Affiliation:
1Department of Clinical Biochemistry and Pharmacology, Ben-Gurion University of the Negev, Be’er Sheva 8410501, Israel
2Laboratory for Biopharmaceutics, E.D. Bergmann Campus, Ben-Gurion University of the Negev, Be’er Sheva 8410501, Israel
ORCID: https://orcid.org/0000-0001-8654-6511
Affiliation:
2Laboratory for Biopharmaceutics, E.D. Bergmann Campus, Ben-Gurion University of the Negev, Be’er Sheva 8410501, Israel
ORCID: https://orcid.org/0009-0002-6771-350X
Affiliation:
1Department of Clinical Biochemistry and Pharmacology, Ben-Gurion University of the Negev, Be’er Sheva 8410501, Israel
Email: sbs@bgu.ac.il
ORCID: https://orcid.org/0000-0001-7551-5488
Affiliation:
2Laboratory for Biopharmaceutics, E.D. Bergmann Campus, Ben-Gurion University of the Negev, Be’er Sheva 8410501, Israel
3Department of Biomedical Engineering, Ben-Gurion University of the Negev, Be’er Sheva 8410501, Israel
Email: asintov@bgu.ac.il
ORCID: https://orcid.org/0000-0001-7995-0276
Explor Drug Sci. 2026;4:1008157 DOI: https://doi.org/10.37349/eds.2026.1008157
Received: January 08, 2026 Accepted: February 26, 2026 Published: April 15, 2026
Academic Editor: Xiqun Jiang, Nanjing University, China
Aim: Chitosan (CHS)-based nanoparticulate systems have gained much interest due to their high drug loading capacity and the simplicity of their fabrication. The physical properties of two types of curcumin-loaded CHS nanoparticles (CHS-NPs) were determined and compared. A new in-vitro release method was developed based on mathematical modeling in which the drug is first released from the NP into the surrounding medium and subsequently diffuses through the membrane.
Methods: Curcumin-loaded CHS-NPs were fabricated by ionotropic gelation using sodium tripolyphosphate (TPP) and sodium hexametaphosphate (SHMP) crosslinking, and characterized by NP tracking analysis, loading capacity, zeta potential, Fourier transform infrared spectroscopy (FTIR), and in-vitro release rates.
Results: The data showed that compared to SHMP crosslinked CHS-NPs, TPP crosslinking demonstrated a decrease in entrapment efficiency at a relatively high concentration of the agent, probably by narrowing the space between the polymeric chains. As indicated by zeta potential measurements, TPP crosslinking at all levels was more uniformly distributed inside the NPs, whereas the higher molecular weight SHMP at a low concentration creates NPs mostly by binding onto the surface. It was found that the release rates of curcumin from CHS-NPs crosslinked by SHMP at higher concentrations were about twice as high as the release rates of curcumin from TPP-crosslinked CHS-NPs, accompanied by notable lag times.
Conclusions: This significant increase in the release rates of curcumin from SHMP-crosslinked CHS-NPs is explained by the large spatial structure of this crosslinker compared to the small TPP molecules. This study advances the literature on drug diffusion by making it possible to accurately determine its release from nanoparticulate systems.
Nanoparticles (NPs) based on natural polysaccharides have been intensely studied due to their biocompatibility, biodegradability, non-immunogenicity, and environmentally benign nature. Besides its general properties as a natural polysaccharide, chitosan (CHS), a linear cationic polysaccharide, displays many attractive properties. To name a few, it has been shown to be mucoadhesive [1–3] and is known to have wound healing activities [4–6]. CHS has gained a widespread interest in the last couple of decades as an adequate nanoparticulate carrier [4, 7–9], as its NPs have demonstrated enhanced delivery and biodistribution of drugs by opening tight junctions [10, 11]. In addition, CHS-NPs have shown increased cellular uptake by cationic ligands with protein domains on the cell surface [12].
There are various methods that have been utilized for manufacturing CHS-NPs. They are basically classified into ‘coacervation’ and ‘emulsification’ methods [13, 14]. Coacervation methods include ionotropic gelation, polyelectrolyte complexation, and desolvation [15], while emulsification methods include emulsion droplet coalescence, emulsion solvent diffusion, and reverse micellization. CHS-NPs loaded with various drugs were demonstrated to enhance drug activity and improve cellular uptake of therapeutics [16–18]. The most popular and commonly preferred method of CHS-NP preparation is the coacervation method based on ionotropic gelation and ionotropic gelation/polyelectrolyte complexation [19], mainly due to its simplicity and because it does not involve addition of an organic solvent. Ionotropic gelation of CHS was first described by Calvo et al. [20], who used sodium tripolyphosphate (TPP) as the polyanionic cross-linker. TPP has frequently been used as the negatively charged polyanion, however, sodium hexametaphosphate (SHMP) has also been reported, although less frequently, as ionotropic crosslinker agent for the preparation of CHS-NPs [21–23]. In the present paper, we used curcumin (CUR) as the model agent. CUR is a natural polyphenolic compound possessing many beneficial properties, including anticancer, antioxidant, antimicrobial, anti-inflammatory, and neuroprotective activities [24–29]. CHS-NPs loaded with CUR were prepared using various methods [30–33]. CUR-containing CHS-NPs (combined with other antioxidants) have been prepared using SHMP [34], but the overall literature on this nano-system is lacking. A comparison of CUR-containing CHS-NPs prepared using the two different commonly-used cross-linkers has not previously been studied and reported. The aim of the current study was to explore the properties of CUR-loaded CHS-NPs fabricated using SHMP crosslinker and to compare them with particles prepared by the more commonly used TPP crosslinker. It was hypothesized that due to the structural differences between the two crosslinking molecules, TPP and SHMP, NPs with different characteristics would be formed. Hence, the physical properties of CUR-loaded CHS-NPs, which were prepared by TPP and SHMP, were analyzed, determined, and compared. Furthermore, the release rates of CUR from the various NPs were determined using the Franz cell system as previously published by our group [35, 36], providing an improved precision.
