Affiliation:
1Departamento de Psicología Social y de las Organizaciones, Universidad Nacional de Educación a Distancia (UNED), 28040 Madrid, Spain
Email: a.borrego@psi.uned.es
ORCID: https://orcid.org/0000-0002-4699-3031
Affiliation:
2Departamento de Microbiología, Universidad de Málaga, 29071 Málaga, Spain
ORCID: https://orcid.org/0000-0002-2174-0652
Explor Neuroprot Ther. 2026;6:1004153 DOI: https://doi.org/10.37349/ent.2026.1004153
Received: December 31, 2025 Accepted: April 10, 2026 Published: May 13, 2026
Academic Editor: Antonio Ibarra, Anahuac University, Mexico
The article belongs to the special issue Role of Microbiota in Neurological Diseases
By individual examination, the present review provides an overview of the potential involvement of various human microbiomes, including the gut, oral, skin, and nasal, in the pathophysiology of neurodegenerative diseases. Research has demonstrated that gut microbiome dysbiosis is linked to the pathogenesis of neurodegenerative conditions, including Alzheimer’s, Parkinson’s, and Huntington’s diseases, through mechanisms involving microbial metabolites, neuroinflammation, amyloid aggregation, and altered neurotransmission. Emerging evidence suggests that the oral, skin, and nasal microbiomes may also influence neurodegenerative diseases through mechanisms such as microbial translocation, immune modulation, metabolite production, and interactions with the gut-brain axis. Although the potential of microbiome-based interventions for neurodegenerative diseases has been highlighted, several gaps remain, such as variability between human and animal models, a lack of standardized multi-omics approaches, and a limited understanding of individual microbial roles. Future studies should focus on clarifying the mechanisms by which dysbiosis in human host microbiomes impacts the pathophysiology of neurodegenerative diseases, identifying reliable biomarkers, and developing safe and effective microbiome-based therapies.
Neurodegenerative diseases (NDDs) encompass a diverse range of conditions that vary genetically, clinically, and pathologically. These diseases are marked by neurodegeneration, selective dysfunction, and progressive loss of both neurons and synaptic connections, often coupled with the accumulation of misfolded proteins with altered physicochemical characteristics in both the brain and peripheral organs [1, 2]. Major NDDs, including Alzheimer’s disease (AD), Parkinson’s disease (PD), and Huntington’s disease (HD), are primarily linked to cognitive deficits that represent a decline from a previous level of performance across one or more cognitive domains, including complex attention, language, executive function, learning and memory, social cognition, and perceptual-motor control, particularly as the disease progresses [3]. In addition, behavioral and psychiatric symptoms are frequently observed in NDDs, significantly impacting the patient’s quality of life, heightening caregiver stress, and being associated with higher healthcare utilization, greater rates of institutionalization, and increased mortality rates [4]. For instance, the prevalence of psychosis within NDDs varies according to the underlying condition, with rates around 30% in AD and 50% in PD [5]. In fact, both NDDs and neuropsychiatric disorders share underlying pathological mechanisms, including epigenetic factors, stress-related pathways, and microbiota–brain interactions, which complicate disease progression and therapeutic approaches [6].
A variety of genetic, epigenetic, medical, lifestyle, and environmental factors have been identified as contributing to the onset of NDDs [7–10]. Genetic risk variants across the genome, identified through genome-wide association studies, have been strongly linked with late-onset NDDs such as AD and PD, enabling the use of polygenic risk scores to estimate an individual’s genetic liability years before clinical symptoms arise [9]. For instance, the presence of the apolipoprotein E (APOE 4 allele) constitutes a significant genetic risk factor, which has been shown to elevate the risk of AD and PD [11]. In turn, environmental factors such as exposure to air pollution and pesticides have been shown to increase the risk of neurodegeneration, with pollutants contributing to neuroinflammation and the deposition of harmful proteins like amyloid beta (Aβ) plaques and Lewy bodies in AD and PD [8]. In addition, cardiovascular risk factors such as hypertension, inflammation, and dyslipidemia, which are closely related to cardiovascular disease (CVD), have also been linked to the onset of AD [10]. Both CVD and AD share long development periods, suggesting that CVD risk factors could predict the future onset of AD [10]. Furthermore, imbalances in the composition, diversity, and functionality of the gut microbiome (i.e., dysbiosis) have been associated with epigenetic modifications that affect gene expression and may contribute to the development of common NDDs such as AD and PD, opening new clinical perspectives for promoting neurological health [7]. In fact, the role of the gut microbiome in AD pathophysiology is pivotal, as gut microbiome dysbiosis can influence brain function through mechanisms such as inflammation, amyloidosis, neurodegeneration, and disruption of the blood-brain barrier (BBB), potentially exacerbating AD progression [12, 13]. Recently, a range of microbial metabolites, such as short-chain fatty acids (SCFAs), secondary bile acids (BAs), tryptophan derivatives (e.g., kynurenine, serotonin, tryptamine, and indoles), and trimethylamine/trimethylamine N-oxide (TMAO), have been linked to host pathways involved in the pathogenesis of NDDs [14]. Some of these metabolites are capable of crossing the BBB and can influence brain activity in multiple ways. These effects include modulation of neurotransmitter release, alterations in neuronal function, induction of oxidative stress and inflammation, and disruption of synaptic processes. Moreover, microbial metabolites may affect the central nervous system (CNS) through immune, enteroendocrine, and enteric nervous system pathways. Such disturbances can lead to changes in gut barrier integrity, immune responses in the periphery, and BBB functionality, as well as influencing neuronal homeostasis, neurogenesis, and the activation and maturation of glial cells [15, 16].
Research has indicated that the bidirectional communication between the brain and peripheral organs is essential for modulating and preserving overall homeostasis within the organism [17–19]. This interplay encompasses several components, such as the CNS, neurotransmitters, the mind-body connection, the peripheral nervous system, hormones, and chemical signals, all of which play a role in shaping emotional and behavioral responses [17, 20]. As a result, a comprehensive understanding of brain-organism communication is pivotal for elucidating the mechanisms that drive the development of NDDs.
The Human Microbiome Project was launched to create resources, methodologies, and insights aimed at exploring the connections between the human microbiome and health outcomes [21]. Although the terms “microbiota” and “microbiome” are often used interchangeably, they represent distinct concepts. The microbiome refers to the complete genetic material of all microorganisms present, including a diverse range of organisms such as archaea, bacteria, fungi, and viruses, as well as their structural components, metabolites, and the environmental factors influencing them. This comprehensive view underscores the intricate relationships and dependencies of microorganisms within their environment [22]. In contrast, “microbiota” is a narrower term, specifically referring to the community of microorganisms, whether commensal, symbiotic, or pathogenic, found in a particular environment [23, 24].
