Affiliation:
Laboratory of Microbiology and Infectious Diseases, School of Veterinary Medicine, Faculty of Health Sciences, Aristotle University of Thessaloniki, 54124 Thessaloniki, Greece
Email: iangelopoulos@auth.gr
ORCID: https://orcid.org/0000-0003-0777-1820
Explor Neurosci. 2026;5:1006126 DOI: https://doi.org/10.37349/en.2026.1006126
Received: September 15, 2025 Accepted: February 02, 2026 Published: March 10, 2026
Academic Editor: Aurel Popa-Wagner, University of Medicine and Pharmacy Craiova, Romania
Hydrocephalus is a complex neurological disease characterized by abnormal cerebrospinal fluid (CSF) accumulation, ventricular enlargement, and progressive neurologic dysfunction. Existing therapies are predominantly surgical, with high complication rates, with high complication rates, prompting ongoing efforts to develop alternative modalities. This review integrates developments across mechanistic platforms and integrated disease models—including genetic and induced animals, patient-derived organoids, and organ-on-a-chip systems—to evaluate their ability to recapitulate CSF kinetics and subventricular zone (SVZ) biology. New therapies, including surgical enhancements, drugs, stem cell-based repair, and gene-targeted therapies, are discussed for translation potential. Ethics and regulatory frameworks, 3Rs, and validation and scalability issues are discussed critically. Finally, computational modeling and AI are introduced as ways to integrate multi-scale data and enable precision medicine. Each of these perspectives outlines a roadmap in which bioengineering, precision medicine, and ethical rigor converge to accelerate discovery and improve outcomes for patients with hydrocephalus.
Hydrocephalus is a neurological disorder caused by abnormalities in cerebrospinal fluid (CSF) circulation, leading to intracranial hypertension, ventricular dilation, and progressive neurological impairment if untreated [1–4]. Clinical findings range from the neonatal phase of ventriculomegaly to the presentation of idiopathic normal-pressure hydrocephalus in the elderly patient [4–7]. This etiological and temporal heterogeneity underscores the need for experimental models capable of capturing age- and cause-specific pathophysiology. The epidemiological burden of hydrocephalus is still significant globally, with congenital hydrocephalus occurring in 1–2 per 1,000 live births, with considerable long-term socio-economic burden through frequent re-hospitalization, surgical revisions, as well as intellectual or psychosocial disabilities [7–11]. At the biological level, hydrocephalus alters the structure and functioning of the ventricular zone (VZ) and subventricular zone (SVZ), which are the neurogenic layers lining the lateral ventricles. Additionally, the ependymal layer, which is of VZ origin, is made up of multi-ciliated cells with motile cilia that are responsible for the regulation of CSF circulation. Beneath this region is the SVZ, a unique neurogenic niche that is rich in neural stem cells and transit-amplifying progenitors, and is important for tissue repair [12–17]. Abnormal regulation of ciliary motility, impaired ependymal differentiation, or trophic gradient deficits caused by Foxj1, GemC1/Mcidas, L1CAM, or CCDC39 defects contribute to hydrocephalus development [18–24]. These studies provide essential mechanistic insight into ventricular physiology but do not constitute integrated models of hydrocephalus. Here, we use the term “hydrocephalus model” to denote systems that reproduce core disease hallmarks—such as sustained CSF-flow disruption and/or impaired absorption, ventriculomegaly, pressure-volume relationships, and downstream tissue responses—rather than isolated upstream mechanisms. Despite numerous advances in basic research using animal models, to date, no such system is capable of replicating human ventricular morphology or the interactions of VZ integrity with SVZ neurogenesis that contribute to hydrocephalus pathophysiology [25–28]. Surgical approaches such as ventriculoperitoneal shunt (VPS) implantation and endoscopic third ventriculostomy (ETV) remain the standard of care [29–31], but their frequent revisions and hardware-related complications indicate limited predictive value of current preclinical models for clinical translation [32–34]. These limitations have accelerated the development of novel platforms that seek to replicate key physiological elements of hydrocephalus that are more relevant to human disease. There have been recent breakthroughs in stem cell biology and bioengineering that have led to the development of patient-derived brain organoids, CSF-interactive assembloids, and microfluidic organ-chip models that seek to replicate elements of ventricular biology, cilia biology, or fluid dynamics [35–42]. These models enable human-genetic interrogation and controlled perturbation of ventricular-relevant biology for the development of human-specific models of pathophysiology, testing of causative human genetic variants, and analysis of therapeutic approaches in a controlled biomechanical setting. Despite such advancements, the challenges remain significant with respect to vascularization, reproducibility, scalability, and bioregulation. To fill the gaps in concepts and translation, this review classifies models of hydrocephalus into a continuum of models progressing from concepts developed in vivo using animal models through ex vivo, in vitro biological systems, and finally developed bioengineered microphysiological models. In this context, there are mechanistic studies that individually investigate such physical phenomena as ciliary movement or gene mutations, contrasted with integrated models of disease that replicate recognizable indicators of hydrocephalus, such as production and circulation of CSF, pressure-volume relations, ventricular dilation, or biologic responses. This distinction helps ensure that a strong conceptual framework is maintained with a logical connection between clinical physiology, biological mechanisms, and engineering models. This structured framework clarifies the role and limitations of each model class and supports a more coherent integration of mechanistic insight with translational relevance.