CHS [low molecular weight: 50–190 kDa, degree of deacetylation ≥ 75%, viscosity: 20–300 cP, 1% (w/w) in 1% acetic acid], SHMP, Tween 80 and tablets for phosphate buffered saline (PBS) preparation were obtained from Sigma Aldrich (Sigma-Aldrich Israel Ltd., Rehovot, Israel). TPP from Santa Cruz Biotechnology Inc. (Dallas, TX, USA). CUR was obtained from Glentham Life Sciences Ltd. (Corsham, United Kingdom). Acetic acid glacial was purchased from Merck (Darmstadt, Germany). Ethyl alcohol AR was purchased from BioLab Ltd. (Jerusalem, Israel). Methanol and high-performance liquid chromatography (HPLC) grade solvents were obtained from J.T. Baker (Mallinckrodt Baker, Inc., Phillipsburg, NJ).
CUR-loaded CHS-NPs were prepared using the ionic gelation method [37–41]. CHS (12 mg) was dissolved by consecutive addition of acetic acid glacial (150 µL) and double-distilled water (DDW) (40 mL). This mixture was stirred using a magnetic bar at 700 rpm for 10 min, and the pH was measured (pH = 3.0–3.2). NaOH 1 N solution was then added at 0.2 mL/min by a syringe pump (NE-300, New Era Pump Systems, Farmingdale, NY, USA) with continuous pH monitoring until pH = 5. Separately, CUR was dissolved in ethanol (5 mg/2.5 mL) and sonicated for 480 s. Tween 80 (90 µL volume) was added, and the mixture was vortexed. Even though CUR had been dissolved in ethanol [42, 43], Tween 80 was added to avoid CUR precipitation prior to the formation of NPs in the aqueous environment. The use of Tween 80 was previously described in the context of ionic gelation method [44]. Various TPP or SHMP quantities (Table 1) were dissolved in DDW and added to the CUR solution and the total volume was made up to 4 mL for all combined components. This mixture dripped into the CHS solution using a syringe pump at 0.1 mL/min for TPP or 0.05 mL/min for SHMP with under stirring (500 rpm) (Figure 1). The lower dripping rate of SHMP was required since larger particles were formed at higher rates. The mixture of the combined solutions was further stirred for 1 h at 400 rpm. This step was performed at a slightly slower stirring rate to allow complete cross-linking and particle stabilization, while minimizing aggregation.
A summary of the prepared CHS nanoparticulate formulations.
| Formulation | Cross-linker | CUR added (mg CUR/mg CHS) | Polyanion added (mg/mg CHS) | Polyanion level (mmol/mg CHS) |
|---|---|---|---|---|
| CHS-T1 | TPP (sodium salt) | 0.417 | 0.025 | 0.068 |
| CHS-T2 | 0.050 | 0.136 | ||
| CHS-T3 | 0.083 | 0.226 | ||
| CHS-T4 | 0.125 | 0.340 | ||
| CHS-S1 | SHMP | 0.417 | 0.042 | 0.068 |
| CHS-S2 | 0.083 | 0.136 | ||
| CHS-S3 | 0.139 | 0.226 | ||
| CHS-S4 | 0.208 | 0.340 |
CHS: chitosan; CUR: curcumin; SHMP: sodium hexametaphosphate; TPP: sodium tripolyphosphate.
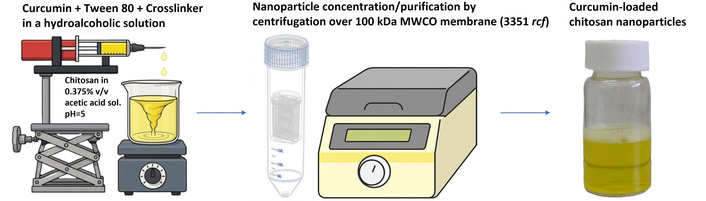

Schematic representation of batch process manufacturing of curcumin-loaded chitosan nanoparticles.
The NPs dispersions were filtered using ultra centrifugal filter with 100 kDa MWCO membrane (Amicon Ultra from Merck Millipore Ltd., Cork, Ireland) at 3,351 rcf, 30 min, 25°C (Figure 1). The liquid that was obtained after filtration was collected, diluted ×10 with methanol and analyzed by HPLC. This liquid that crossed the membrane represented the free unloaded CUR molecules. The samples were injected into a HPLC system (1260 Infinity II, Agilent Technologies Inc., Santa Clara, CA), equipped with a prepacked column (BetaSil C18, 5 μm, 250 mm × 4.6 mm, Thermo Fisher Scientific). The samples were chromatographed using a mobile phase consisting of acetonitrile/acetic acid 0.2% solution (75:25) at a flow rate of 1 mL/min. Calibration curves, peak areas measured at 425 nm for CUR versus drug concentration, were constructed by running standard drug solutions in methanol for each series of chromatographed samples. The limit of detection (LOD) was 0.1 µg/mL and the limit of quantification (LOQ) was 0.25 µg/mL. The mass of CUR in the NPs was calculated by subtracting the obtained data of unloaded CUR from the total mass of CUR added in the preparation process.
