Affiliation:
1Food and Drug Investigation Laboratory, The Indonesian Food and Drug Authority, Jakarta Pusat 10560, Indonesia
Email: alfi.sophian@pom.go.id
ORCID: https://orcid.org/0000-0002-5206-2110
Explor Foods Foodomics. 2026;4:1010140 DOI: https://doi.org/10.37349/eff.2026.1010140
Received: January 23, 2026 Accepted: March 25, 2026 Published: April 27, 2026
Academic Editor: Xianhua Liu, Tianjin University, China
Foodborne pathogen outbreaks impose a substantial and escalating burden on global public health, food systems, and economies, with the World Health Organization estimating over 600 million illness episodes and 420,000 deaths annually. Effective outbreak investigation requires harmonizing microbiological detection, molecular source tracing, and quantitative risk assessment within a single, coherent analytical architecture—a capacity that current fragmented approaches consistently fail to deliver. This review presents a novel, food-system-centered integrated framework for foodborne pathogen outbreak investigation that, for the first time, explicitly unifies conventional microbiology, molecular and whole-genome sequencing (WGS)-based typing, foodomics (metagenomics, proteomics, metabolomics), artificial intelligence and machine learning (AI/ML)-driven source prediction, geographic information systems (GIS)-based spatial epidemiology, and iterative quantitative microbial risk assessment (QMRA) within a single investigative architecture. The framework is further differentiated by a three-tiered adaptive implementation model designed explicitly for resource-limited settings and by dedicated protocols for informal food supply chains—two critical gaps absent from existing WHO/FAO and CDC/EFSA guidelines. A systematic literature search was conducted in PubMed/MEDLINE, Scopus, and Web of Science (1997–2025), with emphasis on evidence published between 2021 and 2025. The framework addresses three structural limitations of current practice: investigative fragmentation, under-integration of risk assessment, and inapplicability in low- and middle-income country (LMIC) contexts. By anchoring investigation in food and production environments rather than in clinical surveillance alone, and by embedding iterative risk assessment from the earliest investigative stage, the proposed framework supports more rapid, accurate, and equitable outbreak responses. Limitations of the review and directions for future validation research are discussed.
Foodborne diseases remain one of the most pervasive and underestimated threats to global public health. The World Health Organization estimates that approximately 600 million people—nearly one in ten individuals worldwide—suffer a foodborne illness episode annually, resulting in 420,000 deaths and the loss of 33 million disability-adjusted life years (DALYs) [1, 2]. The economic burden is equally staggering: conservative estimates place the productivity losses associated with foodborne illness in low- and middle-income countries (LMICs) alone at USD 95 billion per year [1]. Pathogens including Salmonella spp., Listeria monocytogenes, Shiga toxin-producing Escherichia coli (STEC), Campylobacter spp., and norovirus collectively account for the majority of outbreak-associated morbidity and mortality across diverse food matrices, from fresh produce and dairy to ready-to-eat meat products and seafood [3–5].
The investigation of foodborne pathogen outbreaks is inherently complex. Contamination events can originate at any node in the farm-to-fork continuum—primary production, processing, distribution, retail, or domestic food preparation—and may propagate silently across international supply chains before clinical recognition occurs [6]. Conventional outbreak investigations have historically relied on a two-pillar approach: clinical epidemiology to identify case clusters and demographic risk factors, and culture-based microbiological analysis to confirm implicated pathogens [7, 8]. While these pillars remain foundational, they are insufficient in isolation. Culture-based detection is constrained by the viable but non-culturable (VBNC) state of stressed pathogens, analytical turnaround times of several days, and limited discriminatory resolution for source attribution. Epidemiological approaches, meanwhile, depend on accurate case ascertainment and food exposure recall—both prone to significant bias and delay, particularly in diffuse, low-attack-rate outbreaks distributed across globalized supply chains [7, 8].
Advances in molecular biology and analytical sciences have transformed the theoretical capacity for outbreak investigation over the past decade. Whole-genome sequencing (WGS) has emerged as the definitive standard for pathogen strain discrimination, enabling single-nucleotide polymorphism (SNP)-level phylogenetic clustering that links cases, food isolates, and environmental strains with unprecedented precision [9–11]. The emergence of foodomics—encompassing shotgun metagenomics, proteomics, and metabolomics applied to food matrices—has further expanded the investigative toolbox by enabling culture-independent characterization of complex microbial communities, pathogen–food matrix interactions, and the biochemical signatures of contamination events [12, 13]. Concurrently, artificial intelligence and machine learning (AI/ML) algorithms trained on large genomic and epidemiological datasets are demonstrating capacity for automated source attribution and outbreak prediction [14, 15], while geographic information systems (GIS) and spatial epidemiology tools are enabling the geographic visualization of contamination spread and supply chain linkage [16].
Despite these technological advances, a critical and persistent gap remains: the absence of an integrated, operationally coherent framework that assembles these complementary methodologies into a unified investigative architecture. Current practice is characterized by fragmented workflows in which microbiological detection, molecular typing, source attribution, and risk assessment are executed as discrete, largely independent activities [17]. The consequent loss of data synergy means that high-resolution molecular evidence often fails to inform timely risk management decisions, and that quantitative microbial risk assessment (QMRA)—when conducted at all—is applied retrospectively rather than as a dynamic decision-support tool embedded in the investigative process [18, 19]. The result is that outbreak investigations routinely fall short of their potential to accelerate contamination source identification, proportionate risk communication, and preventive intervention.
A further structural gap concerns equity of application. The majority of existing frameworks—including Codex Alimentarius guidelines, the EFSA WGS framework, and CDC surveillance protocols—are designed for and validated in high-income country contexts with well-established food safety infrastructure [18, 20]. LMICs, which carry a disproportionate share of the global foodborne disease burden [1, 2], lack actionable guidance adapted to their laboratory capacities, regulatory frameworks, and food system structures—including the informal markets and street food sectors that dominate food access for hundreds of millions of people [21].
