. 2024;5:800