Encapsulation (or entrapment) efficiency percentage (EE%) was calculated according to the following equation:
Measurements were performed using a NanoSight NS300 instrument (Malvern Instruments Ltd., Worcestershire, UK), equipped with a 642 nm red laser module and 450 nm long-pass filter, and a camera operating at 25 frames per second, capturing a video file of the particles moving at a constant flow rate from a syringe on a syringe pump. The software for capturing and analyzing the data (NTA 3.4) calculated the hydrodynamic diameters of the particles by using the Stokes-Einstein equation. Depending on the NP concentration, each measurement provided data about size distribution (in diameter) for approximately 5,000–15,000 particles.
The samples were diluted to appropriate concentration using DDW and transferred to U-tube cuvette (DTS1060C, Malvern). Measurement was performed using Zeta sizer (ZN-NanoSizer, Malvern, UK). The particles were measured in automatic mode at 25°C.
TEM images were recorded with Talos F200C (Thermo Fisher Scientific) transmission electron microscope operating at 200 kV. Samples of CHS-NPs dispersions were deposited on a copper grid, coated with Formvar and carbon (Electron Microscopy Sciences, Fort Washington, PA, USA) and left to stand for 1 min, then the excess liquid was adsorbed using filter paper. The images were taken with Ceta 16M CMOS camera.
CHS-NPs and their individual components (CHS, CUR, TPP and SHMP) were analyzed using Nicolett 6700 FTIR spectrometer (Thermo Fisher Scientific). Samples of CHS-NPs were lyophilized for 24 h prior to examination. Infrared (IR) spectra of the various samples were recorded over a range of 4,000–500 cm–1.
The permeability of CUR through a hydrophobic polytetrafluoroethylene (PTFE) membrane (Starlab Scientific, 0.1 µm pore size, 25 mm diameter) was measured using a Franz diffusion cell system (Permegear, Inc., Bethlehem, PA, USA). The diffusion area was 1.767 cm2 (15 mm diameter orifice), and the receiver compartment volumes were 12 mL. The solution in the receiver compartment (“receiver solution”) consisted of PBS (0.01 M, pH 7.4) containing 10% (v/v) Tween 80, 0.01% (w/v) butylated hydroxytoluene (BHT), and 5% (v/v) methanol. CUR was dissolved in dimethyl sulfoxide (DMSO) (11 mg/mL), further diluted with the receiver solution and aliquoted to a volume of 0.58 mL containing 120 µg of CUR (DMSO final concentration = 1.9%). The CUR solution was placed in the donor compartment. The solution on the receiver side was stirred by externally driven, Teflon-coated magnetic bars and maintained at 37°C. At predetermined time points, samples of 1 mL from the receiver solution were withdrawn and kept for HPLC analysis. After every sampling, 1 mL of fresh solution was replenished to the volume. Each set of experiments was performed in triplicate (n = 3). A perfect sink condition in the receiver compartment was maintained throughout the permeation and the release experiments by using a relatively large volume of a receiver solution containing alcohol and a surfactant as described above. These conditions and the constant dilution ensured CUR solubility far in excess of the maximum theoretical receiver concentration of CUR.
Aliquots (0.5–0.6 mL) of CHS-NP dispersions were placed in the donor compartment. The two compartments (donor and receiver) were separated by a membrane permeable to CUR but impermeable to the NPs (PTFE membrane, 0.1 µm pore size, 25 mm diameter). The NPs were placed in the donor compartment, and the released CUR permeated through the membrane to the receiver compartment. Various CHS-NP formulations, each containing 120 µg of CUR in 0.5–0.6 mL volumes, were placed in the donor compartment. The experiments with the NPs were performed as mentioned above, except they were performed for 66 h instead of 48 h. Each set of experiments was performed in triplicate (n = 3).
Because of the sampling of large volumes from the receiver solution (and their replacement with equal volumes of buffer), the receiver solution was constantly being diluted. Taking this process into account, the cumulative drug that permeated out into the receiver [
Where
The release of CUR from the NPs was performed in a Franz diffusion cell system using a synthetic membrane. The permeability coefficient of the membrane was calculated by linear regression of the experimental data of cumulative permeation of the free drug in plain solutions (P) as previously described by Sintov [36]. Drug transport across the membrane is governed by the integral form of Fick’s law:
where,
where,
and after simplifying:
Denoting:
then,
whose solution is:
Noting that this leads to:
or:
and since,
Hence:
Thus, a plot of
The release rate of the drug from NPs can be calculated according to Gupta et al. [45]. Assuming the release of CUR from the NPs follows a zero-order process, then:
where,
After integration:
and rearrangement:
Since
after dividing by
At t = 0, the NPs were suspended in buffer and immediately applied into the donor compartment. For this case, an initial free drug
Combining Equation 6 with Equation 25 gives:
Equation 26 is solved by Laplace transforms and its expression is simplified by defining a as:
A plot of
The statistical differences between the release data of CUR obtained from the various CHS-NP formulations were analyzed employing the two-way unweighted means analysis of variance (ANOVA) test and Student’s t-test for multiple comparisons. The differences among groups were considered significant when p values < 0.05.
