Affiliation:
1First Cardiology Clinic, School of Medicine, Hippokration General Hospital, National and Kapodistrian University of Athens, 11527 Athens, Greece
Affiliation:
1First Cardiology Clinic, School of Medicine, Hippokration General Hospital, National and Kapodistrian University of Athens, 11527 Athens, Greece
Email: christosfragoulis@yahoo.com
ORCID: https://orcid.org/0000-0001-5598-9950
Affiliation:
1First Cardiology Clinic, School of Medicine, Hippokration General Hospital, National and Kapodistrian University of Athens, 11527 Athens, Greece
Affiliation:
1First Cardiology Clinic, School of Medicine, Hippokration General Hospital, National and Kapodistrian University of Athens, 11527 Athens, Greece
Affiliation:
2Diabetes Center, 2nd Department of Internal Medicine, Medical School, Hippokration General Hospital of Athens, National and Kapodistrian University of Athens, 11527 Athens, Greece
Affiliation:
3First Department of Internal Medicine, Laiko General Hospital, National and Kapodistrian University of Athens, 11527 Athens, Greece
Affiliation:
1First Cardiology Clinic, School of Medicine, Hippokration General Hospital, National and Kapodistrian University of Athens, 11527 Athens, Greece
Affiliation:
1First Cardiology Clinic, School of Medicine, Hippokration General Hospital, National and Kapodistrian University of Athens, 11527 Athens, Greece
Affiliation:
1First Cardiology Clinic, School of Medicine, Hippokration General Hospital, National and Kapodistrian University of Athens, 11527 Athens, Greece
Affiliation:
2Diabetes Center, 2nd Department of Internal Medicine, Medical School, Hippokration General Hospital of Athens, National and Kapodistrian University of Athens, 11527 Athens, Greece
Affiliation:
1First Cardiology Clinic, School of Medicine, Hippokration General Hospital, National and Kapodistrian University of Athens, 11527 Athens, Greece
Affiliation:
1First Cardiology Clinic, School of Medicine, Hippokration General Hospital, National and Kapodistrian University of Athens, 11527 Athens, Greece
Explor Endocr Metab Dis. 2026;3:101461 DOI: https://doi.org/10.37349/eemd.2026.101461
Received: December 13, 2025 Accepted: February 04, 2026 Published: March 08, 2026
Academic Editor: Victor Gault, Ulster University, United Kingdom
The article belongs to the special issue Innovative Strategies for Diabetes and Metabolic Disorders: Current and Future Directions
Diabetes and metabolic dysfunction represent growing global health challenges, and current therapies mitigate hyperglycemia without sufficiently altering the underlying disease processes. Rapid advances in regenerative medicine, gene editing, and nano-enabled drug delivery are reshaping therapeutic possibilities, offering the potential to restore β-cell function, enhance insulin sensitivity, and personalize care at an unprecedented scale. This review synthesizes emerging therapeutic strategies with the greatest translational promise, including stem cell-derived islet replacement, immune-evasive encapsulation devices, CRISPR-based gene correction, and targeted or glucose-responsive nanocarriers engineered for noninvasive insulin delivery. Complementary advances in multi-omics profiling, proteogenomics, microbiome science, and artificial intelligence are enabling stratification of patients based on molecular signatures, optimizing therapeutic selection, and predicting clinical outcomes. Despite remarkable breakthroughs—such as the early-phase clinical success of stem cell-derived β-cell replacement and organ-targeted lipid nanoparticles (LNPs)—significant barriers remain, including immunogenicity, off-target editing, scalability, ethical concerns, and long-term safety. By integrating mechanistic insights with translational developments, this review outlines a forward-looking perspective on next-generation treatments poised to shift diabetes care from glycemic management toward disease modification and personalized metabolic restoration.
Diabetes mellitus and its associated metabolic disturbances constitute one of the most pressing global health challenges of the 21st century. The global prevalence of diabetes has more than doubled in recent decades among adults, rising from approximately 4.6% (151 million) in 2000 to 10.5% (537 million) in 2021, driven by rapid urbanization, sedentary lifestyles, population aging, and increasing rates of overweight and obesity [1, 2]. Type 2 diabetes (T2D), which represents most of all cases worldwide, is now understood as a key clinical manifestation of metabolic dysfunction, rather than a singular disorder of glucose metabolism. This broader construct integrates insulin resistance (IR), chronic inflammation, ectopic lipid deposition, and dysregulated energy homeostasis, all of which contribute to cardiovascular, renal, hepatic, and neurologic complications that markedly increase morbidity, mortality, and healthcare burden [3, 4]. Metabolic dysfunction refers to impaired metabolic processes that disrupt normal energy homeostasis, including IR, dysregulated glucose and lipid metabolism, chronic inflammation, and ectopic fat accumulation, often culminating in T2D and related cardiometabolic disorders [1].
Despite meaningful therapeutic advances—including glucagon-like peptide-1 receptor agonists (GLP-1RA) and sodium-glucose cotransporter 2 inhibitors (SGLT2i)—current treatments predominantly target hyperglycemia rather than the deeper metabolic pathophysiology. Clinical goal attainment remains suboptimal: fewer than one-quarter of patients globally achieve recommended targets for glycemia, blood pressure, and lipids, with persistent ethnic and socioeconomic disparities [3, 5]. These gaps underscore the need for disease-modifying therapies capable of restoring β-cell health, improving insulin sensitivity, reducing systemic inflammation, and preventing long-term organ damage.
Emerging scientific developments now offer unprecedented opportunities to address the root causes of diabetes. Advances in stem cell biology are enabling the generation of functional insulin-secreting β-cells for transplantation. Breakthroughs in gene-editing technologies, especially CRISPR-based platforms, allow direct correction of pathogenic mutations and modulation of metabolic pathways. In parallel, innovations in nanotechnology have catalyzed the development of noninvasive, targeted, and glucose-responsive drug delivery systems that may revolutionize therapeutic administration. Finally, integration of multi-omics analyses—including genomics, proteomics, metabolomics, and microbiome profiling—together with artificial intelligence (AI) is providing deeper insights into patient heterogeneity and supporting the emergence of precision therapeutics.