Against this background, the present review makes three distinct and original contributions to the field. First, we present what is, to our knowledge, the first investigation framework to explicitly integrate conventional microbiology, WGS-based molecular typing, foodomics (metagenomics, proteomics, metabolomics), AI/ML-driven source prediction, GIS-based spatial tracing, and iterative real-time QMRA within a single, architecturally coherent model. Second, we introduce a three-tiered adaptive implementation model that operationalizes the framework across resource settings—from foundational culture-and-epidemiology approaches available in all LMICs, to advanced WGS-GIS-AI pipelines applicable in high-resource environments—directly addressing the equity gap in existing guidelines. Third, we provide the first systematic treatment of informal food supply chain investigation within an integrated outbreak framework, including dedicated sampling and tracing protocols for traditional markets and street food sectors. By anchoring investigation in food systems and production environments rather than relying primarily on clinical surveillance, and by embedding QMRA as a dynamic iterative process rather than a retrospective addendum, the proposed framework establishes a new standard for evidence integration in foodborne outbreak investigation.
This study is a comprehensive narrative review with elements of structured synthesis, designed to develop and validate a novel integrated framework for foodborne pathogen outbreak investigation. The review synthesizes evidence across five interrelated domains: (1) conventional microbiological detection methods; (2) molecular and genomic typing techniques; (3) foodomics-based analytical approaches; (4) AI/ML and GIS-based investigative tools; and (5) QMRA and risk management. The scope was intentionally broad to capture the full methodological landscape, consistent with the objective of framework development rather than meta-analytic effect estimation.
A systematic literature search was conducted in three major electronic databases: PubMed/MEDLINE, Scopus, and Web of Science. The search covered publications from January 1997 to February 2025, with priority given to high-impact evidence published between 2021 and 2025 to ensure currency of the framework. Search terms were applied using Boolean operators (AND/OR) and included the following MeSH and free-text terms: “foodborne outbreak” OR “foodborne pathogen”; AND “outbreak investigation” OR “source attribution” OR “source tracing”; AND “whole-genome sequencing” OR “WGS” OR “foodomics” OR “metagenomics” OR “proteomics” OR “metabolomics”; AND “artificial intelligence” OR “machine learning” OR “deep learning”; AND “geographic information systems” OR “GIS” OR “spatial epidemiology”; AND “quantitative microbial risk assessment” OR “QMRA”; AND “food safety” OR “food supply chain” OR “food systems”. Additional targeted searches were conducted for informal food supply chains, LMIC food safety, and tiered laboratory frameworks. Reference lists of included reviews were manually screened for additional relevant citations.
Studies were included if they: (1) were published in peer-reviewed journals or constituted authoritative guidelines from WHO, FAO, EFSA, or CDC; (2) addressed one or more of the five framework domains defined above; (3) involved foodborne bacterial, viral, or parasitic pathogens in food matrices, food production environments, or associated supply chains; and (4) were available in English. Studies were excluded if they: (1) focused exclusively on clinical disease management without reference to food systems or investigation methodology; (2) addressed environmental pathogens not transmitted via food; or (3) were conference abstracts, editorials, or letters without primary data or substantive methodological content. No date restriction was applied to foundational methodological papers; however, for rapidly evolving areas (AI/ML, GIS, WGS), priority was given to publications from 2018 onward.
Relevant methodological information was extracted from included studies, including study design, pathogen(s) investigated, food matrices and supply chain contexts examined, analytical methods applied, source attribution approaches, risk assessment frameworks employed, and implementation requirements. For quantitative studies, performance metrics (sensitivity, specificity, turnaround time, cost) were extracted where reported. Quality of primary studies was assessed using a structured appraisal approach considering study design, sample size, methodological rigor, and relevance to framework components. Authoritative guidelines and systematic reviews were given highest evidential weight. Framework development followed an iterative process of evidence synthesis, expert judgment, and structured gap analysis. The initial database search identified a total of 412 records, of which 50 studies and authoritative guidelines met the inclusion criteria and were incorporated into the review. Of these, the distribution across the five framework domains was as follows: (1) conventional microbiological detection methods: 14 studies; (2) molecular and genomic typing techniques: 12 studies; (3) foodomics-based analytical approaches: 6 studies; (4) AI/ML and GIS-based investigative tools: 10 studies; and (5) QMRA and risk management: 8 studies. Among the 50 included references, 18 (36%) were published within the past five years (2021–2025), with the remainder retained as foundational methodological references for established techniques.
The proposed integrated framework for foodborne pathogen outbreak investigation is organized around six interconnected investigative stages—outbreak detection and hypothesis generation, food and environmental sampling, multi-layered pathogen detection, advanced analysis, source attribution and data integration, and risk-based decision-making and prevention—supported throughout by two cross-cutting analytical dimensions: AI/ML-augmented data processing and GIS-enabled spatial analysis. Critically, the framework departs from linear, sequential investigation models by adopting an iterative, bidirectional architecture in which findings from each stage continuously inform and refine preceding and subsequent stages. The result is an adaptive investigative system that improves in precision as evidence accumulates, while simultaneously enabling early-stage risk management under uncertainty. The overall framework structure is illustrated in Figure 1.
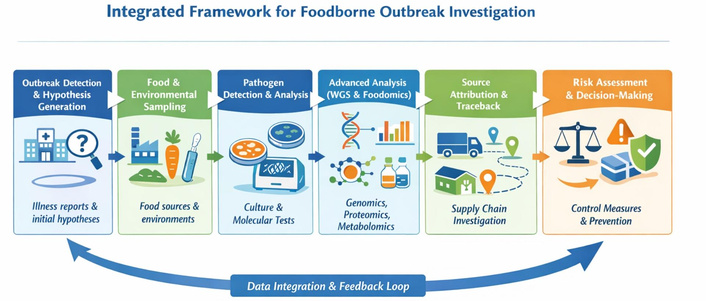
Architecture of the integrated framework for foodborne pathogen outbreak investigation. WGS: whole-genome sequencing.