The human organism harbors a diverse array of microbiota, which can be classified into distinct microbiome types, such as those found in the bladder, eyes, gut, nasal cavity, lungs, oral cavity, skin, and vagina [25–28]. The gut microbiome, which is located in the gastrointestinal tract, has gained considerable attention due to the growing body of evidence highlighting its role in regulating brain and gut homeostasis through the gut-brain axis [17]. Research suggests that gut microbiome dysbiosis may play a significant role in the onset and progression of neuropsychiatric conditions, including NDDs [17, 29–31]. However, research investigating the role of other microbiomes in the context of NDDs remains scarce, likely due to the lower microbial abundance in these regions compared to the gut microbiome [25]. Exploring the microbiota in other organs is pivotal, as these regions may impact the pathophysiology of NDDs and interact with the gut microbiome through immune and inflammatory pathways [28, 32], potentially influencing disease development. This review provides an overview of the roles that various human microbiomes may play in the context of NDDs, including the gut microbiome, oral microbiome, skin microbiome, and nasal microbiome. Thus, by examining these microbiomes individually, this review aims to deepen the understanding of how dysbiosis in diverse microbial communities within the organism may contribute to the development and progression of NDDs.
It is an established fact that every bodily niche is inhabited by a microbiome, which refers to the collective presence of microorganisms within a specific environment. These microbiomes consist of components from all branches of the tree of life, including archaea, bacteria, eukarya, and viruses. Thus, animals should not be viewed merely as individual organisms based on physiological criteria. Considering the wide array of symbionts that interact with their hosts, animals possess additional and unique metabolic pathways that contribute to crucial physiological functions [27].
The relationship between the host and its microbiome has been conceptualized in various ways by different authors. Some have described it as a human organ [33], while others view it as a biological entity in its entirety [34], an ecosystem [35], or a selection unit within the hologenome concept of the holobiont [36]. Regardless of the framework, it is clear that due to their inherent complexity, eukaryotes cannot be considered independent organisms. Instead, they should be understood as natural units that include not only the host itself but also its associated symbionts and their collective metagenomes [34].
The role of human microbiomes in maintaining host health and contributing to various diseases has garnered significant interest within the scientific community [15]. The host-microbiota relationship involves a complex network of microbial interactions, including those within the eye, gut, nasal cavity, lungs, oral cavity, skin, and vagina microbiomes. Notably, the oral microbiome and nasal microbiome serve as critical entry points for potential pathogens, which can then spread to the CNS [37]. However, it is important to recognize that the influence of one microbiome on another is not always consistent across all body sites. For instance, the interconnection between the oral microbiome and the gut microbiome is particularly prominent within the host and plays a pivotal role in determining overall health, as well as influencing the development of various diseases throughout the organism [15, 32].
The human adult gut harbors a highly diverse microbial community, with microbial density (microbiota per gram) varying across different regions of the gastrointestinal tract: the stomach (101), duodenum (103), jejunum (104), ileum (107), and colon (1012) [38, 39]. However, it is crucial to note that approximately 70% of the microorganisms in the gut are not culturable, necessitating the use of molecular sequencing techniques to study this complex microbiome [40]. While the composition and diversity of the gut microbiome vary across these regions, research typically focuses on fecal samples, which are most closely associated with the colon and are easier to analyze. The gut microbiome consists of various microbial taxa, including archaea, bacteria, fungi, protozoa, and viruses, with bacteria being the most dominant group [29]. The human gut is predominantly colonized by several bacterial phyla, including Verrucomicrobiota, Bacillota, Campylobacterota, Actinomycetota, Fusobacteriota, Bacteroidota, Thermodesulfobacteriota, and Pseudomonadota [27, 29, 38, 41]. The distribution of microorganisms across different parts of the intestines has been the subject of considerable research. Studies on the upper gastrointestinal tract have identified Actinomyces, Fusobacterium, Gemella, Neisseria, Prevotella, Pseudomonas, Streptococcus, and Veillonella as the most prevalent bacterial genera [42]. In contrast, the lower gastrointestinal tract exhibits a distinct microbial profile, with genera such as Bacillus, Exiguobacterium, Lysinibacillus, Oceanobacillus, Paenibacillus, and Solibacillus being more abundant in the duodenal mucosa [43]. In the jejunum and ileum, common bacterial genera include Enterococcus, Escherichia, and Lactobacillus in the jejunum [44], and Clostridium, Escherichia, and Streptococcus in the ileum [45]. Finally, the colon is predominantly populated by genera such as Bacteroides, Coprobacillus, Enterococcus, and Escherichia/Shigella [46].
Increasing evidence indicates that the gut microbiome may play a central role in modulating behavior and brain function via multiple mechanisms, including the production of neurotransmitters such as γ-aminobutyric acid (GABA), dopamine, kynurenic acid, and serotonin, as well as a range of metabolites derived from microbial activity, such as BAs, SCFAs, D-amino acids, and tryptophan catabolites [14–16]. In this context, research has consistently shown a link between changes in the composition and diversity of the gut microbiome and the development of psychiatric and brain-related conditions, including various NDDs [47–52].
In the case of AD, Bostanciklioğlu [53] proposed three distinct mechanisms linking the gut microbiome to the pathogenesis of AD. These mechanisms include: (i) CNS inflammation and cerebrovascular degeneration induced by bacterial metabolites and amyloids; (ii) disruption of autophagy-mediated protein clearance due to an altered gut microbiome; and (iii) modulation of brain neurotransmitter levels via the vagal afferent pathway influenced by the gut microbiome. Research has shown that several bacteria, such as Bacillus subtilis, Escherichia coli, and Salmonella enterica serovar Typhimurium, are capable of producing amyloid fibers [54]. Interestingly, while microbial amyloids share tertiary structural similarities with human CNS amyloids, they act as prion-like agents through molecular mimicry, leading the amyloidogenic protein to adopt a pathogenic β-structure [55, 56]. In elderly individuals, these bacterial amyloid fibrils have been observed to cross both the intestinal barrier and the BBB, promoting an increase in Aβ accumulation in the brain and amplifying the inflammatory responses to cerebral amyloids, such as Aβ and α-synuclein [57]. In addition, the presence of bacterial lipopolysaccharides in the brain has been noted, suggesting that these bacterial components might play a role in AD development. It is hypothesized that lipopolysaccharides can cross both the intestinal tract and the BBB, where they interact with microglial TLR4-CD14/TLR2 receptors, leading to increased Aβ levels through elevated cytokine production [58]. Furthermore, Aβ1–42, which is a TLR4 agonist, has been shown to stimulate NFκB-mediated cytokine production, which in turn raises Aβ levels, damages oligodendrocytes, and contributes to the myelin degeneration observed in AD [59]. Elevated secondary BAs have also been linked to Aβ production and accumulation by disrupting the cholesterol metabolic pathway [60]. Moreover, the by-products of gut microbiome dysbiosis can influence the expression of genes and synaptic proteins involved in neuroinflammation, leading to the accumulation of inflammatory proteins in the brain, activation of astrocytes, neuronal apoptosis, and microglial inflammatory responses [61–63].