This review synthesizes these transformative developments in regenerative medicine, gene therapy, nano-enabled drug delivery, and precision medicine. By emphasizing innovation, translational potential, and future clinical directions, the review highlights how next-generation therapeutic strategies may shift diabetes management from symptomatic glucose control toward true metabolic restoration and personalized disease modification. Throughout this review, we explicitly distinguish between well-established therapeutic strategies supported by clinical evidence and emerging or experimental approaches that are currently based on preclinical or early-phase translational data. Unlike existing reviews that primarily focus on single therapeutic domains—such as stem cell replacement, gene editing, or nanomedicine—this review adopts an integrative framework that unifies regenerative medicine, genetic engineering, nano-enabled drug delivery, and multi-omics–driven precision medicine. By examining how these approaches converge at mechanistic, translational, and clinical levels, we aim to provide a holistic perspective on next-generation disease-modifying therapies for diabetes and metabolic dysfunction.
This manuscript is a narrative review that synthesizes key preclinical and clinical evidence on regenerative medicine, gene editing, nano-enabled drug delivery, and precision-medicine approaches for diabetes and metabolic dysfunction.
Literature searches examined innovative regenerative, genetic, and nano-enabled therapeutic strategies for diabetes and metabolic dysfunction. Databases included MEDLINE, PubMed, EMBASE, Web of Science, Cochrane Central Register of Controlled Trials, and reference lists of relevant reports. Keywords comprised “diabetes mellitus”, “metabolic dysfunction”, “stem cell therapy”, “gene therapy”, “CRISPR”, “β-cell regeneration”, “nanomedicine”, “drug delivery systems”, “personalized medicine”, “multi-omics”, “precision therapeutics”, combined with Boolean operators AND/OR. The search strategy was designed to identify representative, high-impact, and recent studies rather than to exhaustively capture all available literature. English-language articles from 2005 onward were included, with the final search in December 2025.
A comprehensive literature search was conducted to identify studies reporting emerging therapeutic strategies for diabetes and metabolic dysfunction, including stem cell-based therapies, gene-editing approaches, nanotechnology-enabled drug delivery systems, and precision medicine applications. Eligible studies included original research articles, clinical trials, preclinical experiments, and high-quality review articles published in peer-reviewed journals. Priority was given to high-quality systematic reviews, meta-analyses, landmark preclinical studies, and recent clinical trials with clear translational relevance. Emerging or experimental approaches were included when supported by robust mechanistic data or early-phase clinical evidence. Studies were excluded if they lacked relevance to innovative therapeutic mechanisms, provided insufficient methodological detail, or focused solely on conventional antihyperglycemic medications without mechanistic or translational implications. Only English publications were considered.
Data extraction was performed independently by two reviewers (MKS, CF) from eligible original studies, meta-analyses, and systematic reviews using standardized forms, with cross-checking for accuracy. Study quality was considered. Extracted information included study design, population or model characteristics, therapeutic modality, molecular or metabolic mechanisms, primary outcomes, safety findings, and limitations. Discrepancies in study inclusion or data interpretation were resolved through discussion and consensus. This structured approach ensured that the synthesized evidence reflected high-quality, mechanistically informative, and clinically relevant advancements aligned with the aims of this review.
The development and progression of T2D involve complex interactions among IR, β-cell dysfunction, chronic inflammation, oxidative stress, ectopic lipid deposition, and alterations in the gut microbiome. A detailed understanding of these mechanisms is essential for designing innovative therapeutics aimed at restoring metabolic homeostasis rather than simply controlling hyperglycemia.
IR is a central feature of metabolic dysfunction, often preceding the onset of T2D. IR is strongly associated with visceral adiposity, metabolic dysfunction-associated steatotic liver disease (MASLD), and reduced skeletal muscle mitochondrial capacity [2]. Ethnic variation plays a critical role; South Asian populations, for example, experience disproportionately high IR at lower body mass index (BMI) thresholds due to reduced lean mass and increased visceral adiposity [3].
At the molecular level, IR results from impaired IR substrate phosphorylation, defective PI3K/Akt signaling, and reduced GLUT4 translocation in adipose and muscle tissue [4]. Epigenetic mechanisms also contribute: hyperglycemia-induced downregulation of TET3 has been shown to dysregulate β-cell gene expression across generations [5]. These pathogenic insights underpin therapeutic strategies targeting Akt activation, epigenetic modulation, and mitochondrial restoration.
Obesity-induced inflammation is a major driver of IR. Hypertrophic adipocytes recruit macrophages and activate inflammatory pathways such as NF-κB, JNK, and the NLRP3 inflammasome, leading to the secretion of cytokines, including TNF-α and IL-1β, that impair insulin signaling [6]. Systemic inflammation contributes to metabolic target organ damage such as nephropathy, retinopathy, neuropathy, and impaired wound healing. For example, activation of STAT3 signaling has been implicated in poor immune responses and delayed tissue repair in diabetic foot ulcers [7]. These discoveries provide a strong rationale for novel anti-inflammatory biologics and gene-editing strategies targeting inflammatory cascades.
Nutrient excess and hyperglycemia increase mitochondrial reactive oxygen species (ROS) generation, overwhelming antioxidant defenses and amplifying metabolic dysfunction [8]. ROS activate stress-signaling pathways such as JNK and NF-κB and impair lipid metabolism through mTORC1 hyperactivation. Oxidative stress also drives β-cell dysfunction, renal fibrosis, and retinal microvascular damage, making antioxidant and mitochondrial-targeted therapies attractive metabolic interventions.
Ectopic accumulation of lipotoxic intermediates—including diacylglycerol, ceramides, and long-chain acyl-CoAs—disrupts insulin signaling by activating PKC isoforms that inhibit IRS-1 and Akt phosphorylation [2]. Overactivation of mTORC1 promotes lipogenesis, β-cell stress, and worsening IR. Therefore, therapeutic modulation of ceramide synthesis, PKC inhibition, and mTOR signaling represents an emerging strategy to reverse metabolic dysfunction at its biochemical core.
Alterations in gut microbiota composition are increasingly recognized as contributors to IR and chronic inflammation. High-fat diets elevate circulating lipopolysaccharides (LPSs), triggering TLR4-mediated inflammatory signaling, metabolic endotoxemia, and impaired insulin action [9]. Increased intestinal permeability further exacerbates systemic inflammation. These findings support novel microbiome-modulating therapies—including engineered probiotics, dietary extracellular vesicles (EVs), and microbial metabolites—as potential modulators of metabolic health.
Progressive loss of β-cell function and mass is a hallmark of T2D. β-cell dysfunction arises from glucotoxicity, lipotoxicity, oxidative stress, and chronic inflammation, which impair insulin gene transcription factors such as PDX1 and MAFA [10]. Upregulation of stress markers such as ALDH1A3 indicates β-cell dedifferentiation, while β-cell apoptosis, senescence, pyroptosis, and ferroptosis all contribute to cellular decline. These mechanisms directly motivate regenerative strategies such as stem cell-derived β-cell therapy, β-cell reprogramming, and genetic enhancement of β-cell resilience.