The investigative process is initiated when anomalous signals—clusters of clinically similar illness, unusual patterns in food microbiological monitoring, consumer complaint surges, or automated biosurveillance alerts—exceed pre-defined thresholds. In contrast to conventional frameworks that rely predominantly on clinical case detection, the proposed framework treats food and environmental monitoring programs as co-equal signal sources. Systematic routine testing of high-risk food categories, processing environment surveillance, and water quality monitoring can detect contamination events before clinical manifestations become apparent, enabling pre-symptomatic intervention—a capacity with significant public health leverage in outbreak scenarios involving pathogens with short incubation periods or high case fatality rates [22].
Signal integration draws on multiple concurrent data streams: microbiological surveillance data, antimicrobial resistance (AMR) profiling trends, foodomics-derived early warning indicators from food matrices, and epidemiological case clustering algorithms [23]. The formal process of hypothesis generation—identifying plausible food vehicles, contamination routes, and supply chain nodes—is structured around evidence triangulation from these sources, with explicit documentation of competing hypotheses and their evidential support. This structured approach directly improves the efficiency and targeting of downstream analytical activities, reducing both false-negative risks from premature hypothesis closure and the time-to-source-identification that determines outbreak control effectiveness.
Sampling design represents the critical interface between hypothesis generation and laboratory analysis, and its quality directly determines the investigative value of all subsequent analytical activities. Sampling plans must be risk-stratified, accounting for pathogen-specific ecology, food matrix heterogeneity, contamination probability distributions along the supply chain, and the sensitivity requirements of downstream detection methods [24]. For low-prevalence pathogens such as L. monocytogenes, composite sampling strategies with statistical power calculations are essential to achieve adequate detection probabilities. The temporal dimension of sampling—capturing contamination at the moment of outbreak versus during remediation—requires coordinated, time-stamped collection protocols.
Environmental sampling within food production facilities serves as an indispensable complement to food sampling, enabling identification of harborage niches, cross-contamination routes, and persistent environmental reservoirs. Evidence from multiple outbreak investigations demonstrates that environmental persistence of L. monocytogenes in processing niches—drains, conveyor belt junctions, cold room door seals—constitutes a recurrent contamination source that can be identified and eliminated only through systematic environmental monitoring integrated with food isolate phylogenomics [25]. The framework recommends that environmental sampling be conducted in parallel with food sampling from the outset of investigations, and that environmental isolates be subjected to the same high-resolution molecular analysis as food isolates to enable phylogenomic convergence.
Informal food supply chains—encompassing traditional wet markets, street food vendors, mobile food sellers, and home-based food producers—present investigation challenges that are categorically distinct from those of formal, regulated food systems. These settings are characterized by: absence of formal traceability records or product lot coding; multiple simultaneous food sources with indeterminate provenance; inadequate temperature control and cold chain infrastructure; high vendor density and rapid product turnover; and limited regulatory oversight. These factors create an investigative environment in which standard trace-back procedures fail and in which contamination source attribution requires alternative approaches, including geo-tagged sampling surveys, vendor network mapping, phylogeographic analysis of pathogen isolates, and community-based epidemiological investigation. Dedicated sampling protocols for informal markets must account for rapid product turnover and the need for on-site, rapid screening methods to enable proportionate sampling of implicated products before they are sold or discarded [21]. The WHO FERG and FAO have recognized informal food systems as a major unaddressed gap in global food safety governance [1], and the present framework provides the first structured investigative approach for this context.
Culture-based microbiological methods constitute the foundational tier of the framework’s analytical hierarchy, serving functions that no other method currently replicates: confirmation of pathogen viability, provision of isolates for phenotypic characterization and AMR susceptibility testing, and generation of legally defensible regulatory evidence [26]. ISO and AOAC standardized methods for selective enrichment, plating, and biochemical confirmation remain the regulatory reference for all major foodborne pathogens and are indispensable for enforcement actions including product recalls and facility closures. The framework treats culture as a non-negotiable first step, even when rapid molecular screening is applied in parallel, because molecular detection of pathogen DNA in a food matrix does not confirm the presence of viable, infectious organisms—a distinction critical for both risk characterization and regulatory action [27].
The principal limitation of culture-based methods—discriminatory resolution insufficient for source attribution in genomically diverse pathogen populations—is addressed within the framework by treating culture as the isolate generation step that enables molecular and genomic analysis. The turnaround time constraint (typically 2–5 days for enrichment and confirmation) is addressed through parallel deployment of rapid molecular screening assays during the enrichment phase, ensuring that actionable presumptive results are available within hours while culture confirmation proceeds. This parallelized approach, rather than sequential application, is a key architectural feature distinguishing the integrated framework from conventional sequential workflows.
Polymerase chain reaction (PCR) and quantitative PCR (qPCR) assays provide the rapid screening capacity required for initial food vehicle exclusion and confirmation during the acute investigative phase [28]. Multiplex real-time PCR platforms targeting pathogen-specific virulence genes and serotype markers can deliver actionable results within 3–6 hours of sample receipt, enabling timely hypothesis confirmation or revision. For VBNC organisms and stressed cells—which evade culture-based detection but retain infection potential—direct PCR on food matrices provides critical detection capacity not available through conventional methods.
WGS has displaced pulsed-field gel electrophoresis (PFGE) and multilocus sequence typing (MLST) as the definitive method for outbreak strain discrimination and source attribution in high-resource settings [9–11, 29]. WGS-based SNP phylogenomics enables discrimination of strains separated by as few as 0–5 SNPs, providing resolution orders of magnitude greater than PFGE while simultaneously characterizing virulence genes, AMR determinants, plasmid content, and mobile genetic elements—data of direct relevance to both source attribution and risk characterization. National genomic surveillance networks, including GenomeTrakr (US), COMPARE (EU), and the Global Microbial Identifier initiative, have demonstrated that real-time WGS data sharing across regulatory and public health agencies can identify multinational outbreak clusters and implicated supply chains within days of initial case detection—a capability that fundamentally changes the epidemiology of outbreak investigation [9, 11]. The framework recommends WGS as the standard for source attribution whenever isolates are available, with bioinformatics pipelines standardized according to ECDC and FDA guidance to ensure cross-institutional comparability.