CHS-NPs were prepared by ionotropic gelation using two polyphosphate crosslinkers, TPP and SHMP, at 0.068, 0.136, 0.226, and 0.340 mmol/mg of CHS. Table 2 summarizes the particle size, particle size distribution [polydispersity index (PDI)], particle concentration, and ζ potential as well as CUR loading and EE of the various nanoparticulate systems. The systems were prepared with four different polyanion levels, i.e., four TPP levels and four SHMP levels. Figure 2 shows the morphological shapes of representative NPs, a TPP-crosslinked CHS-NP and a SHMP-crosslinked CHS-NP. Both NPs had spherical or elliptical shapes ranging in size between 200 to 300 nm. Using NTA, the mean particle size was within the range of 226–284 nm and 264–317 nm for TPP-crosslinked NPs and SHMP-crosslinked NPs, respectively.
A summary of curcumin (CUR) encapsulation (or entrapment) efficiency percentage (EE%), particle size, distribution [polydispersity index (PDI)], particle concentration, and zeta (ζ) potential of CUR-loaded chitosan nanoparticle (CHS-NP) formulations.
| Formulation | Sodium tripolyphosphate (TPP)-crosslinked CHS-NPs | Sodium hexametaphosphate (SHMP)-crosslinked CHS-NPs | ||||||
|---|---|---|---|---|---|---|---|---|
| T-1 | T-2 | T-3 | T-4 | S-1 | S-2 | S-3 | S-4 | |
| Cross-linker level (mmol/mg CHS) | 0.068 | 0.136 | 0.226 | 0.340 | 0.068 | 0.136 | 0.226 | 0.340 |
| CUR loading (mg/mg NPs) | 0.31 | 0.32 | 0.38 | 0.25 | 0.32 | 0.33 | 0.31 | 0.32 |
| EE% | 74 | 77 | 91 | 59 | 78 | 79 | 74 | 78 |
| Particle size (nm) (mean ± SD) | 283.6 ± 3.3 | 284.2 ± 21.0 | 226.6 ± 71.6 | 242.4 ± 21.2 | 264.2 ± 4.4 | 311.5 ± 15.2 | 276.3 ± 3.0 | 317.1 ± 1.3 |
| PDI | 0.28 | 0.24 | 0.14 | 0.16 | 0.2 | 0.1 | 0.13 | 0.1 |
| D10 (nm) | 137.3 | 149.7 | 152.5 | 140.7 | 144.8 | 203 | 167 | 204 |
| D50 (nm) | 231.6 | 249.2 | 237.3 | 225.6 | 241.7 | 292.5 | 261.5 | 301.4 |
| D90 (nm) | 507.5 | 475 | 372.2 | 364.6 | 399.1 | 445.1 | 409.5 | 449.5 |
| NP concentration (NPs/mL) (mean ± SD) | 2.0 × 1010 ± 1.08 × 109 | 4.02 × 1010 ± 2.14 × 109 | 3.11 × 1010 ± 1.91 × 109 | 5.07 × 1010 ± 2.37 × 109 | 2.81 × 1010 ± 2.02 × 109 | 9.27 × 1010 ± 4.41 × 109 | 2.05 × 1011 ± 5.67 × 109 | 1.04 × 1011 ± 2.89 × 109 |
| ζ potential (mV)-CUR-loaded NPs (mean ± SD) | 34.7 ± 0.7 | 31.1 ± 1.7 | 34.4 ± 3.6 | 32.2 ± 3.3 | 24.8 ± 1.35 | 30.8 ± 1.7 | 29.6 ± 2.3 | 29.0 ± 0.4 |
| ζ potential (mV)-unloaded (blank) NPs (mean ± SD) | 26.8 ± 3.4 | 26.3 ± 5.0 | 23.6 ± 2.7 | 32.1 ± 5.2 | 15.6 ± 1.6 | 23.1 ± 3.0 | 25.0 ± 2.2 | 34.6 ± 1.9 |

Transmission electron microscopy (TEM) images of chitosan nanoparticle (CHS-NP) formulations crosslinked with sodium tripolyphosphate (TPP) (A) and sodium hexametaphosphate (SHMP) (B) both at 0.340 mmol polyphosphate/mg of CHS. Images taken at ×45,000 magnification.
As shown in Table 2, CUR loading and the EE, were not dependent on the SHMP concentrations within the tested range, whereas approximately 0.3 mg/mg and 74–79% were determined for CUR loading and EE, respectively. Unlike SHMP crosslinking, CHS-NPs crosslinked by TPP at a concentration of 0.340 mmol/mg CHS demonstrated a significant decrease in CUR loading and in the calculated EE, 0.25 mg/mg and 59%, respectively.