Experimental hydrocephalus models range from basic systems that break down individual physiological processes into isolated modules to integrated models of disease that seek to capture various characteristics of hydrocephalus. At the mechanistic end of the spectrum, models that explored ciliary movement, CSF transport anomalies, or damage to the VZ have played crucial roles in uncovering the molecular players in hydrocephalus pathogenesis [43–45]. These models offer a detailed understanding of individual pathophysiologic processes but, by definition, do not mirror the complex dynamic processes of CSF production, circulation, pressure-volume relations, or ventricular expansion. At a more complex biological network level are models of disease that employ gene manipulations or injuries that recapitulate key hallmarks of hydrocephalus. These gene models introduce disease-causing mutations for congenital hydrocephalus, while induced models employ methods of CSF obstruction or tissue damage by surgical, toxin-mediated, or hemorrhage models [27, 46, 47]. At the same time, human-relevant models like induced pluripotent stem cell (iPSC)-derived organoids and bioengineered organ-on-chip (OOC) systems are starting to leverage human cells, extracellular signals, and biophysical forces to study ventricular physiology in a more translationally relevant context [35, 48, 49]. Table 1 summarizes significant classes of hydrocephalus models with their biological basis, strengths, and limitations. While it is true that models using other species have played critical roles in understanding hydrocephalus development, genetics, or injury-mediated pathophysiology, it should also be noted that they broadly capture only selected aspects of hydrocephalus processes, do not provide human-specific hydrodynamics or VZ-ependymal interactions, or dynamic biophysical forces of CSF in humans. Thus, their extrapolability to human disease remains limited, particularly for human-specific ventricular architecture, SVZ organization, and long-term outcomes.
Comparative overview of hydrocephalus models.
| Model type | Features & mechanism | Advantages | Limitations | References |
|---|---|---|---|---|
| Genetic animal models | L1CAM, CCDC39, Rsph9, Foxj1, GemC1 mutants | Define mechanisms, mimic congenital hydrocephalus | Species-specific CSF dynamics, embryonic lethality, systemic abnormalities, and limited translation | [18–22, 24, 52, 54, 116] |
| Induced animal models | Kaolin injection, neuraminidase, intraventricular hemorrhage | Temporal control, mimic secondary hydrocephalus, reproducible | Variable phenotypes, methodology-dependent, less suited for developmental studies | [47, 61, 65–67] |
| Large animal models | Pig, dog (natural or induced hydrocephalus) | Closer ventricular size and CSF hydrodynamics to humans | Costly, ethical concerns, and fewer genetic tools | [43, 50, 68] |
| iPSC-derived organoids | Patient-derived, CRISPR-edited | Human genetic background, personalized modeling, and drug screening | Lack of vascularization and CSF flow, heterogeneity, and scalability issues | [35, 48, 49, 69, 70, 77] |
| Assembloids | Fusion of brain + vascular or choroid plexus organoids | Introduce barrier and vascular elements, model CSF interactions | Early stage, structural heterogeneity, scalability challenges | [41, 77, 134] |
| Bioengineered organ on chip | SVZ-on-chip, CSF flow chips | Dynamic CSF-like flow, biomechanical fidelity, human relevance | Fabrication complexity, standardization, and regulatory challenges | [36, 83, 84, 134, 135] |
The table summarizes major experimental platforms, detailing their origin, mechanisms, advantages, disadvantages, and representative references. In addition to traditional genetic, induced, and large-animal models, it encompasses new ones such as organoids from iPSCs, assembloids, and organ-on-chip platforms reflecting the evolution of hydrocephalus research. CSF: cerebrospinal fluid; iPSC: induced pluripotent stem cell; SVZ: subventricular zone.