Despite the presence of inter- and intra-individual variability, as well as factors such as gender, diet, and geographic location, a substantial proportion of AD patients exhibit a distinct gut microbiome profile compared to age-matched healthy individuals [64]. A meta-analysis that included both discovery and replication cohorts confirmed a significant association between 20 specific bacterial genera and AD [65]. Notably, two bacterial species, Collinsella spp. and Veillonella spp., were strongly linked to the APOE rs429358 risk allele, whereas Eubacterium fissicatena was associated with a protective factor against AD. These results indicate that host genetic factors may play a substantial role in determining the abundance of specific bacterial genera, which could serve as potential biomarkers or therapeutic targets for AD [65]. Furthermore, meta-analytic data show that, regarding the phylum level, Bacillota are considerably reduced in AD patients in comparison with healthy controls, while Bacteroidota are significantly elevated in individuals with mild cognitive impairment compared to controls [66].
A substantial proportion of patients with PD exhibit gastrointestinal symptoms, including constipation, long before the appearance of motor deficits. This observation suggests a potential link between changes in gut motility and gut microbiome dysbiosis. The protein α-synuclein, which aggregates in the brains of individuals with PD to form Lewy bodies, has also been found to aggregate in the enteric nervous system [67]. Current hypotheses propose that the pathological α-synuclein could begin in the gut and later propagate to the brain through the vagus nerve, thereby contributing to PD pathogenesis [47, 49, 68, 69]. Tansey et al. [70] suggested that gene-environment interactions, combined with an aging immune system, may promote the development and progression of PD. Although several studies have linked gut microbiome dysbiosis to the pathogenesis of PD [71, 72], there is limited research on how the fecal microbiota is altered in PD patients with cognitive impairment. Wallen et al. [73] identified three distinct microbial clusters associated with PD: Cluster 1 consisted of opportunistic pathogens, all of which were more abundant in PD; Cluster 2 included SCFA-producing bacteria, all of which were less abundant in PD; and Cluster 3 contained carbohydrate-metabolizing probiotics, which were more abundant in PD. At the genus level, Bifidobacterium, Corynebacterium, Lactobacillus, Porphyromonas, and Prevotella were found to be more prevalent in PD patients compared to controls, while Agathobacter, Blautia, Butyricicoccus, Faecalibacterium, Fusicatenibacter, Lachnospira, Oscillospira, and Roseburia were less abundant. In a meta-analysis of PD patients from five countries, Hirayama and Ohno [74] reported that the Akkermansiaceae and Lactobacillaceae families were dominant in the gut microbiome of PD patients, with Akkermansia and Christensenella (formerly Catabacter) being the most prominent genera. In contrast, Faecalibacterium and Roseburia genera, together with the Lachnospiraceae family, were found to be reduced in PD patients. The reduction of SCFA-producing bacteria and the increase in mucin-degrading bacteria observed in PD may lead to increased intestinal permeability, potentially allowing toxins like lipopolysaccharides to reach the intestinal neural plexus. This could contribute to the abnormal accumulation of α-synuclein fibrils, further exacerbating PD pathology.
HD is an inherited neurodegenerative disorder characterized by progressive motor decline, cognitive dysfunction, and neuropsychiatric symptoms. This condition results from the age-dependent expansion of cytosine-adenine-guanine (CAG) repeats in the IT15 gene [75], leading to the production of a polyglutamine-expanded, nonfunctional huntingtin protein. The accumulation of this protein has been shown to form aggregates that ultimately cause neuronal damage [76]. The neuronal loss associated with these protein aggregates disrupts various physiological processes. Research indicates that HD is linked to impaired communication, organizational skills, and cognitive decline. These impairments can lead to a range of adverse outcomes, such as dementia, depression, and suicidal tendencies [77]. Beyond cognitive, motor, and neuropsychiatric symptoms, which are closely tied to brain changes, individuals with HD also experience a variety of gastrointestinal issues, including diarrhea, nutrient deficiencies, gastritis, and unintentional weight loss. These gastrointestinal disturbances are considered clinical features of the disease [51]. Although much of the research on HD pathogenesis has focused on brain atrophy and its associated cognitive, behavioral, and psychiatric symptoms, the role of peripheral pathology, particularly in the gut, in relation to the core symptoms of HD remains underexplored [78]. Mukherjea et al. [79] suggested that the gut microbiome in HD patients may present an increase in genera such as Akkermansia and Intestinimonas, whereas the Lachnospiraceae family and the Butyrivibrio genus are typically decreased. Figure 1 presents a hypothetical representation of microbial features associated with gut microbiome dysbiosis in the context of NDDs.
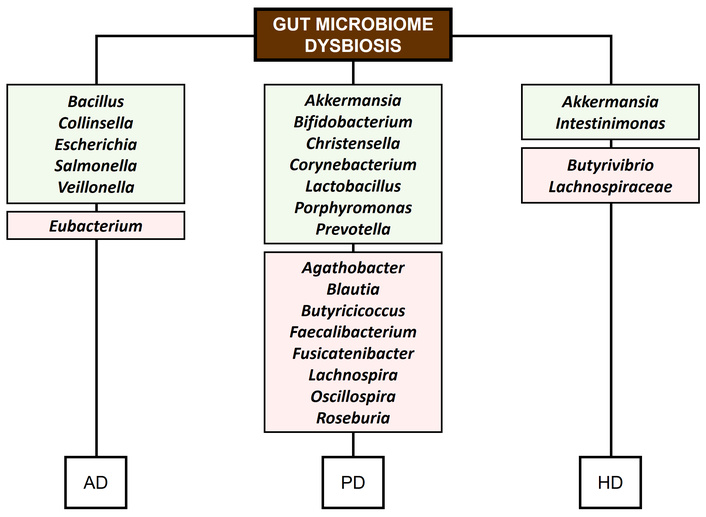
Microbial features associated with gut microbiome dysbiosis and NDDs. Green rectangles: increase. Red rectangles: decrease. AD: Alzheimer’s disease; PD: Parkinson’s disease; HD: Huntington’s disease.