Foodomics approaches extend outbreak investigation beyond single-pathogen, culture-dependent paradigms by enabling comprehensive molecular characterization of entire food ecosystems [12, 13]. Shotgun metagenomics applied directly to food matrices and environmental samples provides culture-independent detection and characterization of all microorganisms present, including pathogens in VBNC states, emerging or atypical pathogens for which culture methods may be unavailable, and complex polymicrobial contamination events [30]. Metagenomic source tracking algorithms, including SourceTracker2 and FEAST, enable probabilistic attribution of contamination to environmental reservoirs by comparing microbial community signatures between outbreak isolates and potential source environments—a capability with demonstrated utility in complex, multi-source outbreaks where WGS alone cannot resolve attribution [20].
Proteomics contributes to outbreak investigations through characterization of pathogen stress response proteins—including heat shock proteins, efflux pump overexpression, and biofilm-associated surface proteins—that indicate adaptation to specific food processing environments and predict persistence behavior relevant to contamination ecology and dose–response modeling [31]. Metabolomics provides complementary information on the biochemical milieu of contaminated food matrices, identifying metabolite signatures associated with pathogen growth, stress adaptation, and toxin production that can serve as early contamination indicators in food surveillance programs. The integration of multi-omics data streams through systems biology approaches—while currently at the frontier of research application rather than routine regulatory practice—represents the investigative direction most likely to resolve the most challenging outbreak scenarios: diffuse, multi-source contamination events in complex processed food matrices with low pathogen concentrations. The framework identifies foodomics as the advanced tier component for source attribution, to be deployed when conventional microbiological and WGS-based approaches fail to achieve attribution resolution.
The application of AI/ML to foodborne outbreak investigation represents one of the most rapidly evolving areas in food safety science, with demonstrated potential to fundamentally alter the speed and accuracy of source attribution. Supervised ML models trained on curated collections of WGS data with known source metadata have demonstrated accuracy rates of 80–95% in attributing Salmonella Typhimurium and Campylobacter isolates to their reservoir source (poultry, cattle, environment) from genomic signatures alone, without requiring epidemiological case linkage [32]. Random forest classifiers applied to pan-genome data have been used to predict the likely food category implicated in L. monocytogenes outbreaks based on genomic attributes of clinical and food isolates, dramatically accelerating the hypothesis generation stage of investigations [33].
Deep learning convolutional neural networks applied to whole metagenome sequencing data have shown promise for rapid pathogen detection and community profiling in complex food matrices, reducing analysis time from days to hours while maintaining sensitivity comparable to culture-based reference methods. Graph-based ML approaches for supply chain network analysis enable mathematical modeling of product flow through complex distribution networks, identification of high-risk nodes at which contamination events would generate the observed case distribution, and prioritization of trace-back investigations in scenarios where standard epidemiological methods cannot resolve the contamination source [16]. The integration of these tools into national surveillance systems—as exemplified by the FDA’s Genome-TrackR network and the EFSA’s One Health surveillance framework—demonstrates the regulatory and operational feasibility of AI-augmented outbreak investigation. The framework incorporates AI/ML as a core component of the advanced tier, with explicit acknowledgment of the data infrastructure, standardization, and algorithmic transparency requirements necessary for responsible regulatory deployment. Challenges in model interpretability (‘black box’ predictions) and the risk of systematic bias in training datasets derived predominantly from high-income country contexts must be explicitly addressed before AI/ML tools can be deployed equitably across food safety systems globally.
GIS and spatial epidemiology provide the geographic intelligence layer that connects microbiological and molecular findings to the physical and logistical architecture of food supply chains. In outbreak investigations, GIS-based spatial cluster analysis (using tools such as SaTScan, ArcGIS, or QGIS) enables identification of case hotspots, delineation of the geographic extent of contamination, and visualization of the relationship between case distribution and the geographic footprint of implicated food supply chains or production facilities [16]. The overlay of case maps with supply chain geographic data, product distribution records, and pathogen phylogeographic data—a technique termed integrated spatial genomic epidemiology—has been used to resolve the contamination source in complex multi-state and multinational outbreaks where epidemiological evidence alone was insufficient [21].
Spatial temporal modeling using kernel density estimation and space-time interaction tests enables real-time tracking of outbreak propagation fronts and identification of secondary contamination events arising from initial source dissemination. Environmental GIS layers—including watershed mapping, agricultural land use data, and climate variables—can inform contamination risk modeling for produce-associated outbreaks where environmental pathogen reservoirs play a critical role. The framework positions GIS as a core cross-cutting component, integrated at the hypothesis generation, source attribution, and risk communication stages. However, effective GIS-based investigation requires reliable geocoding of case residential and food purchase locations, digitized supply chain distribution records, and robust spatial data infrastructure—requirements that present significant implementation challenges in LMIC contexts where these data systems are often absent or incomplete. Capacity building in GIS for food safety purposes is identified as a priority investment for LMICs seeking to develop intermediate-tier investigation capabilities.
Source attribution—the process of linking contaminated foods, environmental sources, and pathogen populations to specific supply chain nodes or production practices—represents the central analytical objective of outbreak investigation and the ultimate test of framework integration capacity. In the proposed framework, source attribution is achieved through convergent evidence integration: the systematic triangulation of epidemiological, microbiological, molecular, foodomics, GIS, and supply chain data to generate a weighted, probabilistic attribution that is continuously updated as new evidence becomes available [20, 34]. This convergent approach directly addresses the principal failure mode of single-method attribution: the inability of any individual data stream to resolve attribution in complex, multi-source scenarios with confounded evidence. WGS phylogenomics provides the strain-linkage scaffold; metagenomics source tracking provides ecosystem context; GIS supply chain analysis provides geographic plausibility weighting; and QMRA provides quantitative risk calibration of competing attribution hypotheses.