Genetic models of hydrocephalus have played a crucial role in identifying the molecular and developmental processes involved in congenital hydrocephalus, especially those that relate to ciliary biology. Deficiencies in multiciliogenesis genes such as Foxj1 or regulators of the GemC1/Mcidas axis impair ependymal differentiation and ciliogenesis, supporting a mechanistic link between ciliary dysfunction and ventriculomegaly [18–21, 50]. Mutations in other axonemal proteins, including CCDC39 and RSPH1, that affect the ultrastructure of motile cilia also support the role of impaired ciliary assembly in the pathophysiology of CSF transport defects and the development of ventricular dilation [51, 52]. While these studies offer important mechanistic insight, they primarily address upstream biological processes and do not capture the integrated phenotype of hydrocephalus. Mice lacking L1CAM develop aqueduct stenosis with ventriculomegaly that closely mimics human X-linked hydrocephalus, making these mice one of the best characterized models of congenital diseases [22, 53]. Similarly, the deletion of genes in mice for CCDC39 or Rsph9 results in hydrocephalus in the early postnatal stage owing to dysfunction of ependymal cilia with severely compromised fluid movement that leads to ventricular distension [24, 52, 54]. These models provide a good representation of the pathophysiological aspects of hydrocephalus; hence, they provide crucial models through which the pathophysiological translation of structural or molecular alterations of cilia can be explored in congenital hydrocephalus. Although they provide significant information, genetic models of hydrocephalus have several challenges. These challenges include embryonic or perinatal lethality, generalized developmental anomalies, or species differences in the ventricular system or CSF hydrodynamics that make their translation complex [55–57]. More recent work in biomechanical models demonstrates the sufficiency of moving ependymal cilia in producing near-wall CSF flow and mixing, but that such processes alone do not adequately represent the human form of hydrocephalus, again emphasizing the difference between pathophysiological perturbation and whole-disease modeling [58]. Currently, genetic models, despite their utility in defining pathophysiological causality, are best interpreted alongside other approaches for whole-disease representation.
These models enable time-controlled development of hydrocephalus, allowing for the study of secondary hydrocephalus caused by trauma, infection, or intraventricular hemorrhage [28, 59]. One of the more widely used models of hydrocephalus is the kaolin injection technique, which blocks CSF transport routes, effectively causing ventriculomegaly [60, 61]. Hemorrhagic models, especially models of neonatal intraventricular hemorrhage, provide complementary information by reproducing the alterations of inflammation, fibrosis, and fluid dynamics observed in post-hemorrhagic hydrocephalus [62–64]. Mechanistic improvements have shown the existence of dose-response correlations between the degree of ventricular enlargement, gliosis, and the severity of the obstruction of CSF pathways for kaolin-based models [65, 66]. Similarly, hemorrhagic models of hydrocephalus have shown that the temporal pattern of hemorrhage significantly impacts the degree of damage to the white matter and subsequent ventricular dilation [67]. These studies emphasize the strengths of induced models of hydrocephalus in discerning distinct pathophysiologic events related to injury-based hydrocephalus. At the same time, induced models of hydrocephalus work as a holistic system for the constituents of the disease since they adequately render the major features of hydrocephalus, like ventricular dilation, disturbed CSF dynamics, inflammation, and tissue remodeling. Nonetheless, since induced models do not render the genetic or developmental basis of hydrocephalus, their applications in combating congenital hydrocephalus remain limited [28]. Large animal models such as pigs and dogs more accurately represent human ventricular anatomy and CSF dynamics, although their use is limited by cost, welfare concerns, and genetic manipulability [68]. While induced models offer a flexible, reliable, and controllable experimental system for investigating hydrocephalus caused by injury, they are excellent models of dynamic biological responses to obstruction and hemorrhage but need to be supplemented by genetic models to adequately reflect hydrocephalus pathophysiology.
Brain organoids differentiated from iPSCs are an excellent example of human-specific models that robustly replicate key features of embryonic neurodevelopment, offering patient-specific analysis of models of human diseases [35, 48, 49]. These models developed from individuals with mutations causing hydrocephalus have shown great promise by dissecting the role of genetic impairment in modulating progenitor cell activity, ciliogenesis, or VZ organization [69, 70]. CRISPR/Cas9 gene editing systems extend their utility by allowing the introduction or correction of mutations in a human genetic background [71]. Scalability in organoid production is behind the development of high-throughput approaches, such as screen testing of compounds in neural tissue [72–74], making it feasible for application in the earlier stages of therapeutic development [75]. Nevertheless, such applications are still largely biophysical since classical brain organoids are not vascularized, have poor CSF flow, and present variability between biological batches that hinders their potential to reproduce complex hydrocephalus phenotypes [38, 41, 49, 76]. New engineering techniques try to bypass such shortcomings. Assembloids, produced by merging either brain, vascular, or choroid plexus organoids, provide the capability of forming barriers, besides partially vascularized systems with better fluid interfaces [77–79]. Moreover, the development of CNS barrier-forming organoids that are able to produce CSF opens up a novel research area for studying choroid plexus biology and the process of CSF production in hydrocephalus models [77]. Vascularized organoids that support perfused microvessels make it feasible to study neuromechanical interactions in a human genetic setting [79, 80]. Despite such advancements, there is still a sense in which organoids do not adequately capture the biomechanical environment, the relationship between pressure and volume, or the CSF circulation that is observed in the context of hydrocephalus. This is where their utility is that of a specific human mechanism as opposed to a more global understanding of the disease. This is why it is important that they be used in conjunction with either bioengineered models or in vivo models.