In addition to size and loading parameters, surface charge characteristics were evaluated in order to assess the influence of crosslinker type and CUR incorporation on NP interfacial properties. The ζ potential data obtained for the various CHS-NPs (Table 2) showed the following phenomena: (a) CUR loading significantly increased the ζ potential of NPs crosslinked by TPP or SHMP at concentrations of 0.068, 0.136, and 0.226 mmol/mg polymer; (b) TPP-crosslinked NPs demonstrated a constant ζ potential (an average of 33.1 ± 1.7 mV) when CUR had been added at all crosslinker’s levels tested. In the absence of CUR, a constant ζ potential existed only at crosslinker’s levels of 0.068 to 0.226 mmol/mg polymer (an average of 25.6 ± 1.7 mV), while at 0.340 mmol TPP/mg CHS, the ζ potential was similar in loaded and unloaded NPs (~32 mV); (c) SHMP-crosslinked NPs demonstrated a constant ζ potential when CUR had been added at crosslinker’s levels of 0.136 mmol/mg CHS and above (an average of 29.8 ± 0.9 mV). However, at 0.068 mmol SHMP/mg CHS, ζ potential was lower than the values obtained at higher crosslinker’s levels; (d) In the absence of CUR, ζ potential values of SHMP-crosslinked CHS-NPs were directly dependent on the level of the crosslinker, increasing from 15.6 ± 1.6 mV to 34.6 ± 1.9 mV at SHMP levels of 0.068 mmol/mg up to 0.340 mmol/mg, respectively.
Figure 3 shows the FTIR spectra of two TPP-crosslinked CHS-NPs (0.136 and 0.340 mmol TPP/mg polymer; CHS-T2, T4), and two SHMP-crosslinked CHS-NPs (0.136 and 0.340 mmol SHMP/mg polymer; CHS-S2, S4). As shown in the figure, the spectra of these samples are similar with all their characteristic peaks: N–H and/or O–H stretching vibrations (3,362 cm–1 or 3,373 cm–1; wide bands), C–H stretching vibrations (2,924 cm–1 and 2,857 cm–1), carbonyl stretching vibrations (1,653 cm–1; very small peak), stretching vibrations of amide (1,570 cm–1), and C–O or P–O stretching vibrations (1,099 cm–1).
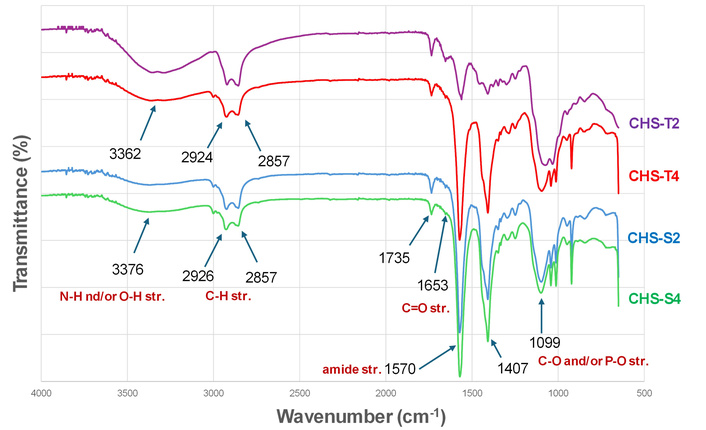
Infrared (IR) spectra of chitosan nanoparticle (CHS-NP) formulations crosslinked with 0.136 and 0.340 mmol polyphosphate [sodium tripolyphosphate (TPP) or sodium hexametaphosphate (SHMP)]/mg of CHS.
The FTIR spectra also indicated that TPP-crosslinked NPs, particularly the CHS-T2 sample, exhibited higher intensity of C–H, N–H, and O–H vibrations (peaks at 3,200–2,900 cm–1) compared with the spectra obtained from the SHMP-crosslinked NPs.
Another spectral feature was observed at 1,570 cm–1, 1,407 cm–1, in which the vibrational intensity is similar for the formulations CHS-T4, CHS-S2, and CHS-S4, and significantly lower (also with a higher transmittance) for the CHS-T2 sample. Spectral data of the raw materials are presented in Figure S1.
Figure 4 presents the cumulative permeation profiles of CUR through the membrane after loaded NPs and plain CUR solution were placed in the donor chamber. Naturally, free CUR in a plain solution permeated through the membrane relatively faster than the entrapped CUR in the CHS-NPs. Figure 5 shows that by plotting
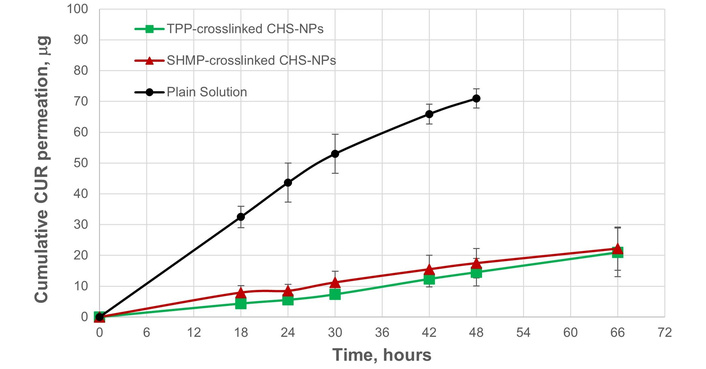
Curcumin (CUR) permeation (expressed as a cumulative quantity in the receiver chamber) through a polytetrafluoroethylene (PTFE) membrane. Free CUR in plain solution was compared with sodium tripolyphosphate (TPP)-crosslinked and sodium hexametaphosphate (SHMP)-crosslinked chitosan (CHS)-based nanoparticles (CHS-T1 and CHS-S1; 0.068 mmol polyanion/mg CHS). CHS-NPs: chitosan nanoparticles.