Harmonized bioinformatics workflows and standardized interpretation criteria are essential prerequisites for cross-institutional evidence convergence [20]. The framework recommends adoption of ECDC-harmonized WGS analysis pipelines (kSNP3, Snippy, or equivalent), standardized nomenclature systems (EnteroBase, MLST schemes), and transparent cluster definition thresholds (SNP cutoffs) to ensure that molecular evidence generated by different laboratories within and across jurisdictions can be meaningfully integrated. The absence of such harmonization—a persistent gap in current international practice—remains one of the principal barriers to timely multinational outbreak resolution. Table 1 provides a structured comparison of the three analytical tiers within the framework.
Comparative characteristics and roles of analytical methods within the integrated framework.
| Aspect | Conventional microbiology | Molecular methods (PCR/WGS) | Foodomics approaches |
|---|---|---|---|
| Detection target | Viable culturable pathogens | Specific genes, genomes, virulence markers | Microbial communities, metabolites, proteomes |
| Discriminatory resolution | Low-moderate (serotyping, biochemical) | High (SNP-level via WGS) | Very high (system-level, culture-independent) |
| Turnaround time | 2–5 days (culture) to weeks | Hours (qPCR) to 1–3 days (WGS) | Variable: days (metabolomics) to weeks (metagenomics) |
| Key strengths | Gold standard for viability; essential for AMR testing; regulatory acceptance | High sensitivity/specificity; strain-level tracing; real-time surveillance integration | Culture-independent; holistic ecosystem view; characterizes pathogen-food matrix interactions |
| Key limitations | Cannot detect VBNC cells; limited strain discrimination; slow | Requires reference genome databases; cannot confirm viability | Bioinformatic complexity; high cost; limited regulatory standardization |
| Regulatory status | ISO/AOAC standardized; fully accepted | Increasingly accepted; WGS mandated in EU/US for select pathogens | Research-grade; regulatory integration emerging |
| Role in framework | Foundation: confirmation, AMR profiling, isolate preservation | Core: source attribution, phylogenomics, strain linkage | Extended: risk contextualization, contamination ecology, hypothesis generation |
AMR: antimicrobial resistance; PCR: polymerase chain reaction; qPCR: quantitative polymerase chain reaction; SNP: single-nucleotide polymorphism; VBNC: viable but non-culturable; WGS: whole-genome sequencing.
As illustrated in Table 1, the three analytical tiers are not alternatives but complementary layers whose integration generates investigative capacity exceeding the sum of their individual contributions. Conventional microbiology provides the legally defensible evidentiary base; molecular methods provide discriminatory resolution for attribution; and foodomics provides ecosystem intelligence for complex scenarios and novel contamination events. The transition from sequential to parallel and iterative deployment of these tiers is the defining architectural feature of the integrated framework. Table 2 summarizes the major foodborne pathogens, their principal food vehicles, epidemiological significance, and severe outcome profiles.
Major foodborne pathogens: food vehicles, global frequency, and clinical significance.
| Pathogen | Primary food vehicle(s) | Global frequency | Severe outcomes (CFR/HUS) | Key reference(s) |
|---|---|---|---|---|
| Salmonella spp. | Poultry, eggs, fresh produce, spices | Very high (~93.8 million cases/year globally) | ~155,000 deaths/year | WHO [1]; Scallan et al. [3] |
| Campylobacter spp. | Poultry, raw milk, contaminated water | Highest in EU; ~96 million cases/year globally | Rare mortality; Guillain-Barré risk | EFSA [4]; Havelaar et al. [2] |
| Listeria monocytogenes | RTE meats, soft cheeses, smoked fish | Low incidence but highest CFR (~20–30%) | Perinatal, elderly: highest risk groups | Swaminathan and Gerner-Smidt [7] |
| Norovirus | Shellfish, fresh produce, ready-to-eat foods | Highest frequency: ~125 million cases/year | Low mortality; high transmission rate | Scallan et al. [3] |
| Staphylococcus aureus | Dairy, deli meats, bakery products | Common; frequently underreported | Self-limiting; rare severe cases | Todd et al. [35] |
| Clostridium perfringens | Cooked meat/poultry dishes, gravies | High; 2nd most common in US outbreaks | Low CFR; high morbidity burden | Scallan et al. [3] |
| Bacillus cereus | Rice, starchy foods, spices | Moderate-high; widespread in Asia | Low mortality; emetic/diarrheal syndromes | EFSA [4] |
| Unidentified agents | Multiple/unknown matrices | ~38–40% of reported outbreaks globally | Unknown; likely underestimated burden | CDC [8] |
STEC: Shiga toxin-producing Escherichia coli.
Table 2 highlights the heterogeneity of the foodborne pathogen landscape and underscores why a pathogen-agnostic, multi-method investigation framework is essential. The persistence of L. monocytogenes in processing environments and the high case fatality rate demand environmental surveillance integration not required for self-limiting Campylobacter outbreaks; the very high frequency of norovirus contamination demands molecular (RT-qPCR) screening methods unavailable through culture; and the substantial proportion of outbreaks with unidentified agents (~38–40%) demands metagenomics capacity for etiology-agnostic detection. No single-method framework can adequately serve this breadth of scenarios. The key references cited in Table 2 (WHO [1], EFSA [4], Scallan et al. [3]) have been supplemented with recent evidence where applicable. The epidemiological landscape described remains current as confirmed by the EFSA One Health Zoonoses Report 2022 [4] and the CDC Annual Surveillance Report 2022 [8], both published within the past five years. Recent systematic analyses further corroborate the frequency and severity data presented [17, 36].