OOC platforms combine microfluidics, extracellular matrix (ECM) with high biologic fidelity, and human-derived cells to replicate biologic microenvironments and expose cells to controlled biophysical forces [39, 81, 82]. Within the context of hydrocephalus research, OOC systems provide a unique opportunity for hypothesis testing that is difficult or impossible with live organisms or fixed cell culture systems. A novel area of research that is emerging is that of the ‘SVZ-on-Chip’ model, which places human-derived cells from the SVZ in a dynamic CSF environment mimicking perfusion conditions to investigate the role of cilia-mediated biophysical forces and ECM in the regulation of ventricular volume homeostasis [83, 84]. These systems enable detailed analysis of individual mechanisms like ciliary beat patterns in controlled fluid environments, with accurate control of those parameters that contribute to ventricular biology. More advanced microphysiological designs can impose controlled shear, pressure gradients, and directional flow—features that are difficult to achieve in static organoids and are only partially captured in animal models [38, 39, 81]. Accordingly, they complement organoids by imposing controlled biophysical boundary conditions on human-derived tissue. Hybrid organoid-on-chip systems integrating neurovascular units with microfluidic flow support such applications by offering research into neurovascular coupling and barrier integrity in a space- and time-controlled human framework [85]. In spite of such benefits, the production of an OOC is still a complex procedure with hurdles in standardization, reproducibility, and scalability [86, 87]. However, OOCs are also increasingly being investigated for their utility in preclinical drug testing applications, especially where dynamic CSF flux, controlled microenvironmental cues, and patient-derived cells provide a distinct advantage over static culture models [88–90]. Their ability to simulate fluid dynamics phenomena or multi-cellular interactions places the OOC systems as a significant additional resource for research using animal or organoid models. Overall, bioengineered OOC systems offer a human-relevant and biophysically controlled setting for the study of ventricular system physiology. These systems encompass both basic modeling solutions for certain phenomena and more complex systems-based models, but still need development for translation into practice. Their strength is realized when used either with an organoid or in vivo models for studying molecular phenomena as well as total system-based CSF dynamics.
Notwithstanding impressive progress, none is a complete replica of human hydrocephalus. Animal models clarify mechanisms but are handicapped by variation in ventricular anatomy and CSF physiology among species [25, 28]. Induction models offer control of onset and mimicking of injury-related hydrocephalus but are inapplicable to congenital and genetic hydrocephalus. Organoids offer genetic and patient specificity but have limitations in structure and reproducibility [38, 76]. OOC platforms simulate biomechanical forces and fluid flows, but are not scalable and are not universally accepted by regulators [39, 86, 87]. VZ and ependymal injury models provide an important mechanistic dimension, particularly for pediatric and post-hemorrhagic hydrocephalus, where the ependymal lining is still maturing. Disruption or denudation of the VZ frequently represents the earliest pathological event, preceding secondary alterations in the SVZ. Studies in rodent models have shown that cell-junction defects in neural stem cells lead to VZ breakdown, abnormal neurogenesis, and hydrocephalus development [91]. More recently, loss of the ependymal-regulatory protein RGS22 was demonstrated to impair multiciliogenesis and compromise ependymal integrity, resulting in ventricular enlargement and hydrocephalus [92]. Complementary in vitro VZ-injury models further support this sequence: exposure of the VZ to blood components induces epithelial disruption, glial activation, and cytopathology consistent with early post-hemorrhagic injury [93]. Together, these findings emphasize that ependymal denudation and VZ vulnerability are central to early-life hydrocephalus and highlight the need for experimental models that capture both ependymal-level and SVZ-level mechanisms. The next innovation is presumably in hybrid devices that merge organoid genetic fidelity and OOC biomechanical fidelity [94] and embed computational models to predict CSF flow perturbations and drug results [95, 96]. This will only result from global consortia and translation benchmarking programs to develop protocols, offer reproducibility, and expedite translation [87, 88]. When viewed together, the current models offer informative but incomplete data, since each type of model recapitulates certain aspects of hydrocephalus pathophysiology. Additionally, species differences make it difficult for data from animal models to be extrapolated to humans, while the lack of biomechanical realism and CSF dynamics in organoids is a major issue, with OOC systems still being technically challenging. This situation creates a need for a more convergent approach that incorporates genetic, biomechanical, and human aspects into one framework. This would become crucial for improving the predictivity of models for translation, allowing for therapeutic challenges to be evaluated in physiologically relevant contexts.