Stem cell-based therapies represent one of the most promising and innovative approaches for achieving durable β-cell replacement and restoring physiological glucose regulation in diabetes. Advances in pluripotent stem cell engineering, differentiation protocols, encapsulation devices, and immune-modifying strategies have accelerated translation from experimental models to early clinical trials. Throughout this review, therapeutic strategies are discussed in relation to their primary relevance to type 1 diabetes (T1D), T2D, or advanced insulin-deficient states where pathophysiological features overlap.
Stem cell-derived β-cell replacement is primarily relevant to T1D and to advanced T2D characterized by severe β-cell loss and insulin deficiency, rather than to early insulin-resistant stages of T2D. In T1D and advanced T2D, progressive β-cell loss results from autoimmune destruction, glucolipotoxicity, chronic inflammation, and metabolic stress. Because an estimated 40–60% of β-cell mass is lost before clinical diagnosis [11], restoring functional β-cell numbers is a central goal of regenerative medicine. Human pluripotent stem cells (hPSCs) offer a renewable source of β-like cells capable of glucose-stimulated insulin secretion [12]. Innovations in transcription factor programming (e.g., PDX1, NGN3, MAFA) and three-dimensional culture systems have improved the maturation, functionality, and safety of stem cell-derived islets. As shown in Figure 1, stem cell-based strategies aim to restore functional β-cell mass through differentiation, immune protection, and encapsulation approaches.
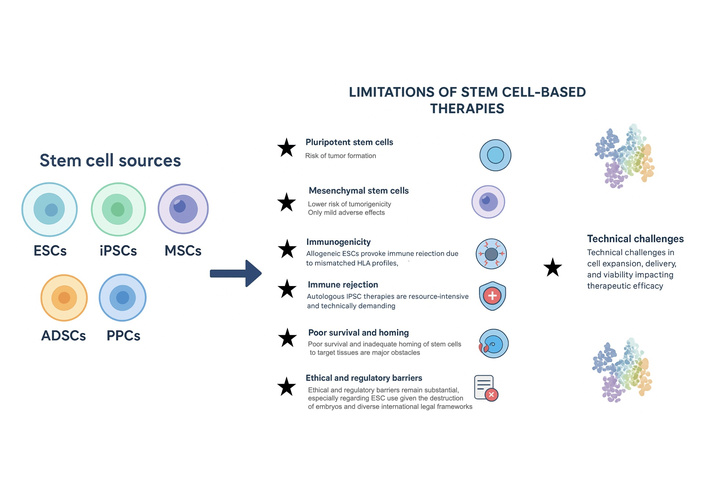
Sources of stem cells and major limitations of stem cell-based therapies. Overview of commonly used stem cell types—including embryonic stem cells (ESCs), induced pluripotent stem cells (iPSCs), mesenchymal stem cells (MSCs), adipose-derived stem cells (ADSCs), and perinatal progenitor cells (PPCs)—and key challenges limiting their translational application. Major barriers include tumorigenicity in pluripotent cell derivatives, immunogenicity due to mismatched HLA profiles, immune rejection in autologous and allogeneic settings, inadequate survival and homing after transplantation, technical difficulties in large-scale expansion and delivery, and ethical or regulatory concerns, particularly regarding ESC use. This figure mainly covers preclinical and early translational ideas; we still need to confirm how relevant they are to human clinical practice.
Emerging preclinical data suggest that further optimization of differentiation protocols may enhance β-cell maturity and functional stability; however, long-term efficacy in humans remains to be established. Directed differentiation protocols now mimic embryonic pancreas development by sequentially activating signaling pathways such as activin A, WNT, and retinoic acid to generate definitive endoderm, pancreatic progenitors, and finally insulin-secreting β-like cells. Recent advances have enhanced β-cell maturity through suppression of ALDH1A3, incorporation of endothelial and mesenchymal support cells, and adjustment of metabolic cues that drive oxidative phosphorylation, a hallmark of mature β-cells [13, 14]. Single-cell transcriptomic profiling has enabled refinement of these protocols by identifying off-target populations such as enterochromaffin-like cells and undifferentiated clusters that may pose safety risks [15].
Immune-evasive strategies and encapsulation technologies are primarily designed to address autoimmune rejection in T1D, although they may also mitigate alloimmune responses in allogeneic transplantation settings relevant to advanced T2D. Immune rejection remains a major barrier to β-cell replacement therapy. Encapsulation devices—such as alginate hydrogels, planar macroencapsulation systems, and nanoporous membranes—aim to protect transplanted cells from host immune attack while permitting nutrient, oxygen, and insulin exchange [16]. Chemical modification of alginate polymers with zwitterionic or ultrapurified variants has been shown to reduce fibrosis and foreign-body responses. Parallel strategies include gene-editing approaches to produce immune-evasive stem cell lines that lack HLA class I/II molecules or express “don’t eat me” signals like CD47 [17]. While these approaches directly target autoimmune mechanisms characteristic of T1D, their broader applicability in T2D is contingent upon disease stage and transplantation context. These next-generation platforms aim to eliminate the need for chronic immunosuppression. These immune-evasive and encapsulation-based strategies remain experimental, with human relevance and long-term safety yet to be fully determined.
Early-phase clinical studies have demonstrated the feasibility and partial restoration of endogenous insulin production using stem cell-derived pancreatic progenitors. Well-established clinical evidence from early-phase human studies demonstrates that stem cell-derived pancreatic progenitors can engraft and produce detectable C-peptide in selected individuals with T1D. ViaCyte’s PEC-01 cells encapsulated in macro-devices achieved engraftment and C-peptide production in individuals with T1D [18]. Clinical trial data indicate that stem cell-derived islet replacement can substantially reduce exogenous insulin requirements in carefully monitored settings. Vertex Pharmaceuticals’ VX-880, a stem cell-derived fully differentiated islet therapy delivered via portal vein infusion, has shown particularly promising results: treated patients achieved substantial reductions in exogenous insulin use and improvements in glycemic control, including instances of insulin independence [19]. These clinical approaches have been developed mainly for individuals with T1D; however, they may also be applicable to selected patients with advanced T2D who exhibit profound β-cell failure. Although these findings represent major milestones, questions remain regarding long-term graft durability, immune rejection, and optimal implantation sites. Key quantitative clinical outcomes from early-phase stem cell-derived β-cell replacement studies are summarized in Table 1.