Table 1 provides an overview of studies exploring the relationship between gut microbiome dysbiosis and the clinical pathophysiology of NDDs, thereby summarizing alterations in specific bacterial taxa and their reported effects on disease-related processes.
Gut microbiome dysbiosis and clinical features in NDDs.
| NDDs | Gut microbiome dysbiosis | Clinical features | References |
|---|---|---|---|
| AD | Bacillus subtilis, E. coli, Salmonella enterica serovar Typhimurium | Production of amyloid fibers | [54] |
| AD | Bacteroides fragilis, E. coli | Neuroinflammation | [58] |
| AD | Faecalibacterium, Pseudomonas | Systematic inflammatory reactions | [64] |
| AD | Collinsella | Interaction with the APOE rs429358 allele | [65] |
| PD | Akkermansia | Intestinal inflammationPermeability of the gut mucosal layer | [48, 74] |
| PD | Butyricicoccus, Clostridium XIVb | Non-motor symptomsCognitive impairment | [72] |
| PD | Lactobacillaceae, Prevotellaceae | Decrease ghrelin and α-synuclein accumulation | [80] |
| HD | Anaerobutyricum (formerly Eubacterium) hallii | Cognitive performanceMotor signs | [51] |
| HD | Bilophila | Immune response | [76] |
NDDs: neurodegenerative diseases; Aβ: amyloid beta; AD: Alzheimer’s disease; PD: Parkinson’s disease; HD: Huntington’s disease.
The oral microbiome refers to the diverse community of microorganisms residing in the human oral cavity, constituting the second-largest microbial ecosystem in the human body. This microbiome includes a wide range of microorganisms, such as archaea, bacteria, fungi, protozoa, and viruses [81, 82]. Bacteria dominate the oral microbiome in terms of both abundance and variety. The major bacterial phyla in the oral cavity include Actinomycetota, Bacillota, Bacteroidota, Fusobacteriota, Pseudomonadota, and Spirochaetota [83]. In addition, several minority and candidate phyla have been identified, including Gracilibacteria (GN02/BD1-5), SR1, Chloroflexota, Saccharibacteria (TM7), Chlorobiota, WPS-2, and Synergistota [81]. Cho et al. [84] suggested categorizing the oral cavity into three separate metaniches. In this respect, the P-GCF metaniche includes the bacterial genera that are present in both the gingival crevicular fluid and the supragingival dental plaque. This niche is primarily populated by Aggregatibacter (Pseudomonadota), Actinomyces and Corynebacterium (Actinomycetota), Fusobacterium and Leptotrichia (Fusobacteriota), and Capnocytophaga and Tannerella (Bacteroidota). The S-T-HP metaniche, consisting of the bacterial communities on the tongue, saliva, and hard palate, is mainly dominated by Alloprevotella, Porphyromonas, and Prevotella (Bacteroidota), Granulicatella and Veillonella (Bacillota), and Neisseria (Pseudomonadota). Lastly, the U-C metaniche, covering the cheek and sublingual area, is predominantly populated by Actinobacillus and Haemophilus (Pseudomonadota), and Gemella and Streptococcus (Bacillota). In contrast, Li et al. [85] proposed a different distribution of the oral microbiome into three principal niches: soft tissue surfaces, tooth surfaces, and saliva, which comprise the buccal mucosa, palate, tongue, and gingiva. In a study of the salivary microbiota within the context of a Qatari population, it was noted that the most prevalent genera were Prevotella, Haemophilus, Gemella, Neisseria, Porphyromonas, Veillonella, and Streptococcus [86]. However, research on the salivary microbiota of a Japanese cohort found a predominance of Rothia mucilaginosa, Streptococcus salivarius, Granulicatella adiacens, Streptococcus mitis, and Neisseria flavescens [87]. Such variations in microbial composition can be influenced by a range of factors, including environment, diet, and genetic predispositions [88]. In this context, the tooth surface offers an optimal niche for bacterial proliferation and the establishment of dental plaque. Research shows that dental plaque generally exhibits higher microbial richness, α-diversity, and greater uniformity when compared to microbial samples from the tongue or saliva [89]. Anaerobic bacteria such as Veillonella, Fusobacterium, and Actinomyces are particularly abundant in subgingival plaque [82]. The composition of the microbial community on tooth surfaces is also influenced by the anatomical and physiological characteristics of the tooth surfaces and the perigingival area. For instance, Streptococcus species are commonly found on the labial surfaces of canines and incisors, but are less frequently observed on the lingual surfaces [90]. The tongue, in contrast, supports a more diverse and denser microbial community compared to other mucosal areas [91]. Facultative and obligate anaerobes, including Veillonella, Streptococcus, Actinomyces, Neisseria, Haemophilus, Leptotrichia, Prevotella, and Porphyromonas, are the predominant microorganisms found on the tongue [89].
Given that the oral cavity is the primary gateway to the human body, alterations in the oral microbiome may play a role in the onset and progression of neuropsychiatric conditions, including NDDs [92, 93]. Research has linked specific oral microbiome profiles with individuals suffering from brain-related conditions, giving rise to the hypothesis that a relationship exists between these two elements. Specifically, this hypothesis proposes that the CNS may influence the oral microbiome, promoting the growth of certain microbial populations. Similar to the effects observed in the gastrointestinal tract, this phenomenon has been observed in patients with mental health disorders, who often demonstrate poor oral health [94]. Consistent with this, further evidence supports a link between the oral microbiome, oral diseases, and various NDDs, including AD and PD [95].
It has been suggested that the oral microbiome influences brain function through various direct and indirect mechanisms. One direct route involves the transmission of microbial material to the brain through the olfactory and trigeminal nerve pathways, which connect the oral cavity to the olfactory bulb [96]. Chronic oral conditions can compromise the oral mucosal barrier, permitting bacterial lipopolysaccharides and microorganisms to enter the circulation, frequently via mechanical actions like chewing or brushing. This process elicits a strong inflammatory response by stimulating endothelial cells expressing TNF-α and IL-1β receptors, which subsequently signal perivascular macrophages. In turn, these signals facilitate neuroinflammation and activate the hypothalamic-pituitary-adrenal (HPA) axis [81, 97]. Moreover, these inflammatory mediators can disrupt the BBB, a condition linked to inflammation and cognitive decline in NDDs [98]. Porphyromonas gingivalis, a key bacterium associated with periodontitis, has been shown to breach the BBB and colonize the brain, where it produces cysteine proteases (i.e., gingipains). These gingipains interfere with the metabolism of the transmembrane protein β-amyloid precursor protein (APP), which is crucial for neuronal development and synaptic stability [99, 100]. In turn, the oral microbiome could also influence brain activity in an indirect way via the oral-gut-brain axis [101]. Research has demonstrated that oral microbiome bacterial species can be transported to the gut via saliva, where they alter the composition of the gut microbiome [102]. As a result, the oral cavity may serve as a reservoir for pathogenic microorganisms that typically reside in the gut, triggering immune responses in the gut and inducing persistent inflammation [102].