The treatment of hydrocephalus continues to center primarily upon surgical diversion of CSF. The most widely used technique is VPS, in which CSF is diverted from the ventricles to the peritoneal cavity, effectively decreasing ventricular dilatation and producing better patient outcomes. ETV, as a substitute, is used in select cases, in which a fenestration in the floor of the third ventricle is produced to bypass impaired CSF pathways [97]. Although effective, there are complications of infection, obstruction, and mechanical failure, frequently necessitating repeat surgery and substantial morbidity and health costs [33, 34]. Hydrocephalus is not, however, a disease of children and babies only. In adults, there is idiopathic normal pressure hydrocephalus (iNPH), whose classic presentation is a triad of gait disturbance, intellectual impairment, and incontinence, in a setting of ventriculomegaly and normal opening pressure of CSF [98–101]. VPS remains the initial treatment, with most reliable improvement occurring in gait and quality of life, whereas cognitive outcomes are less trustworthy [102, 103]. Long-term follow-up confirms that despite wide patient improvement, high revision frequency and complications highlight difficulties in achieving lasting control [104]. A summary of current and forthcoming therapeutic methods, status, merits, challenges, and representative references is listed in Table 2. Surgical improvements have focused on greater safety and reduced revision rates. Antibiotic-impregnated ventricular catheters, evaluated in the BASICS randomized controlled trial, lowered shunt infection rates substantially [105]. Programmable valves featuring anti-siphon mechanisms permit easier adjustment of CSF outflow and reduced over- or under-drainage risk [106]. For infants, adjunctive ETV in combination with choroid plexus cauterization (ETV-CPC) has reduced dependency on shunt in select populations [107].
Treatment approaches for hydrocephalus.
| Therapy type | Description | Current status | Advantages | Challenges | References |
|---|---|---|---|---|---|
| Surgical & shunt | VPS, ETV ± CPC, antibiotic catheters, programmable valves | Clinical use | Effective CSF diversion, reduced shunt dependence, and infection reduction | Infection, revision, and mechanical complexity | [29–34, 105, 106] |
| Pharmacological | Acetazolamide, furosemide, NKCC1 inhibitors, antifibrotics | Preclinical/Limited clinical | Non-invasive, adjunct therapy | Limited efficacy, systemic toxicity | [108–110] |
| Regenerative & cell-free | Stem cells, MSCs, EVs, ependymal reprogramming (GemC1/Mcidas) | Preclinical | Repair, neuroprotection, and restore the ependymal lining | Delivery, tumorigenicity, and scalability | [91, 113, 115] |
| Gene & molecular | CRISPR/Cas9, AAV vectors, molecular targets (NKCC1, TRPV4) | Preclinical/Experimental | Targeted therapy, disease-modifying | Immune responses, delivery challenges | [117–119] |
| Modeling platforms | Organoids, organ-on-chip, computational models | Preclinical/Regulatory | Personalized testing, patient-specific modeling | Standardization, reproducibility | [35, 39, 83, 84, 88] |
This table summarizes proven and newly emerging strategies, including surgical and shunt therapies (VPS, ETV, ETV-CPC, programmable/antibiotic valves), drug therapies (acetazolamide, furosemide, NKCC1 inhibitors, antifibrotics), regenerative and cell-free therapies (stem cells, EVs, ependymal reprogramming), molecular and gene therapies (CRISPR/Cas9, AAV vectors, ion channel targets), and bioengineered modeling platforms (organoids, organ-on-chip, computational fluid dynamics). Each entry clarifies therapeutic rationale and status. AAV: adeno-associated virus; CPC: choroid plexus cauterization; CSF: cerebrospinal fluid; ETV: endoscopic third ventriculostomy; EVs: extracellular vesicles; MSCs: mesenchymal stem cells; NKCC1: Na+-K+-2Cl– cotransporter 1; SVZ: subventricular zone; VPS: ventriculoperitoneal shunt; TRPV4: transient receptor potential vanilloid 4.
Surgical procedures being less than optimal, a pharmacological solution has been explored. Diuretics, such as acetazolamide and furosemide, reduce CSF secretion experimentally, but randomized controlled trials in pre-term infants did not reduce shunt placement and were associated with increased neurological morbidity, reflecting their narrow therapeutic window and systemic toxicity [108].
More promising reports have emerged from alternative non-surgical interventions. For example, the Drainage, Irrigation, and Fibrinolytic Therapy (DRIFT) intervention improved longer-term neurodevelopmental outcomes in infants with post-hemorrhagic hydrocephalus, in randomized controlled trials, as well as 10-year follow-up [109, 110]. At a molecular level, NKCC1 cotransporter inhibition, which is involved in most of the CSF secretion in the choroid plexus, has been shown in preclinical models treated with bumetanide to reduce CSF output [111]. Elevated transforming growth factor-beta 1 (TGF-β1) following intraventricular hemorrhage has been demonstrated to play a role in fibrotic scarring and defective CSF absorption [112]. Experimental treatment with decorin, a naturally occurring TGF-β inhibitor, blocked ventricular dilation and fibrosis in animal models, introducing a novel disease-modifying treatment [113]. Regenerative medicine approaches are beginning to reshape the therapeutic landscape. Neural stem cells, ependymal progenitors, and mesenchymal stem cells (MSCs) have demonstrated neuroprotective and reparative effects in preclinical models of germinal matrix hemorrhage-related hydrocephalus, supporting ependymal repair and reducing neuroinflammation. Translation, however, is handicapped by sourcing, delivery, immunogenicity, tumorigenicity, and dubious longitudinal efficacy [91, 114].