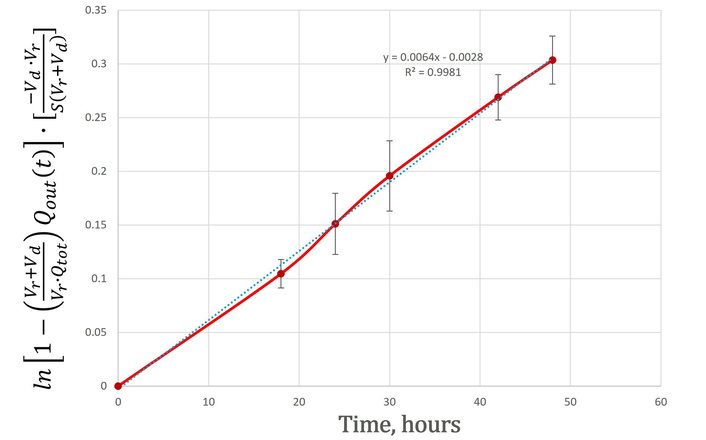
Determination of the permeability coefficient (P) of polytetrafluoroethylene (PTFE) membrane (0.1 µm pore size, 25 mm diameter) used in the release study of curcumin. P was derived from the slope of
As shown in Figure 4, linear permeation rates of CUR were measured through the membrane after its release from TPP- and SHMP-crosslinked CHS-NPs. These linear permeation profiles are the result of the release rates of the entrapped CUR from the NPs, which limit and regulate the permeation fluxes of the free molecules after their release. The release rates
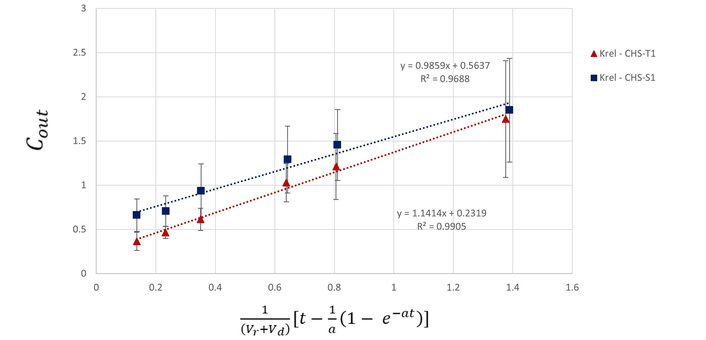
Calculation of the release rate constants of curcumin (CUR)
Figure 7 demonstrates that SHMP levels in the range of 0.136–0.340 mmol/mg CHS resulted in 1.6–1.9-fold increase of the steady state permeation fluxes (0.57–0.66 μg/h) compared to 0.068 mmol SHMP/mg CHS (0.35 μg/h; Figure 4). More importantly, the calculated release rate constants (
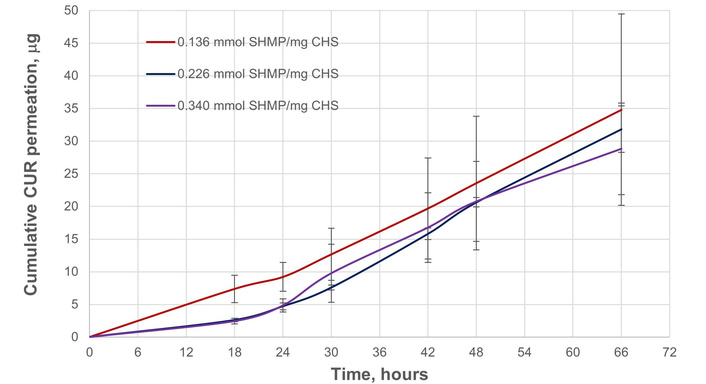
Curcumin (CUR) permeation (expressed as a cumulative quantity in the receiver chamber) through a polytetrafluoroethylene (PTFE) membrane. The permeation of free CUR molecules only through the membrane after they are released from sodium hexametaphosphate (SHMP)-crosslinked chitosan (CHS)-based nanoparticles (CHS-S2–S4; 0.136, 0.226, and 0.340 mmol polyanion/mg CHS).
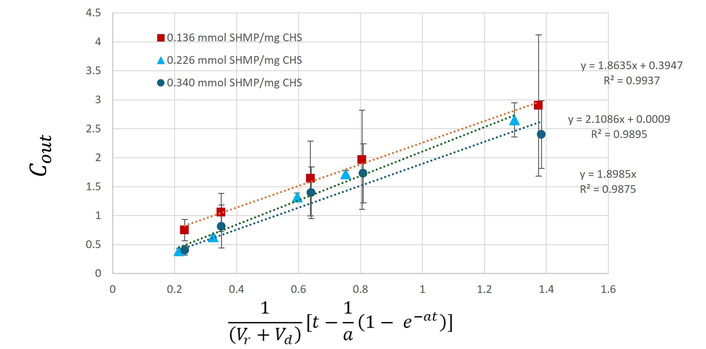
Calculation of the release rate constants of curcumin
The slight increase in the size of SHMP-crosslinked NPs compared to the TPP-crosslinked NPs (Table 2) was probably due to the larger SHMP molecules, which increased the space between CHS linear chains. TPP at a concentration of 0.340 mmol/mg CHS demonstrated a significant decrease in CUR loading and in the calculated EE. As this deviation had been repeatedly confirmed, it was postulated that at the high concentration of TPP, having a smaller molecular size (relative to SHMP), the NPs turned out to be denser and less permeable to CUR molecules. In addition, this finding may also be related to CUR’s poor water solubility. When a water-soluble active agent is used, it usually dissolves in the CHS solution, then the crosslinker is added dropwise [21, 46]. Here, CUR was added concomitantly with the crosslinker, which may cause CUR to compete with TPP resulting in less available CUR during NPs’ formation. Thus, at higher TPP concentrations, denser particles are formed faster, turning out to be less permeable to CUR.