Early clinical outcomes of stem-cell-derived β-cell replacement therapies.
| Study/Program | n | Follow-up | Baseline vs. post-treatment C-peptide | Insulin reduction | HbA1c change |
|---|---|---|---|---|---|
| ViaCyte PEC-Encap (VC-01) [18] | ~20 (early-phase cohorts) | Up to 12–24 months | Baseline: undetectable → Post-treatment: generally low or undetectable due to limited engraftment | No sustained insulin independence; minimal or no durable reduction | No consistent HbA1c improvement reported |
| ViaCyte PEC-Direct (VC-02) [18] | ~17 | Up to 12 months | Baseline: undetectable → Post-treatment: detectable C-peptide in a subset of patients | Partial insulin dose reductions in selected individuals | Modest HbA1c reductions in responders |
| Vertex VX-880 [19, 87] | n = 12 (actual phase 1/2 cohort); specific data | 90–360 days | Baseline: undetectable → Post-treatment: robust fasting and stimulated C-peptide | Insulin independence in 2/6 at 1 year | HbA1c achieved levels < 7.0% to near-normal range (70–180 mg/dL) |
| Vertex VX-264 (encapsulated) [19, 87] | Phase 1 ongoing | Interim | Data pending | Data pending | Data pending |
Reported outcomes reflect early-phase, small-cohort studies with limited follow-up; results should be interpreted cautiously, as long-term durability, safety, and generalizability remain under investigation. HbA1c: glycated hemoglobin.
The potential role of β-cell replacement in T2D is therefore likely to be limited to late-stage disease, underscoring the need for careful patient selection. Current limitations include incomplete β-cell maturation, risk of teratoma formation from undifferentiated cells, limited oxygenation and vascularization of encapsulated grafts, and potential immune recognition of edited or foreign cell lines. Efforts to overcome these challenges include oxygen-generating biomaterials, vascularized scaffolds, controlled-release proangiogenic factors, and organ-specific implantation strategies (e.g., omentum, subcutaneous, intramuscular). The integration of gene editing, single-cell omics, and advanced biomaterials is expected to enable the next generation of safe, durable, and scalable β-cell replacement therapies (Figure 1).
Gene therapy and gene editing have emerged as transformative strategies for diabetes and metabolic dysfunction, offering the potential to correct pathogenic mutations, modulate metabolic pathways, enhance β-cell resilience, and restore endogenous insulin production. Recent advances in CRISPR-based tools, viral and non-viral delivery technologies, and cell-type–specific promoters have accelerated translation toward clinical application. Figure 2 summarizes the principal gene therapy and gene-editing approaches being explored to modulate insulin production, immune responses, and metabolic pathways.
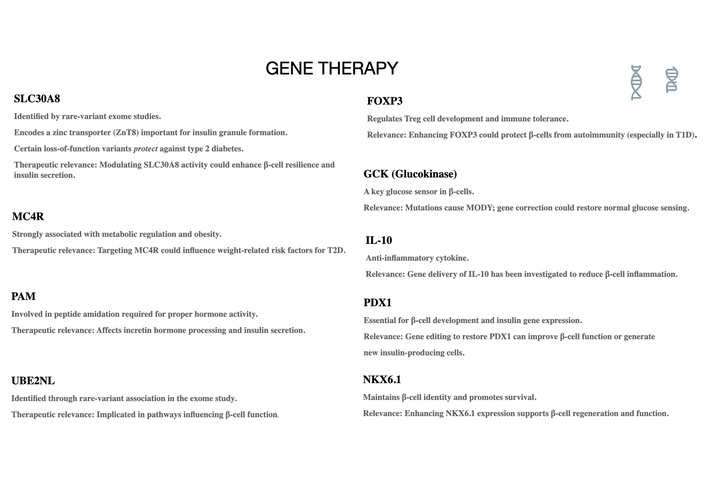

Key gene targets under investigation for gene therapy in diabetes and metabolic dysfunction. Summary of therapeutic gene targets with demonstrated relevance to β-cell function, insulin secretion, immune regulation, and metabolic homeostasis. These include SLC30A8 (zinc transporter ZnT8 involved in insulin granule formation), MC4R (metabolic regulation and obesity), PAM (hormone amidation and incretin processing), UBE2NL (β-cell functional pathways), FOXP3 (T-regulatory cell maintenance and immune tolerance), GCK (glucose sensing and MODY-associated variants), IL-10 (anti-inflammatory modulation), PDX1 (β-cell development and insulin gene expression), and NKX6.1 (β-cell identity and survival). These genes represent high-value candidates for CRISPR-based correction, RNA therapies, or targeted gene modulation. This figure outlines primarily preclinical and early translational concepts, with their relevance to human clinical settings yet to be determined.
Well-established genetic evidence supports the role of monogenic defects in specific forms of diabetes, providing a clear rationale for targeted gene-based interventions. The genetic architecture of diabetes involves numerous loci affecting β-cell function, insulin signaling, immune regulation, and metabolic homeostasis. Monogenic disorders such as maturity-onset diabetes of the young (MODY) and neonatal diabetes are caused by highly penetrant variants in genes including GCK, HNF1A, INS, and KCNJ11 [20]. Additionally, common polygenic variants contribute to β-cell vulnerability and IR. Gene therapy offers a precise strategy to address these defects by restoring gene function, silencing pathogenic alleles, or enhancing protective pathways in metabolic tissues.
Early-phase preclinical studies indicate that CRISPR-based correction of diabetes-associated mutations can restore insulin secretion in animal models. CRISPR/Cas9 has revolutionized metabolic disease research due to its simplicity and programmability. Preclinical studies have demonstrated the feasibility of correcting INS and GCK mutations, improving insulin secretion, and restoring normoglycemia in rodent models [21]. However, classical CRISPR introduces double-strand breaks, raising concerns about off-target activity and genomic instability.
To overcome these limitations, next-generation tools such as base editors and prime editors have been developed. Base editors allow precise conversion of single nucleotides without double-strand breaks, making them ideal for the correction of MODY-related mutations [22]. Prime editing expands this capability to include small insertions, deletions, and all base transitions, offering a more versatile platform for repairing diverse mutations associated with diabetes and metabolic disorders.
Efficient and tissue-specific gene delivery remains a major challenge. Adeno-associated virus (AAV) vectors have been widely studied due to their safety and tropism for hepatocytes, pancreatic cells, and muscle tissue [23]. AAV-mediated delivery has successfully reversed hyperglycemia in diabetic mouse models by enhancing insulin expression, inhibiting glucagon signaling, or improving hepatic glucose metabolism.