Risk assessment in the integrated framework is not a terminal activity performed after source attribution but an iterative, embedded process that generates actionable risk estimates at every investigative stage. This architectural decision—distinguishing the proposed framework most sharply from existing guidelines—reflects the fundamental insight that risk management cannot wait for complete investigative resolution: every hour of delayed intervention in a high-hazard outbreak represents preventable cases. Early-stage risk estimates, while associated with higher uncertainty, can support proportionate precautionary actions (targeted withdrawal, enhanced surveillance, consumer advisories) that are far less socially and economically disruptive than the broad reactive recalls that characterize late-recognition outbreaks [18, 19].
QMRA provides the quantitative bridge between microbiological evidence—pathogen concentration, prevalence, strain virulence characteristics—and public health outcome estimation (probability of illness, hospitalization, death) [37]. In the integrated framework, QMRA models are initialized at the hypothesis generation stage using published dose–response relationships and preliminary concentration estimates, generating an initial risk profile that prioritizes investigative resources toward the highest-risk hypotheses. As culture, molecular, and foodomics data accumulate, QMRA parameters are iteratively updated: culture-based enumeration refines exposure concentration estimates; WGS virulence profiling refines dose–response parameters for specific strains; and metabolomics data on pathogen stress adaptation informs survival probability modeling under food processing conditions [38–40]. This dynamic, Bayesian-framework-compatible approach transforms QMRA from a retrospective reporting tool into a real-time decision-support instrument.
The integration of stochastic Monte Carlo simulation into outbreak QMRA—using software platforms such as @RISK, ModelRisk, or the open-source MCRA platform—enables explicit quantification and communication of uncertainty at each investigative stage, providing decision-makers with probability distributions of outcomes rather than point estimates [37]. This is particularly valuable for communicating the risk implications of partial evidence to non-specialist decision-makers under time pressure.
WGS-based virulence and AMR profiling provide hazard characterization data of direct relevance to QMRA that were unavailable to earlier investigation frameworks. The identification of virulence gene profiles (e.g., stx1/stx2 in STEC, hly/actA in L. monocytogenes, sopB/invA in Salmonella) from food isolates enables strain-specific dose–response estimation, enabling differentiation between high-virulence outbreak strains and background environmental contamination that would be conflated by culture-only analysis [39]. Similarly, AMR profiling identifies outbreak strains with resistance profiles that limit treatment options, elevating their clinical risk classification and potentially modifying the risk management response. Foodomics-derived metabolomic profiles of contaminated matrices—indicating biofilm formation capacity, heat resistance acquisition, or efflux pump upregulation—provide additional hazard characterization inputs that improve QMRA accuracy in complex food processing scenarios [40].
Risk management actions in outbreak contexts span a spectrum from consumer advisories and targeted voluntary recalls to mandatory product withdrawals, facility closures, and criminal enforcement proceedings. The proportionality of these actions to the evidence base and quantified risk magnitude is both an ethical imperative and a practical necessity: disproportionate responses incur economic harm, erode stakeholder trust, and may inhibit future voluntary cooperation with investigations [41]. The integrated framework supports proportionate decision-making by providing decision-makers with a continuously updated, evidence-graded risk characterization—distinguishing between ‘probable contamination source’ (phylogenomic linkage supported), ‘confirmed contamination source’ (culture and WGS convergence), and ‘exposure risk quantified’ (QMRA completed)—and by mapping these evidential grades to pre-defined management response protocols consistent with HACCP principles and Codex Alimentarius guidance [42]. The use of WGS evidence in enforcement proceedings has been upheld by regulatory agencies in the US, EU, and Australia, and its admissibility as legal evidence is increasingly established in international jurisprudence [9, 39].
The communication of outbreak risk findings—including their uncertainty—to diverse audiences (public, industry, regulators, media) is a distinct professional competency that the framework explicitly addresses. Evidence from risk communication research consistently demonstrates that transparent acknowledgment of uncertainty, when framed constructively and accompanied by clear guidance on protective action, maintains rather than erodes public trust [41, 43]. Conversely, premature certainty claims that are subsequently revised generate severe credibility damage and long-term harm to the efficacy of risk communication. The framework recommends staged risk communication synchronized with investigative milestones: preliminary consumer guidance at hypothesis generation; targeted recall notifications at phylogenomic cluster confirmation; and comprehensive outbreak report at QMRA completion. Visualization tools—including GIS-generated contamination maps, risk characterization matrices, and probabilistic exposure figures—are integral to the communication strategy, enabling non-technical audiences to interpret geographically and temporally complex outbreak data [44].
The ultimate measure of outbreak investigation quality is not speed of source identification but the degree to which investigation findings are translated into durable improvements in food system safety. The framework establishes a formal post-outbreak learning cycle in which root cause analysis—identifying the specific practices, equipment failures, supply chain vulnerabilities, or regulatory gaps that enabled the contamination event—generates targeted, evidence-based recommendations for prevention [35, 45–50]. These recommendations are mapped to specific intervention points in the HACCP framework, enabling food business operators to implement corrective actions within their existing food safety management systems rather than requiring the development of entirely new protocols. AI/ML models trained on aggregated outbreak investigation data can identify systemic risk patterns across food sectors and supply chains that are not apparent from individual investigation analyses, enabling prospective risk prioritization and resource allocation in food safety surveillance programs—a ‘learning system’ capacity that the framework is explicitly designed to support [32, 33].
A defining innovation of the present framework is its explicit operationalization across three resource tiers, directly addressing the equity gap that renders most existing guidelines inapplicable in LMIC contexts. The tiered model is not a simplification of the framework for lower-resource settings but a structured mapping of its core investigative principles to the methodological tools and infrastructure available at each resource level. The guiding principle is that every tier enables the same fundamental investigative objectives—detection, attribution, risk assessment, management, prevention—with methods calibrated to available capacity, and with clear upgrade pathways to more advanced tiers as capacity develops. Table 3 compares the framework against existing guidelines, and Table 4 provides a structured operational specification of each tier across investigation stages.