Recent advances in developmental neurobiology hint at a different regenerating approach through reprogramming ependymal cells via GemC1 and Mcidas pathways as a means of restoring ventricular integrity [112]. Gene and genome-guided therapies are in sight as well. L1CAM mutations have demonstrated to cause congenital hydrocephalus [22, 53]. Proof of concept trials using CRISPR/Cas9 have successfully generated hydrocephalus models by L1CAM disruption in rats, demonstrating potential for corrigibility of pathogenic variants [116]. At the same time, intraventricularly delivered adeno-associated virus (AAV) vectors have yielded stable, lifelong expression in ependymal and periventricular cells in animal models [117]. Whereas such progress has appeared, however, there remain substantial amounts of concern regarding delivery efficacy, off-targets, and immune responses [118, 119].
Progress in regenerative as well as genetic therapies is extremely dependent upon human-relevant preclinical models that are closer to mirroring hydrocephalus pathophysiology as well as the SVZ microenvironment [25, 39, 40, 82]. Organ-on-a-chip bioengineered platforms and patient-derived organoids provide novel horizons for individualized testing of treatment modalities and mechanistic exploration [120, 121]. However, challenges in reproducibility, standardization, and scale are current predominant hurdles [39, 121, 122]. Integrated advances in computational fluid dynamics (CFD) models and in vitro CSF flow setups enable simulation of patient-specific hydrodynamics, permitting testing of devices and confirmation of therapeutic hypotheses [123, 124]. From a translation perspective, regulatory agencies such as the FDA, EMA, and OECD are increasingly adopting advanced models as new approach methodologies (NAMs) for preclinical testing [86, 87, 125]. The workshops point out their potential to complement or partially replace animal experiments, as projects of international harmonization and FAIR (Findable, Accessible, Interoperable, Reusable) data principles are gaining momentum in building transparency, reproducibility, and global consistency [39]. Incorporation of hydrocephalus research in such paradigms will enhance scientific robustness and accelerate translation of experimental innovation to safe, effective therapies. Broadly, while VP shunting and ETV remain the cornerstone of treatment, constraints in durability and safety have spurred innovation in pharmacological, regenerative, and gene-based approaches. Integration of bioengineered as well as computational models, coupled with regulatory interaction, will be beneficial in converting such modalities into sustained clinical benefits.
In summary, current therapeutic approaches for hydrocephalus range from well-established surgical procedures to emerging pharmacological, regenerative, and gene-based strategies. While surgical diversion remains the clinical standard, its limitations highlight the need for complementary therapies capable of modifying disease mechanisms rather than simply controlling CSF flow. Pharmacological and regenerative interventions show promise but remain preliminary, requiring more precise human-relevant models for validation. Overall, therapeutic progress depends increasingly on integrating biological, bioengineered, and computational platforms that can predict treatment responses and bridge the gap between mechanistic discovery and clinical translation.
The ethical foundation of animal research is founded upon the 3Rs of replacement, reduction, and refinement, which are increasingly influenced by global strategies and controversies [126, 127]. These 3Rs have formed the foundation of hydrocephalus research in which animal models have been essential in discovering pathophysiological mechanisms and testing therapeutic interventions [25, 28, 128, 129]. Nonetheless, variations among species in brain structure, movement of CSF, and cell behavior limit predictive capability, and drugs and therapies successful in animals prove unsuccessful clinically [130, 131]. These constraints not only create translational barriers but also ethical concerns relating to sustained animal use of negligible clinical applicability [132, 133]. These have been met by bioengineered models like patient-derived organoids and OOC devices, being designed as human-translatable equivalents. These platforms are composed of human cells and simulate under physiological conditions like dynamic movement of CSF, forces, and ECM cues [36, 134, 135]. 3Rs alignment reduces reliance on animals [136], permitting controlled, high-throughput experiments impossible in vivo [137]. Disadvantages are technical complexity, standardization, and scalability [87, 88, 133, 138]. However, OECD and FDA/EMA initiatives announce growing interest in incorporating microphysiological systems into safety testing and drug discovery pipelines [88, 125, 139, 140]. Obstacles, however, are technological sophistication, reproducibility, and scalability [39, 138]. OECD and FDA/EMA initiatives reference enthusiasm for the incorporation of microphysiological systems into safety evaluation and compound discovery; however, harmonized validation platforms are a requirement yet to come [88, 89, 125, 139, 140]. These ethical and translational bottlenecks, along with possible solutions, are tabulated in Table 3. Moral matters reach beyond animal welfare and extend to regenerative and gene therapies, which require vigorous preclinical validation in models that are relevant to humans to guarantee patient safety [114, 118, 119]. Thus, ethical requirements and translation bottlenecks converge in a need for better models that reduce animal use and improve clinical predictivity [132, 133, 137]. Adherence to FAIR data principles and preregistration initiatives is necessary to reduce bias and increase robustness [141, 142]. Similarly, EU directives and harmonization efforts from the FDA, EMA, and OECD emphasize cross-discipline teamwork and standardization as regulatory acceptability requirements [125, 131, 139, 140]. Overall, translation barriers and ethical imperatives converge in a core need: models of relevance to humans that are 3Rs compliant, better preclinical predictors of clinical outcomes, and fully aligned with precision medicine and bioengineering. Their establishment and regulatory acceptability will become crucial in hydrocephalus research and treatment optimization, in terms of efficiency, ethicality, and effectiveness. Overall, ethical and translational considerations reveal both the strengths and limitations of current hydrocephalus models. Animal studies have been central to mechanistic discovery, yet their limited predictive value underscores the need for human-relevant platforms. Emerging organoid and OOC systems align with 3Rs principles and offer greater physiological fidelity, but challenges in standardization, validation, and regulatory readiness remain substantial. Progress in this area will rely on harmonized guidelines, FAIR data practices, and closer interaction between bioengineering, clinical science, and regulatory bodies to ensure that next-generation models are both ethically responsible and translationally robust.