However, AAV vectors have size limitations and may trigger immune responses. As a result, non-viral LNPs have gained traction. LNPs can deliver mRNA, CRISPR components, or regulatory RNAs to metabolic tissues and have already achieved clinical success in other therapeutic areas. Organelle- or tissue-targeted LNPs engineered with specialized lipids or ligands can selectively home to pancreatic islets, adipose tissue, or liver, offering a precision platform for metabolic gene therapy [24, 25]. However, these approaches remain experimental, and their safety, durability, and clinical feasibility in humans require further validation.
Gene editing has enabled reprogramming of non-β-cell populations into insulin-producing cells. Forced expression of pancreatic transcription factors such as PDX1, NGN3, and MAFA in hepatic or gastrointestinal cells can induce β-like phenotypes capable of secreting insulin in response to glucose [26]. Similarly, CRISPR-mediated suppression of β-cell stress pathways (e.g., TXNIP) has been shown to enhance β-cell survival and insulin secretion under metabolic stress [27]. These approaches offer promising avenues to increase functional β-cell mass without the need for stem cell transplantation. Such reprogramming strategies may have shared relevance across T1D and insulin-deficient T2D, as both conditions ultimately converge on insufficient functional β-cell mass. Human relevance remains to be established, particularly for in vivo gene-editing strategies.
RNA-based therapies targeting lipid metabolism pathways have demonstrated clinical efficacy in related cardiometabolic conditions, supporting their translational relevance. RNA interference (RNAi), antisense oligonucleotides (ASOs), and small activating RNAs (saRNAs) provide additional strategies for modulating metabolic gene expression. RNAi therapies targeting enzymes involved in lipid metabolism (e.g., ANGPTL3, ApoC3) have shown potent triglyceride-lowering effects, suggesting potential benefits for metabolic dysfunction [28–30]. In diabetes models, siRNA-mediated silencing of negative regulators of insulin sensitivity—such as PTP1B—has improved glucose tolerance and insulin action.
Major challenges in gene therapy include off-target editing, immune responses to viral vectors or bacterial Cas proteins, vector biodistribution, and long-term safety. Integration-free editing, cell-type–specific promoters, self-limiting Cas9 expression systems, and transient delivery via mRNA are under development to enhance safety. Ethical considerations, particularly for heritable genome editing, require rigorous regulatory oversight. Manufacturing scalability, cost of goods, and access disparities also remain significant barriers as these therapies move toward clinical application.
In vivo gene-editing strategies raise important safety concerns that extend beyond delivery efficiency. Off-target DNA modifications remain a central risk, as unintended edits may disrupt tumor suppressor genes or activate oncogenic pathways [31, 32]. Mosaicism—where only a subset of target cells is successfully edited—can lead to heterogeneous tissue function and unpredictable metabolic outcomes, particularly in organs with limited regenerative capacity such as pancreatic islets [33]. In addition, long-term carcinogenic risk remains incompletely understood, especially in the context of permanent genomic alterations and chronic Cas nuclease expression [34].
Multiple technological advances aim to reduce these risks. High-fidelity Cas9 variants, paired nickases, and truncated guide RNAs have been developed to minimize off-target activity [35]. Base editing and prime editing platforms further reduce genomic risk by avoiding double-strand DNA breaks, thereby decreasing the likelihood of chromosomal rearrangements and p53 activation [22, 36]. Transient delivery approaches—such as mRNA-based editors or ribonucleoprotein complexes delivered via LNPs—limit the duration of nuclease exposure and may improve safety profiles in vivo [37].
Despite these advances, substantial uncertainties remain regarding the long-term genomic stability, immunogenicity, and oncogenic potential of edited cells following in vivo administration. Current human data are limited to small cohorts and short follow-up durations, precluding definitive conclusions regarding lifetime cancer risk or delayed adverse events. As such, rigorous long-term surveillance, post-marketing pharmacovigilance, and cautious patient selection will be essential as metabolic gene-editing strategies move toward broader clinical application.
Advanced drug delivery technologies are redefining therapeutic possibilities for diabetes by enabling targeted, sustained, or stimulus-responsive delivery of insulin and metabolic drugs. Nanotechnology, innovative biomaterials, and engineered biological vesicles provide platforms that can overcome limitations of traditional routes—particularly oral degradation, poor bioavailability, and lack of glucose responsiveness—thereby improving therapeutic precision and patient adherence. Figure 3 illustrates the major nano-enabled drug delivery platforms designed to enhance targeting, stability, and controlled release of metabolic therapies.
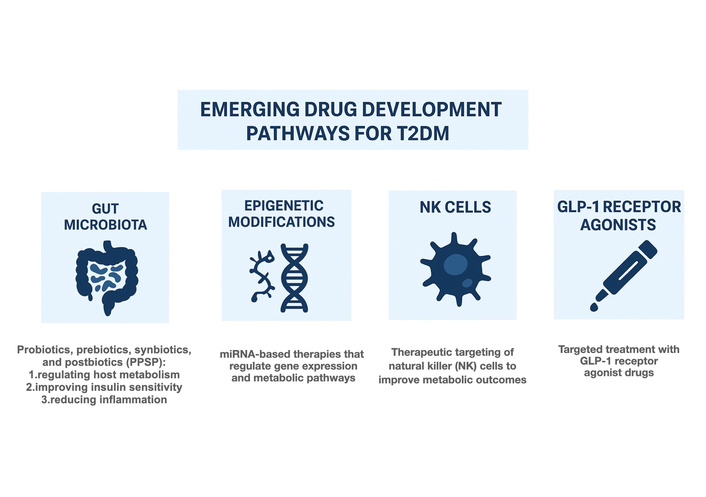
Emerging drug development pathways for type 2 diabetes mellitus. Emerging therapeutic pathways include modulation of the gut microbiota via probiotics, prebiotics, synbiotics, and postbiotics to regulate metabolism and inflammation; epigenetic modification strategies such as miRNA-based therapies to alter gene expression; immune-based approaches targeting natural killer (NK) cells to improve metabolic outcomes; and expansion of incretin-targeted therapies, including glucagon-like peptide-1 (GLP-1) receptor agonists. These pathways represent innovative mechanisms under investigation to enhance metabolic control and ameliorate IR. This figure summarizes predominantly preclinical and early translational concepts; human clinical relevance remains to be established.