Comparison of the integrated framework with existing international guidelines.
| Framework feature | WHO/FAO Codex guidelines | CDC/EFSA surveillance frameworks | This integrated framework (present review) |
|---|---|---|---|
| Investigative perspective | Clinical/epidemiological | Epidemiological + limited food tracing | Food-system & production-environment-centered; integrates farm-to-fork continuum explicitly |
| Pathogen detection approach | Culture-based reference methods | Culture + selective molecular (PCR, WGS) | Tripartite: culture → molecular (PCR/WGS) → foodomics (metagenomics/metabolomics); iterative layers |
| Source attribution methodology | Epidemiological case linkage; food questionnaires | WGS clustering; SNP-based phylogenomics | WGS + metagenomics + supply chain network analysis + GIS spatial tracing; multi-evidence convergence |
| Risk assessment integration | Retrospective QMRA post-outbreak | Post-detection risk profiling | Dynamic, iterative QMRA embedded throughout investigation; real-time dose–response updating |
| AI/ML applications | Not incorporated | Emerging; not systematically integrated | Explicitly incorporated: WGS-based source prediction, deep learning for metagenomics, network analysis for supply chains |
| GIS & spatial epidemiology | Optional; not structured | Recommended for cluster detection | Core structural component: spatial cluster analysis, supply chain geography mapping, contamination front tracking |
| Resource-limited settings (LMICs) | General principles only; no tiered guidance | High-income country focus | Explicit three-tier adaptive model (foundational/intermediate/advanced) with LMIC-specific implementation guidance |
| Informal food supply chains | Not addressed | Not addressed | Dedicated section: traditional markets, street food vendors, traceability challenges in unregulated sectors |
| Novelty basis | Regulatory compliance standard | Surveillance and WGS integration | First framework to explicitly integrate foodomics + AI/ML + GIS + LMIC adaptation + informal supply chains within a single iterative investigation architecture |
AI/ML: artificial intelligence and machine learning; GIS: geographic information systems; LMICs: low- and middle-income countries; PCR: polymerase chain reaction; QMRA: quantitative microbial risk assessment; SNP: single-nucleotide polymorphism; WGS: whole-genome sequencing.
Three-tiered adaptive implementation model: investigation stages, methods, and enabling conditions.
| Investigation stage | Tier 1: foundational (all settings) | Tier 2: intermediate (moderate-resource) | Tier 3: advanced (high-resource) | Enabling conditions required |
|---|---|---|---|---|
| Outbreak detection | Passive clinical reporting; basic food inspection | Active sentinel surveillance; food monitoring programs | Integrated molecular surveillance (PulseNet-equivalent); AI-assisted signal detection | T1: Basic lab + reporting. T2: Trained inspectors, national surveillance. T3: Genomic database, IT infrastructure |
| Pathogen detection | ISO/AOAC culture methods; biochemical confirmation | Culture + PCR/qPCR; ELISA-based rapid assays | WGS (Illumina/Nanopore); metagenomics; proteomics | T1: BSL-2 lab. T2: PCR platform + reagents. T3: Next-gen sequencer, bioinformatics pipeline |
| Source attribution | Case-control interviews; trace-back of suspect food vehicles | PFGE/MLST typing; trace-forward supply chain mapping | SNP phylogenomics; metagenomics source tracking; GIS-supply chain overlay | T1: Trained epidemiologist. T2: Molecular typing capability. T3: WGS platform + GIS software |
| Risk assessment | Semi-quantitative risk ranking; HACCP review | Simplified QMRA using published dose-response data | Dynamic QMRA with real-time omics data inputs; stochastic modelling | T1: HACCP competency. T2: QMRA training + software. T3: Biostatistician + MCRA/R integration |
| Communication & decision | National authority notification; product recall protocol | Multi-agency coordination; stakeholder briefings | Real-time dashboards; GIS risk visualization; AI-assisted decision support | T1: Legal framework. T2: Inter-agency protocols. T3: Data visualization platforms |
AI: artificial intelligence; GIS: geographic information systems; MLST: multilocus sequence typing; PCR: polymerase chain reaction; PFGE: pulsed-field gel electrophoresis; QMRA: quantitative microbial risk assessment; qPCR: quantitative polymerase chain reaction; SNP: single-nucleotide polymorphism; WGS: whole-genome sequencing.
As demonstrated in Table 3, the proposed framework occupies a structurally distinct position in the landscape of outbreak investigation guidance. Its differentiation from existing frameworks is not incremental but architectural: no prior framework has simultaneously integrated all eight of the framework dimensions listed, and the combination of foodomics integration, explicit AI/ML incorporation, iterative real-time QMRA, three-tiered LMIC adaptation, and informal supply chain protocols represents a genuinely novel contribution to the field. The references underpinning the comparisons in Table 3 span both foundational guidelines (Codex [18, 42], CDC [8], EFSA [4, 20]) and recent publications (2021–2025), including Kirby and Teixeira [36] and Manning et al. [14], ensuring that the comparative analysis reflects the current state of the field.
Table 4 illustrates that Tier 1 implementation requires only a biosafety level 2 laboratory, trained microbiologists, and basic epidemiological capacity—prerequisites met by the majority of national food safety authorities in LMICs. The step-up to Tier 2 requires PCR platform access (increasingly available through regional laboratory networks and mobile laboratory programs) and a national molecular typing capability. Tier 3 advancement requires investment in WGS infrastructure, bioinformatics pipelines, and data sharing arrangements with international genomic surveillance networks. The framework deliberately avoids positioning Tier 1 as a ‘lower quality’ investigation; rather, a well-executed Tier 1 investigation using culture, epidemiology, and HACCP review will resolve the majority of food safety incidents that occur in LMIC contexts effectively and at minimal cost. Tiers 2 and 3 are invoked for complex, multinational, or high-consequence events where higher resolution is required.