Ethical and translational concerns in hydrocephalus research and potential solutions.
| Challenge | Description | Potential solutions/Innovations | References |
|---|---|---|---|
| Animal use and welfare | Ethical concerns over animal suffering and limited clinical predictability | Application of 3Rs (replacement, reduction, refinement) and transition to human-relevant models | [126, 127, 136] |
| Translational failure | Poor predictability of animal models; therapies often fail clinically | Development of patient-derived organoids and organ-on-chip systems to improve human relevance | [130–132] |
| Model complexity & regulatory hurdles | Difficulty standardizing advanced in vitro models; evolving rules for regenerative/gene therapies | Cross-disciplinary collaboration, harmonized OECD/FDA/EMA guidelines, and early regulatory engagement | [87, 89, 125] |
| Patient safety, ethics & reproducibility | Long-term safety concerns, need for rigorous validation, bias, and lack of transparency | Rigorous preclinical validation, patient-centered monitoring, open science practices, FAIR data sharing, and preregistration | [133, 141, 142] |
| Economic & accessibility | High costs limit equitable patient access | Consideration of cost-effectiveness and policies for equitable access | [147–149] |
This table compiles major ethical and translational concerns, including animal welfare, translational failure, complexity of models and regulatory hurdles, patient safety, data reproducibility, and economic accessibility, and proposes potential solutions. These are 3Rs implementation, creation of organoids and organ-on-chip models relevant to humans, collaborative research work from different disciplines, harmonized regulatory interaction, implementation of FAIR practices and preregistration, and addressing cost-efficiency and equitable accessibility. FAIR: Findable, Accessible, Interoperable, Reusable.
Translation of hydrocephalus research in the next few decades will depend upon constructing and integrating multi-scale, human-translatable models to bridge the gap between animal models and clinical translation [88, 137, 140]. Despite several decades of progress, though, the bench-to-bedside pipeline is a scattershot process in which a few bottlenecks create rough transitions from preclinical discovery to efficacious therapies. This stepwise progression is shown in Figure 1, which describes crucial checkpoints, ethical considerations, and points of convergence for how to integrate human-translatable systems to make translation faster.
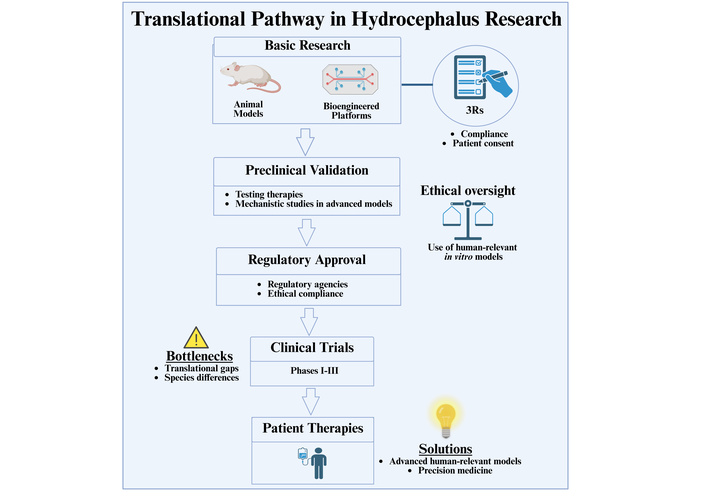
Translational route in hydrocephalus research. Stepwise summary of fundamental research (animal models and bioengineered systems) to preclinical proof, regulatory approval, and clinical investigations, ultimately culminating in patient therapy. Critical checkpoints include ethical concerns (3Rs, patient consent), regulatory advice, and determination of translational bottlenecks (species differences, predictive gaps). The intermingling of high-end human-translatable models and precision medicine strategies offers potential means of overcoming such hurdles as well as rapid acceleration of clinical translation. 3Rs: replacement, reduction, and refinement. Created in BioRender. Angelopoulos, I. (2025) https://BioRender.com/eekhg0f.