Well-established clinical experience supports the feasibility of alternative insulin delivery routes, such as oral or inhaled formulations, in selected patient populations. Oral insulin delivery has long been hindered by enzymatic degradation and low intestinal permeability. Recent nanocarrier formulations—including polymeric nanoparticles, liposomes, and mucoadhesive carriers—protect insulin from gastrointestinal degradation and enhance trans-epithelial transport. Chitosan-based nanoparticles, for example, transiently open tight junctions to facilitate paracellular absorption, while enteric-coated lipid carriers resist gastric acid and release insulin in the small intestine [38]. Inhaled insulin and transdermal microneedle patches also offer noninvasive alternatives, with microneedles providing minimally painful, controllable release of insulin or GLP-1 analogues directly into dermal microcirculation [39–41].
Nanoparticle-based delivery systems have demonstrated reproducible pharmacokinetic advantages in multiple preclinical and translational studies. Nanotechnology enables organ- and cell-specific delivery of therapeutics to pancreatic islets, liver, muscle, and adipose tissue. Ligand-modified nanoparticles—such as those conjugated with GLP-1 peptides, mannose, or integrin-targeting motifs—can selectively home to β-cells, improving intrapancreatic drug accumulation [42, 43]. In the liver, LNPs engineered for hepatocyte uptake have shown success in silencing metabolic genes, reducing hepatic glucose output, and enhancing insulin sensitivity [44]. Precision nanocarriers designed for adipose targeting may improve delivery of insulin-sensitizing drugs, anti-inflammatory agents, or RNA therapeutics to inflamed or hypertrophic adipose depots.
Emerging preclinical data suggest that glucose-responsive and closed-loop delivery systems may improve glycemic control; however, clinical translation remains limited. Glucose-responsive delivery aims to mimic the self-regulating physiology of endogenous β-cells. Smart insulin delivery systems employ glucose oxidase, phenylboronic acid, or glucose-binding lectins to trigger drug release proportional to circulating glucose levels [45]. For example, microneedle patches containing glucose oxidase generate local pH changes in hyperglycemia that activate insulin release from polymeric vesicles. Other systems use polymer networks that swell or contract based on glucose concentration, enabling closed-loop, self-regulating insulin release without external pumps [46–48].
EVs, including exosomes derived from plants, milk, or mammalian cells, are emerging as biocompatible, low-immunogenicity carriers for peptides, RNA molecules, and small drugs. Plant-derived EVs have demonstrated stability in the gastrointestinal tract and can modulate gut microbiota composition, reduce inflammation, and improve glucose tolerance in preclinical studies [49]. Engineered mammalian exosomes loaded with therapeutic RNAs offer targeted approaches for metabolic regulation with enhanced intracellular delivery efficiency [46, 49, 50].
Advanced biomaterials enable sustained, controlled release of insulin or metabolic modulators. Injectable hydrogels—such as thermos-responsive or pH-sensitive formulations—can deliver insulin over hours to days while protecting it from enzymatic degradation [51]. Composite scaffolds combining polymers and nanoparticles can simultaneously release growth factors, anti-inflammatory molecules, and metabolic drugs to support β-cell survival, reduce islet stress, and improve graft function in regenerative therapies [45, 52]. These nano-enabled platforms remain largely experimental, with human efficacy and long-term safety yet to be confirmed.
Despite promising preclinical performance, nanomedicine for diabetes faces several translational challenges. Oral systems must achieve consistent bioavailability across varied human gastrointestinal conditions. Targeted nanoparticles require precise control of biodistribution, avoidance of off-target accumulation, and mitigation of immunogenicity. Long-term safety, cost-effective manufacturing, and regulatory harmonization remain major obstacles. Future directions include bioinspired nanocarriers, universal tissue-targeting ligands, AI-guided nanoparticle design, and hybrid systems integrating glucose responsiveness with real-time biosensing [53, 54].
Beyond biological efficacy, the clinical translation of nano-enabled drug delivery systems for diabetes is strongly influenced by manufacturing and regulatory considerations. Achieving batch-to-batch reproducibility remains a major challenge, as small variations in nanoparticle size, surface chemistry, or payload encapsulation can substantially alter pharmacokinetics, biodistribution, and immunogenicity [55, 56]. From a regulatory perspective, both the U.S. Food and Drug Administration (FDA) and the European Medicines Agency (EMA) require rigorous physicochemical characterization, validated manufacturing processes, and scalable quality-control frameworks for nanomedicines, yet harmonized standards across jurisdictions remain limited [57, 58]. In addition, long-term pharmacovigilance is essential, as delayed immune activation, bioaccumulation, or unforeseen toxicity may not be apparent in early-phase trials. Establishing post-marketing surveillance strategies and real-world evidence frameworks will therefore be critical to ensuring the safety, durability, and public trust of nano-enabled therapies as they move toward broader clinical use.
Innovations in nano-enabled delivery systems hold the potential to transform diabetes care by reducing therapy burden, improving metabolic control, and enabling disease-modifying interventions tailored to individual physiological responses.
Precision medicine is rapidly transforming the understanding and treatment of diabetes by identifying biologically distinct subtypes, predicting therapeutic response, and guiding individualized interventions. Multi-omics approaches—including genomics, transcriptomics, proteomics, metabolomics, lipidomics, and microbiome profiling—allow deep phenotyping of metabolic dysfunction and provide unprecedented insight into the heterogeneity of diabetes. Integration of these data with AI and machine learning (ML) methods has begun to reveal clinically actionable molecular signatures that can refine diagnosis, improve prognostication, and optimize therapeutic selection.
Well-established genomic studies have identified reproducible genetic variants associated with diabetes risk, enabling stratification of disease subtypes. Genome-wide association studies (GWAS) have identified hundreds of variants influencing glycemic traits, insulin secretion, and β-cell vulnerability [59]. Polygenic risk scores (PRS) derived from these variants can predict individual risk of T2D, stratify early-onset disease, and differentiate subtypes with distinct metabolic drivers. Clinical application of monogenic diabetes testing represents a validated example of precision medicine in routine care. Monogenic forms of diabetes, such as MODY, illustrate the power of genomics: patients with HNF1A-MODY respond exceptionally well to sulfonylureas, whereas those with GCK-MODY often require no pharmacologic treatment [20]. Incorporating genomic profiling into clinical decision-making may allow personalized therapeutic strategies based on inherited metabolic architecture [60, 61].
Transcriptomic profiling of pancreatic islets, adipose tissue, liver, and skeletal muscle has uncovered molecular networks associated with β-cell dedifferentiation, IR, and inflammation. Single-cell RNA sequencing (scRNA-seq) has revealed β-cell subpopulations with distinct stress signatures and maturation states, informing targeted therapies to enhance β-cell resilience [62]. Proteomics provides complementary insight by quantifying signaling pathways, post-translational modifications, and secreted factors that govern metabolic adaptation. Proteogenomic integration supports the identification of new biomarkers predictive of drug response or disease progression [63].