As ζ potential is defined as the electrical potential at the interfacial surface of the particles, the decrease in the potential can be explained by the binding of the polyanion molecules on the surface of the unloaded CHS-NPs, thus neutralizing in part the positively charged surface. The increase in the potential noted in CUR-loaded CHS-NPs, can be explained therefore by the attraction of CUR molecules on the surface resulted in protection of the surficial ammonium cations by preventing interaction with the polyanion. This mechanism preserves the original ζ potential of the non-crosslinked CHS. The difference between the two polyphosphate crosslinkers, which is expressed in NPs at the low concentration of 0.068 mmol polyanion/mg CHS, is probably due to the larger molecular structure of SHMP compared to the TPP molecule. At low concentration, SHMP molecules create NPs mostly by binding onto the outer amine groups rather than into their inner core. When higher SHMP concentrations are used, the crosslinking turns out to be more uniformly distributed during the NP formation, and the ionic bonds are evenly created along all the polymer chains.
FTIR analysis provided further insight into these structural differences. Two stretching peaks at 1,735 cm–1 and 1,407 cm–1 did not appear in CHS spectra or in the IR spectra related to pure polyphosphates. These stretching peaks, therefore, seem to be associated with acetate anion stretching and the ionic interactions of the positively charged amine groups on the CHS’s backbones and the negatively charged acetate groups [47]. Bhumkar and Pokharkar [48] noted that in the FTIR spectra of TPP-crosslinked CHS (freeze-dried particles) the band of amide I at 1,655 cm–1 disappeared and two new peaks at 1,645 cm–1 and 1,554 cm–1 appeared, which they attributed to the linkage between the phosphoric and ammonium ions; however, such features were not observed in the present study. The higher intensity of C–H, N–H, and O–H vibrations observed in TPP-crosslinked NPs, particularly CHS-T2, likely reflects closer structural proximity between CHS chains, while reduced vibrational intensity at low TPP concentration is attributable to fewer ionic interactions.
The release profiles of drugs and active agents entrapped in NPs in general and in CUR-loaded CHS-NPs in particular [43] have been usually determined using the conventional dialysis bag method [15, 49]. However, since any method involving a membrane is dependent on its permeability coefficient for the tested drug, a technique allowing improved precision of the permeability coefficient measurement may be advantageous over the commonly used dialysis tubing. The use of Franz diffusion cell system for evaluating drug release from NPs can provide adequate measures of the membrane’s surface area as well as the volumes of the donor and the receiver solutions [35, 36].
As CUR release (and membrane permeation) from TPP-crosslinked CHS-NPs was not dependent on TPP concentrations in the given range, the effect of SHMP levels was also examined. The findings were quite different from TPP crosslinking, as the cumulative permeation profiles and the release rates of CUR were relatively higher.
This interesting finding complemented and supported our previous assessment (see above) that at low levels of SHMP, the crosslinking bonds concentrate mainly in the outer shell of the NPs, thus hindering and delaying the release of the entrapped CUR. At higher concentrations of SHMP, the crosslinking bonds are more uniformly distributed along the polymeric chains, enabling fluent diffusion of CUR outwards after a relatively short lag time. Naturally, a lag time appears as more crosslinking bonds are formed, which prolong the time to reach a steady state release rate. This aligns well with the ζ potential data (Table 2), where SHMP potential increased with concentration from 15.6 ± 1.6 mV to 34.6 ± 1.9 mV at SHMP levels of 0.068 mmol/mg up to 0.340 mmol/mg, respectively, suggesting a change in surface charge density. The other finding that CUR release profiles from TPP-crosslinked CHS-NPs did not have a notable lag time of the release can be explained by (a) the small molecular weight of TPP that formed relatively denser NPs and decreased the release rate of CUR, (b) the slow rate of the steady-state release of CUR resembled the rate of the initial release. The increased permeation and release rates of CUR from SHMP-crosslinked CHS-NPs, which are in contrast with the permeation and release of CUR from TPP-crosslinked CHS-NPs, are due to the high molecular weight and “bulky” structure of SHMP crosslinker. Although the repeating hexametaphosphate units provide more potential negatively charged phosphate sites than the linear triphosphate chain of TPP, it has been demonstrated that NPs formed by reaction with SHMP are larger than those fabricated with TPP. This finding confirms a previous report by Abdelgawad and Hudson [46], with similar NPs also prepared at pH = 5. A quantitative porosity characterization of the NPs was not performed in the present study. It is therefore hypothesized and suggested that if SHMP crosslinks CHS in a way that creates less regular or heterogeneous crosslinking domains, the resultant network can have larger voids or internal free volume, opposite to TPP that possesses shorter anionic chain and can produce a more compact and uniform ionic bridges.