Microfluidics-based OOC models can mimic key elements of the SVZ niche—CSF dynamic flow, ECM signaling, and multicellular interactions—which enhance physiological relevance [36, 134, 135]. These models complement animal and genetic models by allowing mechanistic studies and individualized drug testing using patient-derived cells [36, 138]. At the same time, strategies for precision medicine based on iPSCs and gene editors such as CRISPR/Cas9 are providing a gateway to therapies designed specifically for individual genetic profiles, toward tailored therapies for hydrocephalus and other neurodevelopmental disorders [143–146].
Attainment of this potential will require collaborative, interdisciplinary research spanning bioengineering, neuroscience, clinical research, and regulatory science [88, 130, 140]. Standardization, validation, and scalability will be required to integrate bioengineered platforms into preclinical pipelines and regulatory processes [39, 88, 140]. The 3Rs, and ethical principles generally, remain central. Through reducing reliance upon animal models by adopting sophisticated in vitro and ex vivo systems, researchers can preserve ethical imperatives alongside greater predictive accuracy [126, 127, 132]. Development will also depend upon regulatory clarity, patient safety, and early patient and public engagement in research design [130, 141, 142]. The need for economic feasibility and equitable access must also take center stage to deliver novel therapies to patients globally [147–149].
At the same time, AI and computational modeling are increasingly viewed as transformative players in translational medicine. When patient data, multi-omics, and preclinical outcomes are integrated, AI may increase predictability, hone experimental design, and enable patient stratification [150]. The next step forward will similarly rely on international collaboration and harmonized standards. Initiatives such as the FDA, EMA, and OECD’s GIVIMP (Good In Vitro Method Practices) emphasize validation and reproducibility, while FAIR data practices and multicenter consortia are pivotal in dismantling fragmentation [140]. Table 4 summarizes these next steps, highlighting major challenges and research priorities in hydrocephalus.
Summary of future directions, challenges, and priorities in hydrocephalus research.
| Area | Future directions | Key challenges | References |
|---|---|---|---|
| Human-relevant models | Multi-scale organoids, OOC platforms | Complexity, validation, scalability | [35, 36, 39] |
| Precision medicine | iPSC and CRISPR/Cas9-based therapies, AI-integration | Data integration, personalized delivery | [143–146, 150] |
| Ethical & regulatory | 3Rs compliance, translational ethics, patient/public engagement | Regulatory approval, trial design, and long-term safety | [125–127] |
| Global collaboration & data | International consortia, FAIR data standards | Harmonization, reproducibility | [139, 142] |
| Economic & equitable access | Cost-effectiveness, global access to therapies | Infrastructure, affordability | [147–149] |
The table outlines the main future directions in hydrocephalus research, including the establishment of dynamic, multi-scale organoid and OOC models, use of methods of precision medicine such as personal therapies based on iPSCs and CRISPR coupled with AI, adherence to ethical and regulatory values such as the 3Rs and patient/public participation, and consideration of equitable access and cost-effectiveness of therapies. International collaboration, data sharing, and FAIR principles are highlighted as essential in allowing harmonization, reproducibility, and regulatory convergence. 3Rs: replacement, reduction, and refinement; FAIR: Findable, Accessible, Interoperable, Reusable; iPSC: induced pluripotent stem cell; OOC: organ-on-chip.
Translation of preclinical innovation to clinics will also require adaptive trial design and early patient engagement. Patient engagement extends ethical as well as clinical relevance [142], whereas frameworks of translational ethics emphasize sites of intersection where innovation will need to be balanced against safety in proceeding responsibly to the bedside [141]. Development of hydrocephalus research needs a paradigm shift toward dynamic, clinically relevant, and personalized models. The fusion of bioengineered developments, precision medicine, AI, and ethical rigor signifies a potential direction toward unraveling disease mechanisms as well as contributing to patient-specific therapies. Transcending current limitations, substantial progress will depend upon a holistic approach that balances scientific innovation along with translational, regulatory, and social limitations to bring about substantial therapeutic advancement.
3Rs: replacement, reduction, and refinement
CSF: cerebrospinal fluid
ECM: extracellular matrix
ETV: endoscopic third ventriculostomy
FAIR: Findable, Accessible, Interoperable, Reusable
iPSC: induced pluripotent stem cell
OOC: organ-on-chip
SVZ: subventricular zone
TGF-β1: transforming growth factor beta 1
VPS: ventriculoperitoneal shunt
VZ: ventricular zone
IA: Conceptualization, Methodology, Investigation, Writing—original draft, Writing—review & editing, Visualization, Project administration. The author read and approved the submitted version.
The author declares that there are no competing interests.
Not applicable.
Not applicable.
Not applicable.
This review article does not contain original datasets. All information discussed is available in the published literature and cited accordingly.
This work was supported by the Hellenic Foundation for Research and Innovation (HFRI), 3rd call for postdoctoral fellows (Research Grant No. 7371). The funder had no role in study design, data collection, data analysis, interpretation, writing of the manuscript, or the decision to submit the article for publication.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
View: 1237
Download: 15
Times Cited: 0