Metabolomics and lipidomics characterize alterations in circulating metabolites, lipids, and amino acids that reflect underlying pathophysiological processes. Branched-chain amino acids (BCAAs), acylcarnitines, ceramides, and phospholipids are consistently associated with IR and cardiometabolic risk [64]. Metabolic signatures can identify individuals with disproportionate lipotoxicity, impaired mitochondrial oxidation, or dysregulated glycolysis—pointing toward targeted lifestyle, pharmacologic, or nutraceutical interventions. These omics-based biomarkers may also predict responsiveness to GLP-1 RA, SGLT2 inhibitors, or emerging metabolic drugs [65, 66].
Gut microbiome composition strongly influences glucose metabolism, inflammatory tone, bile acid signaling, and insulin sensitivity. Dysbiosis, characterized by reduced short-chain fatty acid (SCFA)–producing species, increased LPS–producing bacteria, and compromised mucosal barrier function, contributes to metabolic dysfunction [67]. Microbiome sequencing has revealed distinct microbial signatures associated with glycemic response to dietary interventions, metformin, and personalized nutrition plans. Integration of microbiome data into precision medicine frameworks may guide individualized dietary strategies, probiotic therapy, or microbiome-modulating interventions [67–69].
Emerging evidence suggests that multi-omics integration and AI may enable more refined disease stratification. AI and ML methods enable integration of multi-omics datasets with clinical, imaging, and continuous glucose monitoring (CGM) data. These models identify patterns invisible to conventional analysis and can classify diabetes into biologically meaningful subgroups with prognostic relevance. ML-derived clusters have already been shown to predict risk of diabetes complications, β-cell decline, and treatment response [70–72]. AI-assisted platforms can also optimize insulin dosing, forecast glycemic excursions, and enhance closed-loop insulin delivery systems. These approaches remain largely investigational and face substantial barriers to widespread clinical implementation.
Despite their promise, widespread clinical implementation of multi-omics and AI-driven approaches faces substantial real-world barriers. High costs associated with large-scale sequencing, data storage, and computational infrastructure currently limit accessibility beyond specialized centers [73, 74]. Integration into routine care requires robust digital infrastructure, interoperability between electronic health records and omics platforms, and standardized analytic pipelines, which remain inconsistently implemented across healthcare systems [75, 76]. In addition, effective clinical adoption depends on clinician literacy in genomics, data science, and AI-assisted decision-making, as well as multidisciplinary collaboration, which are not yet uniformly embedded in medical training or clinical workflows [77]. These challenges are particularly pronounced in low- and middle-income countries, where resource constraints, limited bioinformatics infrastructure, and workforce shortages may exacerbate existing disparities in diabetes care [78]. Addressing these economic, infrastructural, and educational barriers will be essential to ensure equitable, ethical, and sustainable translation of precision-medicine approaches into global diabetes care.
Future directions include real-time precision nutrition, organ-specific multi-omics (e.g., islet proteogenomics), and cross-platform digital twins that simulate individual metabolic trajectories to optimize therapy.
Together, precision medicine and multi-omics integration hold the potential to fundamentally reshape diabetes care by enabling personalized diagnosis, targeted therapies, and predictive disease modeling grounded in deep biological insight.
Rapid advancements in regenerative medicine, gene editing, nanotechnology, and precision medicine are generating unprecedented opportunities to redefine the therapeutic landscape of diabetes. However, several scientific, regulatory, ethical, and logistical challenges must be addressed before these innovations can be widely adopted in clinical practice. A forward-looking appraisal of emerging technologies clarifies both the transformative potential and the critical barriers that remain.
Stem cell-derived β-cell replacement has shown promising early clinical outcomes, but further refinement is required to ensure durable engraftment, robust glucose responsiveness, and minimized immune rejection. Key priorities include improving β-cell maturation, reducing off-target endocrine cell populations, and optimizing implantation sites to enhance vascularization and survival [13]. Immune-evasive gene editing, such as deletion or modification of HLA molecules and overexpression of CD47, represents a promising strategy for eliminating the need for chronic immunosuppression, but long-term safety data are still lacking [17, 79].
Gene therapy approaches must address several translational barriers, including off-target editing, vector immunogenicity, genomic instability, and precise tissue targeting. While non-viral LNPs offer safer alternatives to viral vectors, achieving consistent and selective delivery to metabolic tissues such as islets, adipose, and muscle remains challenging [24]. Base and prime editors reduce double-strand break-associated risks, but controlling the duration and localization of editing activity is essential to minimize unintended consequences [22]. Regulatory oversight of genome-editing therapies, particularly those with heritable potential, will require harmonized global frameworks. For in vivo metabolic gene editing, clinical translation will ultimately depend on demonstrating a favorable long-term risk-benefit balance through extended follow-up, sensitive off-target detection methods, and robust regulatory oversight.
Nanoparticle-based insulin and drug delivery systems face hurdles related to manufacturing reproducibility, long-term biocompatibility, and interpatient variability in absorption. Oral and glucose-responsive formulations must reliably perform across a broad range of gastrointestinal environments, ages, and comorbidities [38]. Comprehensive long-term safety evaluations—including biodistribution studies, immunogenicity profiles, and elimination kinetics—are essential before broad clinical adoption. Moreover, cost-effective and scalable production processes must be developed to ensure equitable global access.
Although multi-omics and AI-driven stratification have demonstrated significant potential, integration into clinical workflows remains limited. Barriers include variability in omics platforms, lack of standardized reference datasets, limited data diversity, and concerns regarding data privacy and interoperability [70]. Developing federated learning models, harmonized analytic pipelines, and affordable large-scale sequencing strategies will be critical for bringing precision diabetes medicine into routine care. Future systems may incorporate real-time biosensing, digital health monitoring, and metabolic “digital twins” capable of simulating individualized disease trajectories.
As advanced therapies enter clinical testing, ethical considerations become increasingly prominent. Gene editing in germline cells remains prohibited in many jurisdictions due to concerns about long-term societal and biological impacts. Ensuring equitable access to regenerative and genetic therapies—many of which may initially have high costs—will be critical to prevent widening disparities in diabetes outcomes [80, 81]. Transparent regulatory pathways, community engagement, and global collaboration are needed to guide sustainable and ethically responsible implementation.