Several studies have indicated a potential link between periodontal disease, oral malodor, cognitive decline, and an increased risk of developing AD [103, 104]. In this context, P. gingivalis has been demonstrated to trigger the accumulation of neurofibrillary tangles (NFTs) and Aβ plaques after inducing oral infection in murine models. In addition, research indicates that administering gingipain inhibitors orally can reduce P. gingivalis levels and decrease Aβ42 production in neural tissues of mice [105]. Additional research has demonstrated that chronic systemic P. gingivalis infection leads to Aβ accumulation in human models, specifically in the neuroblastoma cell line SH-SY5Y [106]. Moreover, intraperitoneal injection of P. gingivalis lipopolysaccharides into male C57BL/6J and SAMP8 murine models led to diminished neprilysin expression within the hippocampus [107]. In turn, reduced neprilysin levels have been linked to heightened Aβ accumulation, which constitutes a hallmark of AD [108]. The oral bacterium Aggregatibacter actinomycetemcomitans has also been associated with AD due to its capacity to trigger inflammatory responses, affect the CNS, and promote the release of Aβ [109]. Furthermore, A. actinomycetemcomitans has been found to elevate the production of inflammatory cytokines and Toll-like receptors in brain cells, leading to the increased expression of pro-inflammatory cytokines such as TNF-α, IL-6, and IL-1β [110]. Other bacterial species, including Fusobacterium nucleatum and Treponema denticola, have also been linked to AD in mouse models [111, 112].
Recent research emphasizes the possible involvement of the oral microbiome in the context of PD [113]. In fact, correlations have been identified between specific bacterial genera present in the oral cavity and the severity of depressive and anxious symptoms in individuals with PD, indicating that the oral microbiome may influence the psychiatric symptoms associated with PD [114]. In a separate preclinical study, periodontitis induced by ligatures, involving subgingival plaque, was shown to worsen motor dysfunction, dopaminergic neuron degeneration, and microglial activation in mice with PD induced by 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) [115]. This investigation revealed that the oral pathogens Veillonella parvula and Streptococcus mutans necessitate the presence of periodontitis to exacerbate motor dysfunction and neurodegeneration in MPTP-treated mice with PD. The observed effects were associated with microglial activation and the infiltration of T helper 1 (Th1) cells in the colon, ileum, brain, and cervical lymph nodes of the mice. Furthermore, neutralizing interferon-gamma (IFN-γ) showed protective effects on dopaminergic neurons in mice with PD that were treated with V. parvula and S. mutans. Figure 2 presents a hypothetical representation of the relationship between oral microbiome dysbiosis and NDDs.
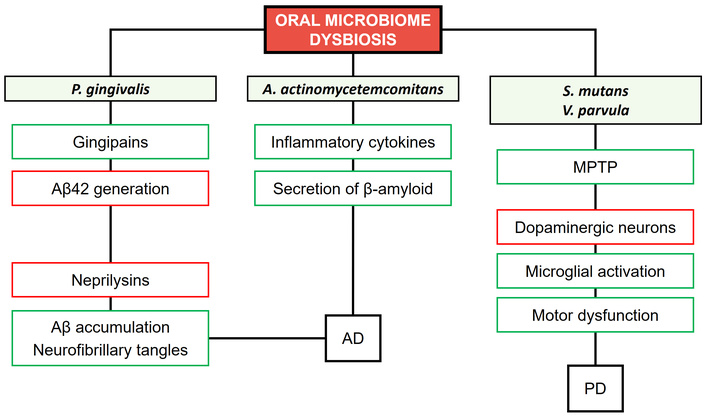
Relationship between oral microbiome dysbiosis and NDDs. Green rectangles: increase. Red rectangles: decrease. P. gingivalis: Porphyromonas gingivalis; A. actinomycetemcomitans: Aggregatibacter actinomycetemcomitans; S. mutans: Streptococcus mutans; V. parvula: Veillonella parvula; MPTP: 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine; AD: Alzheimer’s disease; PD: Parkinson’s disease; Aβ: amyloid beta.
Various pathological conditions can compromise the integrity of the oral-gut barrier, substantially increasing the risk of translocation and colonization by oral-derived microorganisms [116]. Impairments in mucosal defenses, influenced by host-associated factors such as aging and poor oral hygiene, facilitate the migration of microorganisms, while pharmacological interventions, including antibiotics, can downregulate the expression of key barrier proteins such as mucin 2, zonula occludens-1, and occludin. This disruption of the epithelial barrier, together with thinning of the mucus layer, enhances bacterial translocation and may promote the overgrowth and intestinal colonization of oral pathogens, including Streptococcus spp. and P. gingivalis [117]. Similarly, proton pump inhibitors increase gastric pH by reducing acidity, thereby allowing oral bacteria such as Rothia spp. and Streptococcus spp. to survive passage through the gastrointestinal tract [118]. Chronic oral infections, particularly periodontitis, maintain a persistent microbial load in the oral cavity, and pathogens such as P. gingivalis and Klebsiella spp. can reach the gastrointestinal tract via oral or hematogenous routes, contributing to dysbiosis and intestinal inflammation [119]. In addition to direct translocation, some oral bacteria can exploit host immune cells to disseminate systemically, a mechanism often described as a “Trojan horse”, with P. gingivalis serving as a well-characterized example of this immune-cell exploitation strategy capable of reshaping the microbial community [120].
The skin, as the largest organ in the human body, covers an average surface area of approximately 30 m2 and supports a complex and diverse microbial community. Estimates suggest that an adult’s skin hosts from 103 to 106 microorganisms, depending on the specific area being sampled [121]. The dominant bacterial phyla in the human skin microbiome include Actinomycetota, Bacillota, Bacteroidota, and Pseudomonadota. The primary genera found on the skin are Acinetobacter, Cutibacterium, Corynebacterium, Micrococcus, Staphylococcus, and Streptococcus [122]. In addition, fungi from the Ascomycota and Basidiomycota divisions are integral members of the skin microbiome, with Malassezia being the most prevalent genus, including species such as M. sympodialis, M. globosa, and M. restricta [123]. The skin also harbors a variety of viruses, primarily from the families Circoviridae, Papillomaviridae, Poxviridae, and Polyomaviridae [124]. Furthermore, a number of bacteriophages have also been identified recently as part of the skin microbiome [125].