The framework also recognizes that LMIC food safety capacity is not uniformly limited: many LMICs contain urban reference laboratories with Tier 2 or Tier 3 capability, surrounded by regional facilities operating at Tier 1. The tiered model supports this heterogeneity by enabling a hub-and-spoke investigation architecture, in which complex analyses (WGS, metagenomics) are conducted at central reference laboratories while primary sampling, culture, and epidemiological investigation are conducted locally—a model that mirrors the approaches recommended by WHO INFOSAN and the African Centre for Disease Control for regional outbreak investigation capacity building.
This review has several limitations that require explicit acknowledgment. First, as a narrative review with structured synthesis elements—rather than a systematic review with formal PRISMA methodology—it is subject to potential selection bias in the literature included. While the search strategy was comprehensive and conducted in multiple databases, the absence of a pre-registered protocol and formal PRISMA reporting represents a methodological limitation relative to systematic reviews. A formal systematic review with meta-analysis of specific method performance characteristics would complement the framework development objective of the present work, but was beyond its scope.
Second, the evidence base for several framework components—particularly AI/ML-based source attribution and integrated foodomics-QMRA workflows—remains largely confined to proof-of-concept and retrospective validation studies in high-income country contexts. The real-world performance of these tools in prospective outbreak investigations, particularly under the time pressures, data quality constraints, and resource limitations characteristic of LMICs, has not been systematically evaluated. The performance claims made in this review should therefore be understood as evidence-informed projections rather than empirically validated capabilities.
Third, the proposed framework is architecturally comprehensive but has not been prospectively validated in a real-world outbreak investigation. Validation—demonstrating that framework adoption improves investigation efficiency, reduces time-to-source-identification, and decreases the total outbreak case count—requires prospective implementation research that is identified as the highest priority for future work. Fourth, the framework’s treatment of informal food supply chains, while more developed than any existing guideline, is constrained by the limited peer-reviewed evidence base on formal investigation methods in these settings. Much of what is known about outbreak dynamics in informal markets derives from case reports and qualitative assessments rather than systematic investigation studies, limiting the specificity of protocol recommendations. Fifth, AI/ML models discussed in the framework are subject to rapid technological evolution; specific algorithms and platforms referenced may be superseded within the publication cycle, and the framework should be understood as articulating principles and approaches rather than endorsing specific software tools.
Foodborne pathogen outbreaks represent a persistent, evolving threat to global public health that demands investigative capacity commensurate with the complexity of modern food systems. The integrated framework presented in this review advances the field in three distinct ways. First, it resolves the fragmentation problem that characterizes current practice by establishing an architecturally coherent, iterative investigative system that integrates conventional microbiology, WGS-based molecular typing, foodomics, AI/ML, GIS, and real-time QMRA within a single framework—enabling data synergy that no component method can achieve independently. Second, it addresses the equity gap in existing guidelines through an explicit three-tiered adaptive implementation model that operationalizes the framework’s core principles across the full spectrum of resource settings, from LMICs with basic laboratory infrastructure to high-income national reference laboratories with advanced genomic and computational capabilities. Third, it provides the first systematic treatment of informal food supply chain investigation within an integrated outbreak framework, acknowledging and structurally addressing a food safety governance gap with direct relevance to the populations most burdened by foodborne disease globally.
The framework’s most consequential departure from existing guidelines is the repositioning of risk assessment from retrospective reporting tool to iterative real-time decision-support instrument, embedded at every investigative stage. This architectural change—enabling proportionate risk management under uncertainty from the earliest phase of investigation rather than requiring complete investigative resolution—has direct implications for the public health impact of outbreak responses and represents a practical advance with immediate implementability in existing regulatory systems.
Future research should prioritize five directions: (1) Prospective validation of the integrated framework in real-world outbreak investigations across diverse resource settings and food system contexts, measuring time-to-source-identification, total outbreak case counts, and cost-effectiveness as primary outcomes; (2) Development and multi-country validation of AI/ML source attribution models using balanced training datasets that include genomic data from LMIC pathogen populations, addressing the systematic geographic bias in current model training data; (3) Standardization and regulatory validation of metagenomics-based source tracking algorithms for routine outbreak investigation, including establishment of performance benchmarks, quality control standards, and interpretation frameworks; (4) Implementation research on tiered framework adoption in LMIC national food safety authority contexts, including assessment of capacity-building needs, regional laboratory network requirements, and health system integration factors; and (5) Development of formal investigation protocols for informal food supply chains, including rapid on-site screening approaches, community-based epidemiological methods, and supply chain mapping tools adapted to environments without formal traceability infrastructure. The realization of these priorities will determine whether the advances in investigative technology reviewed here are translated into the equitable improvements in global food safety outcomes that the scale of the foodborne disease burden demands.
AI/ML: artificial intelligence and machine learning
AMR: antimicrobial resistance
GIS: geographic information systems
LMICs: low- and middle-income countries
MLST: multilocus sequence typing
PCR: polymerase chain reaction
PFGE: pulsed-field gel electrophoresis
QMRA: quantitative microbial risk assessment
qPCR: quantitative polymerase chain reaction
SNP: single-nucleotide polymorphism
STEC: Shiga toxin-producing Escherichia coli
VBNC: viable but non-culturable
WGS: whole-genome sequencing
The authors wish to acknowledge the institutional support of the Indonesian Food and Drug Authority (BPOM) in facilitating this research. No external writing assistance was used. The authors are solely responsible for the content.
AS: Conceptualization, Methodology, Data curation, Writing—original draft, Writing—review & editing, Visualization. WY: Writing—original draft, Writing—review & editing. FUS: Writing—original draft, Writing—review & editing. All authors read and approved the submitted version.
The authors declare that there are no conflicts of interest.
Not applicable.
Not applicable.
Not applicable.
Not applicable.
Not applicable.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
View: 971
Download: 57
Times Cited: 0