Despite these barriers, the convergence of stem cell technologies, precision gene editing, advanced drug delivery, and multi-omics-guided personalization is moving the field toward true disease-modifying therapies. Integrated platforms—such as gene-edited stem cell-derived β-cells within nano-protective biomaterial scaffolds or AI-informed targeted metabolic interventions—represent likely breakthroughs over the coming decade. Continued interdisciplinary collaboration and robust clinical evaluation will be essential for translating these innovations into therapies that can restore metabolic health, prevent complications, and ultimately transform diabetes management.
Despite the promising future, there are still safety concerns to be addressed.
Stem cell therapies often involve transient issues like backache, meningism, and mild infections, with severe events such as cerebrospinal fluid leaks or neuropathic pain occurring infrequently [82]. Gene editing via CRISPR carries risks of off-target mutations, large DNA rearrangements, and potential cancer promotion from genomic instability [83]. Encapsulation devices for islet cells trigger foreign body responses, leading to pericapsular fibrotic overgrowth that impairs nutrient diffusion and graft viability [84]. Glucose-responsive insulin systems may provoke hypoglycemia, hyperglycemia, or device failures like infusion errors, alongside neuropathy exacerbation in diabetics [85]. LNP platforms, key for mRNA delivery, elicit inflammatory responses via Toll-like receptor activation, protein corona formation disrupting transfection, and toxicity from ionizable lipids [86]. Long-term concerns across therapies include tumorigenesis, immune rejection, and thromboembolism, though clinical trials report mostly mild-to-moderate events resolvable without intervention.
Rigorous long-term follow-up is needed to assess durability, oncogenic risk, immune responses, and delayed adverse events associated with stem cell-based, gene-editing, and nano-enabled therapies.
Greater emphasis should be placed on validating preclinical findings in human-relevant models and well-designed early-phase clinical trials, with careful patient stratification for T1D versus advanced T2D.
Development and clinical validation of high-fidelity editing platforms, transient delivery systems, and sensitive off-target detection methods remain critical for safe in vivo applications.
Standardized, reproducible manufacturing processes and harmonized regulatory frameworks will be essential to enable scalable production and global deployment of advanced therapeutics.
Cost-effective multi-omics platforms, interoperable digital infrastructure, and clinician training programs are required to ensure equitable access, particularly in low- and middle-income countries.
Future studies should focus on embedding advanced therapies, AI tools, and decision-support systems into real-world clinical practice while demonstrating clear benefit over existing standards of care.
Diabetes and metabolic dysfunction continue to impose a profound global burden, driven by complex interactions among genetic, environmental, metabolic, and immunological factors. While current therapies effectively manage hyperglycemia, they do not sufficiently address the underlying drivers of β-cell dysfunction, IR, inflammation, or metabolic derangements that contribute to long-term complications. Advances in regenerative medicine, gene editing, nano-enabled drug delivery, and precision medicine have ushered in a new era of innovative strategies that hold promise for modifying disease progression rather than merely controlling symptoms.
Importantly, the therapeutic relevance of the strategies discussed varies across diabetes subtypes, with β-cell replacement and immune-evasive therapies primarily targeting T1D and late-stage T2D, whereas gene modulation, drug delivery innovations, and precision medicine approaches have broader applicability across the metabolic disease spectrum.
Stem cell-derived β-cell replacement, supported by breakthroughs in differentiation protocols, encapsulation technologies, and immune-evasive gene editing, is progressing from experimental concepts to early clinical translation, with initial data demonstrating restoration of endogenous insulin production [13, 17, 79]. Gene therapy approaches—including CRISPR-based editing, base and prime editing, and RNA-based therapeutics—have shown potential to correct monogenic defects, enhance β-cell resilience, and remodel metabolic pathways [22]. Concurrently, nano-enabled and biologically inspired drug delivery systems are improving therapeutic precision by enabling targeted, noninvasive, and glucose-responsive delivery of insulin and metabolic agents [38]. Multi-omics and AI-driven computational models are redefining diabetes heterogeneity, providing tools for individualized risk prediction, stratified therapies, and personalized interventions [70]. Ensuring equitable global implementation of multi-omics and AI-driven precision medicine will require parallel investments in infrastructure, workforce training, and cost-reduction strategies, particularly in low- and middle-income settings.
Successful integration of nano-enabled therapeutics into diabetes care will depend not only on biological innovation, but also on reproducible manufacturing, regulatory alignment, and robust long-term pharmacovigilance
Despite this progress, significant translational challenges remain, including ensuring long-term safety, achieving consistent delivery to metabolic tissues, reducing manufacturing barriers, enhancing regulatory clarity, and ensuring global accessibility. Ethical considerations, especially those related to gene editing and high-cost regenerative therapies, must be carefully navigated to ensure equity in implementation.
Nevertheless, the convergence of these emerging technologies signals a transformative shift toward disease-modifying and personalized metabolic restoration therapies. Continued interdisciplinary research, robust clinical trials, and strategic integration of omics-guided precision medicine will be critical for realizing the full potential of these innovations. As scientific and technological progress accelerates, the next decade may bring therapies capable not only of improving glycemic control but of fundamentally restoring metabolic health and preventing the long-term complications of diabetes.
While several strategies discussed in this review are supported by early clinical evidence, many remain in preclinical or early translational stages. Continued rigorous clinical evaluation will be essential to determine their long-term safety, efficacy, and real-world applicability.
AAV: adeno-associated virus
AI: artificial intelligence
EVs: extracellular vesicles
GLP-1RA: glucagon-like peptide-1 receptor agonists
IR: insulin resistance
LNPs: lipid nanoparticles
LPSs: lipopolysaccharides
MASLD: metabolic dysfunction-associated steatotic liver disease
ML: machine learning
MODY: maturity-onset diabetes of the young
RNAi: RNA interference
ROS: reactive oxygen species
SGLT2i: sodium-glucose cotransporter 2 inhibitors
T1D: type 1 diabetes
T2D: type 2 diabetes
MKS, CF, and IB: Conceptualization, Investigation, Writing—original draft, Writing—review & editing. KM: Conceptualization, Investigation, Conceptualization. IG, SPL, KD, KA, NK, AT, CC, and KT: Formal analysis, Investigation, Writing—review & editing. All authors read and approved the submitted version.
The authors declare that there are no conflicts of interest.
Not applicable.
Not applicable.
Not applicable.
Not applicable.
Not applicable.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
View: 2306
Download: 50
Times Cited: 0
Huiling Huang ... Dongsheng Li
Huiling Huang ... Dongsheng Li
Dorothy E. Vatner ... Stephen F. Vatner