Recent research has suggested that the skin microbiome may play a role in neuropsychiatric conditions such as NDDs, potentially through its influence on the gut microbiome [126]. This raises the hypothesis that the skin microbiome could indirectly affect the gut-brain axis, contributing to brain functional activities [127]. This interplay might influence the development of NDDs through immune-related pathways [126], and might also involve mechanisms such as increased oxidative stress, DNA damage, epigenetic alterations, impaired cell signaling, and gut dysbiosis. However, research on changes in the skin microbiome of patients with brain-related conditions remains limited. In a recent study, Arikan et al. [128] identified an association between the axillary microbiota and cognitive impairment in PD patients, indicating a potential role of the skin microbiome in cognitive dysfunction. Evidence also suggests that the human skin microbiome can produce neurotransmitters, including trace amines, dopamine, and serotonin, which may modulate classical neurotransmitter signaling or activate trace amine-associated receptors [129]. In this context, Staphylococcus species have been identified as capable of neurotransmitter production, displaying two distinct profiles: one producing tryptamine and phenylethylamine, and the other tyramine and dopamine. These profiles are determined by two bacterial aromatic amino acid decarboxylases: SadA and tyrosine/tyramine decarboxylase [129]. Moreover, a large population-based study indicated that inflammatory conditions marked by systemic TNF-α activity, such as psoriasis, may increase the risk of AD, while treatment with anti-TNF-α agents seems to reduce this risk [130]. However, other investigations have reported no significant link between psoriasis and the likelihood of developing AD or dementia [131]. Nevertheless, further research with larger sample sizes is needed to confirm these findings.
The nasal cavity harbors a wide range of microorganisms that play a role in preserving the health of the nasal mucosa and aiding the overall function of the immune system [132]. The nasal microbiome is primarily composed of several key bacterial taxa, including Actinomycetota (e.g., Corynebacterium and Cutibacterium), Bacillota (e.g., Dolosigranulum, Peptoniphilus, Staphylococcus, and Streptococcus), and Pseudomonadota (e.g., Haemophilus and Moraxella) [132]. Nevertheless, the diversity and composition of the nasal microbiome differ considerably across individuals, shaped by factors such as genetics, environmental exposures, age, lifestyle habits like smoking, and drug use [34, 35].
Although olfactory deficits and related neuropathology are well-established in the early stages of PD, evidence supporting a contributing role of the nasal microbiome remains limited [133]. For instance, one study found no marked differences in the nasal microbiome of the middle meatus between individuals with PD and healthy controls [134]. However, this does not preclude the possibility of localized microbial variations near the olfactory neuroepithelium, given that microbial communities can differ across nasal regions [135]. In AD, olfactory impairment similarly emerges years before classic symptoms and may offer predictive value when considered together with apolipoprotein E4 status [136]. Neuroimaging research has revealed reduced volumes of the olfactory bulb and tract in affected individuals compared to controls, with these changes showing correlations with cognitive performance [137]. In addition, meta-analytic data suggest that infection with the respiratory pathogen Chlamydia pneumoniae is associated with an increased risk of developing AD [138].
Building on the aforementioned, it can be stated that the involvement of the nasal microbiome in neuropsychiatric disorders constitutes a developing area of research. It is thought that the nasal microbiome may impact the established gut-brain axis via several mechanisms, such as direct interactions with the olfactory system, modulation of immune responses, and the synthesis of metabolites or neurotransmitters capable of crossing the BBB [139]. Research indicates that aging has a notable effect on olfactory function, positioning it as a potential early marker for certain NDDs [37]. Indeed, olfactory dysfunction, including anosmia (i.e., complete loss of smell) and hyposmia (i.e., reduced smell sensitivity), is commonly observed in patients affected by these conditions [37, 133]. Despite a scarcity of direct evidence linking the nasal microbiome to NDDs, some research has explored its potential involvement in AD and PD [37, 140]. In this respect, evidence suggests a possible correlation between nasal microbial communities and the onset or progression of these conditions. In particular, due to the frequent occurrence of olfactory dysfunction in the early stages of PD, Lazarini et al. [141] proposed a hypothesis suggesting that the nasal microbiome could be involved in the accumulation of misfolded α-synuclein, which is a protein primarily found in olfactory sensory neurons (OSNs). Misfolded α-synuclein is a key component of Lewy bodies, which are neuronal inclusions characteristic of PD pathology. More specifically, the hypothesis suggests that chronic dysbiosis in the nasal microbiome could trigger local inflammation, which might exacerbate damage to OSNs and lead to the accumulation of misfolded α-synuclein. This process could then spread to the olfactory bulb, substantia nigra pars compacta, and potentially other regions of the brain, ultimately contributing to the motor, cognitive, and psychiatric symptoms seen in PD patients. Furthermore, research suggests that distinct nasal microbial compositions could be associated with variations in olfactory function and cognitive performance, with certain genera such as Corynebacterium linked to lower risk of cognitive impairment and to favorable immunoregulatory profiles [142]. Mechanistically, dysbiosis in the nasal microbiome may facilitate the upward spread of pathogenic or pro-inflammatory bacteria along the olfactory pathway, promoting microglial activation, BBB dysfunction, and propagation of neurotoxic protein aggregates, which constitute processes implicated in NDDs [140]. These inflammatory and neuroimmune perturbations are likely to impact the local microenvironment of the olfactory epithelium and olfactory bulb, which are key sites of adult neurogenesis, thereby potentially compromising the proliferation and differentiation of olfactory progenitor cells and impairing the plasticity and repair of the olfactory system [140, 143]. In turn, research using intranasal delivery of therapeutic cells indicates that substances administered via the nasal route can reach the olfactory bulb and other brain regions, supporting the notion that molecules, microbial metabolites, or inflammatory mediators originating in the nasal cavity can influence neural survival, regeneration, and functional integration [143]. Figure 3 presents a hypothetical representation of direct and indirect relationships between nasal microbiome dysbiosis and NDDs.
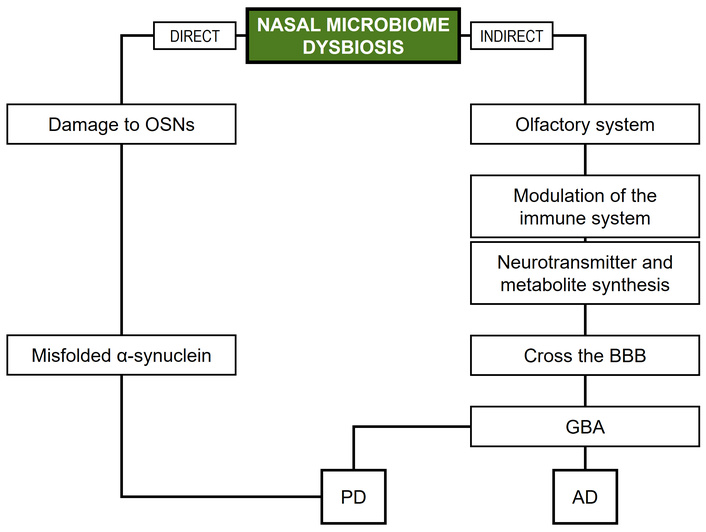
Direct and indirect relationships between nasal microbiome dysbiosis and NDDs. OSNs: olfactory sensory neurons; BBB: blood-brain barrier; GBA: gut-brain axis; AD: Alzheimer’s disease; PD: Parkinson’s disease; NDDs: neurodegenerative diseases.