The zero-order release kinetics obtained in the present study are in agreement with Nair et al. [43], who also examined the release of CUR from CHS-based NPs. Using the dialysis bag method, this previous report showed that 41.5% of CUR were released from TPP-crosslinked CHS-NPs at pH = 5, and only about 19% were released at pH = 7.4 over a 24-hour period [43]. The data obtained in the present report demonstrated comparable results, in which less than 10% of CUR were permeated through the membrane at pH = 7.4 by 24 h and about 17% over 66 h period when TPP-crosslinked CHS-NPs were tested. However, since a membrane permeability is obviously a limiting factor, the calculated release rates of CUR from the CHS-NPs according to Equation 27, were between 1.1–1.2 μg/h, i.e., 22–24% of CUR were released by 24 h with the four TPP concentrations used. In contrast to TPP crosslinking, CHS-NPs crosslinked with SHMP at levels ranging between 0.136–0.340 mmol/mg CHS showed a higher release rates with notable lag times. A different study using CHS-NPs was reported by Gomathi et al. [50], who used direct dissolution testing to evaluate the release rates of 5-fluorouracil from NPs crosslinked by TPP and SHMP. The release patterns of the this drug from both types of NPs included an initial burst followed by a sustained release [50]. These different patterns stem from a highly water soluble drug adherent to the surface of the NPs followed by the release of the drug trapped within the core. The release from CHS-NPs is not only dependent on the drug properties, but on the NP properties as well. CHS-NP composition and mode of manufacturing, such as crosslinker type, CHS to crosslinker ratio, pH, temperature, and stirring speed are important factors that dictate the in-vitro and in-vivo release profiles of the active compound [48, 51, 52].
To conclude, in this study, TPP- and SHMP-crosslinked CHS-NPs were prepared with loaded CUR as a model active agent, characterized, and compared for their physical properties and in-vitro release of their loading. It was shown that although SHMP resulted in a slight increase in the mean particle size in comparison to the particle size of TPP crosslinker, the EE and CUR loading were constant at the various concentrations of this polyanion. TPP crosslinking, however, demonstrated a decrease in EE at a relatively high concentration of the agent, probably by narrowing the space between the polymeric chains. As indicated by ζ potential measurement, TPP crosslinking at all levels was more uniformly distributed inside the NPs whereas the larger molecular weight SHMP at a low concentration (0.068 mmol/mg CHS) creates NPs mostly by binding to the ammonium groups on the surface. As further described in this study, drug released from NPs can be more adequately determined by Franz diffusion cell system. The method which has been established and developed by our group is based on a model that the drug is first released from the NP into the medium inside the donor chamber and subsequently diffuses through the membrane into the receiver chamber. The release rate of the drug from the particles can be evaluated only if the permeability coefficient of the membrane (P) is determined. After calculating P, the release rates were derived. It has been found that the release rates of CUR from CHS-NPs crosslinked by SHMP at concentrations higher than 0.136 mmol/mg CHS were about twice as high than the release rates of CUR from TPP-crosslinked CHS-NPs, accompanied by notable lag times. This significant increase in the release rates of CUR from SHMP-crosslinked CHS-NPs is explained by the high molecular weight and the large structure of this crosslinker compared to the small tripolyphosphate molecules. To conclude, this report demonstrates the complexity of CHS-based nano systems. The results can be considered as one more step in the mechanistic understanding of CHS formulations, therefore, they are practically useful for any pharmaceutical scientist working in the field.
CHS: chitosan
CHS-NPs: chitosan nanoparticles
CUR: curcumin
DDW: double-distilled water
DMSO: dimethyl sulfoxide
EE%: encapsulation (or entrapment) efficiency percentage
FTIR: Fourier transform infrared spectroscopy
HPLC: high-performance liquid chromatography
IR: infrared
NPs: nanoparticles
NTA: nanoparticle tracking analysis
PBS: phosphate buffered saline
PTFE: polytetrafluoroethylene
SHMP: sodium hexametaphosphate
TEM: transmission electron microscopy
TPP: sodium tripolyphosphate
ζ: zeta
The supplementary figure for this article is available at: https://www.explorationpub.com/uploads/Article/file/1008157_sup_1.pdf.
The authors are grateful for the professional assistance and technical support of the staff at Ilze Katz Institute for Nanoscale Science & Technology.
IE: Investigation, Formal analysis, Visualization, Writing—original draft. DY: Investigation, Formal analysis, Writing—review & editing. SBS: Conceptualization, Writing—review & editing. ACS: Conceptualization, Supervision, Formal analysis, Visualization, Writing—original draft, Writing—review & editing. All authors read and approved the submitted version.
Amnon C. Sintov, who is the Editorial Board Member of Exploration of Drug Science, had no involvement in the decision-making or review process of this manuscript. The other authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Not applicable.
Not applicable.
Not applicable.
The data that supports the findings of this study are available from the corresponding author upon reasonable request.
This research received funding from B.G. Negev (BGN) (the tech-transfer company of Ben-Gurion University of the Negev). The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
View: 873
Download: 22
Times Cited: 0