Neurological disorders are an increasingly urgent global public health issue, potentially leading to substantial mortality and/or long-term disability [144]. In this context, NDDs require urgent attention due to the progressive and often rapid degeneration of neurons, accompanied by functional decline. Although numerous interventions are employed to address NDDs and promote healthy aging, their effectiveness remains limited due to factors such as the heterogeneity and diversity of populations and contexts, methodological weaknesses (e.g., insufficient sample sizes, lack of intervention descriptions, biases that lead to the overestimation of treatment effects), as well as the fact that most approved therapies only help treat the associated manifestations and fail to control the progression of the disease [145–147]. In addition, their design, evaluation, and implementation necessitate further advancements in research, particularly regarding a deeper understanding of the underlying mechanisms of pathophysiology, as well as the incorporation of multidisciplinary approaches to address these complexities.
Although several factors have been recognized as contributing to neuronal loss, including aging, genetic mutations, neuroinflammation, and mitochondrial dysfunction, research has highlighted the human microbiome as a potential pivotal player in the pathophysiology of certain NDDs [22, 79]. Indeed, evidence suggests that dysbiosis across various human microbiomes, such as the gut microbiome, oral microbiome, skin microbiome, and nasal microbiome, may play a role in the development and progression of NDDs. Moreover, research on other microbiomes, such as those of the vagina and bladder, suggests that they may also be linked to psychological states and/or psychiatric symptoms [22, 148, 149]. The existence of a microbiome-brain axis has been demonstrated in several studies, with evidence highlighting the involvement of inflammatory and immune system pathways [25, 29]. Nevertheless, further research is needed to understand how different human microbiomes interact with each other and to identify the biological mechanisms through which shifts in host-microbiota dynamics contribute to NDDs.
According to Lu et al. [150], communication between the host and the gut microbiome occurs through three primary pathways: neural, immune, and neuroendocrine. The neural pathway of the gut-brain axis involves the enteric nervous system, the vagus nerve, and the CNS. The enteric nervous system provides a habitat for gut microorganisms, which in turn are essential for its development, structural integrity, and proper functioning. Microbial metabolites can influence the distal CNS via these neural connections, transmitted through both the circulatory system and the vagus nerve. For instance, the intestinal metabolism of glutamate can rapidly signal glutamatergic neurons in the dorsal hippocampus via the vagus nerve, allowing the brain to sense gut activity and regulate taste preferences. In addition, dysregulated microbial metabolism or microbial toxins can induce neurotoxic effects in the CNS through the same neural pathways. Beyond signal transmission, the vagus nerve also regulates gastrointestinal functions, including motility, innervation of intestinal villi, and modulation of blood glucose levels. In turn, the immune pathway serves to maintain intestinal and neurological homeostasis, while the enteric nervous system can influence gut immune barriers through IL-18 signaling, with a balanced gut microbiome-immune barrier modulating CNS perception of visceral pain and emotional states via 5-hydroxytryptamine release. Both the intestinal barrier and the BBB are pivotal for controlling the passage of signals between the gut and the brain. Dysbiosis of the gut microbiome can impair the intestinal barrier by disrupting tight junction proteins, allowing inflammatory mediators to enter the nervous system and potentially contribute to cognitive deficits. On the other hand, the neuroendocrine pathway, primarily mediated by the HPA axis, links the CNS with gastrointestinal function. The HPA axis regulates responses to stress and other stimuli by releasing corticotropin-releasing factor, adrenocorticotropic hormone, and cortisol, which influence sympathetic and parasympathetic activity, metabolism, immune function, and CNS activity. Disruption of the gut microbiome has been shown to activate the HPA axis, affecting emotion, cognition, and behavior. Moreover, neurotransmitters produced by enteroendocrine cells, including ghrelin, 5-hydroxytryptamine, somatostatin, cholecystokinin, norepinephrine, adrenaline, and dopamine, represent additional key pathways through which the gut communicates with the brain. Although the role of the gut microbiome in the processes described above is relatively well established, the impact of other microbiomes in similar processes or even the interactions among them remain areas requiring further investigation. The oral microbiome, for instance, may influence the CNS through both local pathways, such as the direct influence of oral bacteria, and systemic pathways, such as the translocation of oral bacteria and their metabolites via the bloodstream. Similarly, the skin microbiome, which interacts with the immune system and can influence local inflammation, might play a role in systemic immune responses that affect CNS function. The nasal microbiome, in turn, which has direct connections to the brain through the olfactory system, may also have a role in NDDs by contributing to local inflammation and signaling pathways. Despite their potential influence, our understanding of how these microbiomes interact with the gut microbiome and each other to shape brain function and health is still in its early stages. For this reason, research is still needed to clarify the specific mechanisms through which these microbiomes affect the brain and their collective impact on NDDs. Importantly, much of the existing work has been conducted in animal models, and translating these findings to human populations is a major challenge. Therefore, as our understanding expands, it will be crucial to explore their roles in both health and disease, particularly in humans, which may provide novel insights into the diagnosis, prevention, and treatment of NDDs. Figure 4 illustrates the hypothesized influence of various human microbiomes on NDDs, integrating the available evidence into a cohesive synthesis.
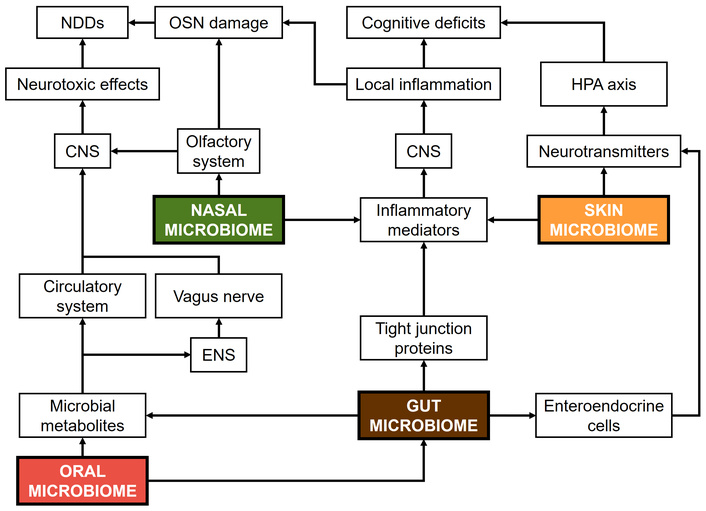
Summary of hypothetical pathways of various human microbiomes on NDDs. NDDs: neurodegenerative diseases; CNS: central nervous system; ENS: enteric nervous system; OSN: olfactory sensory neuron; HPA: hypothalamic-pituitary-adrenal.
A key question in human microbiome research is whether manipulating the microbiota can lead to effective clinical interventions that improve patient outcomes. To advance the treatment of NDDs, it is crucial to optimize microbiome functionality and ensure that therapeutic results are reproducible. In this respect, the study of the microbiome-brain axis is central for developing potential interventions that target the CNS. However, a central challenge in translating microbiota-based therapies into clinical practice lies in determining whether the microbiota alone is sufficient to enable successful treatments. As previously noted, one significant obstacle is the disparity between animal and human models, which is driven by differences in microbial composition and intestinal cell types. In addition, the lack of integrated multi-omics approaches and standardized bioinformatics tools complicates the task of elucidating microbiota functionality and ensuring reproducible results. The inherent variability of microbiomes over a lifetime, particularly of the gut microbiome, together with their individual specificity, further underscores the need for personalized therapeutic strategies that are influenced by factors such as diet and geographic location. Moreover, the dual role of certain microorganisms highlights the importance of long-term studies to better understand their specific impacts on health. In fact, longitudinal studies investigating human microbiome dynamics over the course of NDDs progression remain scarce, leaving gaps in the characterization of temporal relationships between microbial composition and clinical outcomes. In turn, behavioral phenotypes relevant to cognition, mood, and non-motor neurological symptoms are difficult to assess, further highlighting the need for innovative research. One promising approach is metabolomics, a technique that enables the comprehensive profiling of microbiome-derived metabolites and their neurological impacts [45, 151]. Prion-like protein aggregates, commonly found in NDDs, are being explored as early biomarkers, with therapeutic strategies aimed at identifying pathways to degrade these aggregates and prevent their spread [152]. Given the limitations of traditional animal models, innovative technologies such as organ-on-a-chip and computational simulations are emerging as valuable tools to improve the translational accuracy of research findings. Furthermore, randomized controlled trials are essential to rigorously assess the safety, efficacy, and durability of microbiome-targeted interventions, encompassing probiotics, prebiotics, dietary modifications, and fecal microbiota transplantation. Adopting personalized medicine frameworks that account for individual microbiome profiles, genetic predispositions, and immune system characteristics may also enable more precise and effective therapeutic strategies, ultimately improving clinical outcomes and advancing our understanding of microbiome-brain interactions, such as the exact mechanisms through which microbial communities influence neuronal signaling, neuroinflammation, and oxidative stress. Finally, it should be emphasized that, as research moves forward, it is pivotal to proceed with caution during both preclinical and clinical development, carefully monitoring and mitigating any potential adverse effects associated with microbiome-targeted therapies for NDDs. Table 2 presents a concise overview of the potential involvement of human microbiomes in NDDs.
Overview of the potential involvement of human microbiomes in NDDs.
| Microbiome | Potential involvement | Evidence | Main limitations |
|---|---|---|---|
| Gut | Modulation of neuroinflammation, oxidative stress, and neurotransmitter synthesis. ENS/vagus-mediated gut-brain communication. Immune, endocrine, and metabolite-mediated CNS effects. Influence on neurogenesis and protein aggregation. | High | High inter-individual and temporal variability. Reliance on animal models. Confounding factors (e.g., diet, drugs). |
| Oral | Circulation-mediated CNS effects via microbial metabolites. Oral pathogen-induced systemic inflammation. Modulation of CNS function via oral-gut interactions. | Medium | High inter-individual variability. Limited longitudinal human studies. Confounding factors (e.g., oral hygiene). |
| Skin | Immune-mediated modulation of neuroinflammation. | Very low | Extremely low empirical evidence. Hypothetical mechanistic pathways. |
| Nasal | Olfactory system-mediated CNS effects. | Low | Low empirical evidence. High inter-individual variability. Confounding factors (e.g., smoking). |
NDDs: neurodegenerative diseases; CNS: central nervous system; ENS: enteric nervous system.
Emerging evidence underscores the potential role of various microbiomes, including the gut microbiome, oral microbiome, skin microbiome, and nasal microbiome, in the pathogenesis of NDDs such as AD, PD, and HD. Dysbiosis across these microbiomes may influence disease progression via mechanisms such as microbial metabolite production, inflammation, amyloid accumulation, and neurotransmitter modulation. Despite promising results, the link between these microbiomes and NDDs remains complex and requires further investigation. The potential involvement of the gut-brain axis, oral-gut-brain axis, and other pathways suggests that these microbiomes may serve as both diagnostic and therapeutic targets, but further investigations are required for establishing clear causal relationships and optimizing therapeutic strategies. Thus, future studies should focus on clarifying the mechanisms by which dysbiosis in human host microbiomes impacts the pathophysiology of NDDs. Identifying reliable biomarkers associated with these microbiomes might improve diagnosis, early detection, and assessment of such diseases.
AD: Alzheimer’s disease
Aβ: amyloid beta
BAs: bile acids
BBB: blood-brain barrier
CNS: central nervous system
CVD: cardiovascular disease
HD: Huntington’s disease
HPA: hypothalamic-pituitary-adrenal
MPTP: 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine
NDDs: neurodegenerative diseases
OSNs: olfactory sensory neurons
PD: Parkinson’s disease
SCFAs: short-chain fatty acids
ABR: Conceptualization, Investigation, Writing—original draft, Writing—review & editing. JJB: Conceptualization, Investigation, Writing—original draft, Supervision. Both authors read and approved the submitted version.
The authors declare that they have no conflicts of interest.
Not applicable.
Not applicable.
Not applicable.
Not applicable.
Not applicable.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
View: 429
Download: 18
Times Cited: 0
Adnan Akhtar Shaikh ... Niveditha Nair
Salomón Páez-García ... Miguel Germán Borda
Zhengrui Li ... Jing Li
Diego Fernández-Lázaro ... Juan Mielgo-Ayuso
Ericka C. Loza López ... Felipe Esparza Salazar
Fernando Leal-Martínez, Irene Alejandra Berumen Coronado
Natalia Rojas-Sánchez ... Antonio Ibarra
