Affiliation:
1Department of Computer Engineering, Faculty of Engineering, Igdir University, 76000 Igdir, Turkey
ORCID: https://orcid.org/0009-0002-5721-1100
Affiliation:
1Department of Computer Engineering, Faculty of Engineering, Igdir University, 76000 Igdir, Turkey
ORCID: https://orcid.org/0009-0008-7227-9182
Affiliation:
1Department of Computer Engineering, Faculty of Engineering, Igdir University, 76000 Igdir, Turkey
2Department of Electronics and Information Technologies, Faculty of Architecture and Engineering, Nakhchivan State University, AZ 7012 Nakhchivan, Azerbaijan
3Department of Computer Engineering, Faculty of Engineering and Architecture, Fenerbahçe University, 34758 Istanbul, Turkey
Email: ishak.pacal@igdir.edu.tr
ORCID: https://orcid.org/0000-0001-6670-2169
Explor Med. 2026;7:1001382 DOI: https://doi.org/10.37349/emed.2026.1001382
Received: November 12, 2025 Accepted: January 22, 2026 Published: February 27, 2026
Academic Editor: Lindsay A. Farrer, Boston University School of Medicine, USA
The article belongs to the special issue Artificial Intelligence in Precision Imaging: Innovations Shaping the Future of Clinical Diagnostics
Aim: Interobserver variability continues to limit the consistency of breast ultrasound interpretation. This study compares two Vision Transformer (ViT) models and two Convolutional Neural Network (CNN) models for automated three-class breast ultrasound classification, with a specific focus on the tradeoff between predictive performance and computational efficiency.
Methods: Swin Transformer Base and DeiT Base were evaluated alongside InceptionV3 and MobileNetV3 Large using the public Breast Ultrasound Images (BUSI) dataset, which contains 780 images labeled as benign, malignant, and normal. A consistent on-the-fly augmentation pipeline was applied during training to promote robustness and reduce sensitivity to incidental image variations.
Results: Swin Transformer Base achieved the highest test accuracy (0.9167) and F1 score (0.8981). MobileNetV3 Large reached an accuracy of 0.8583 with substantially lower computational demand. The efficiency contrast was pronounced, with Swin requiring 30.33 GFLOPs versus 0.43 GFLOPs for MobileNetV3 Large.
Conclusions: On this benchmark, ViT models can yield higher classification performance, while lightweight CNNs offer a strong efficiency profile that may better match deployment-constrained settings. These results suggest that model selection should be guided by both predictive accuracy and operational feasibility within the target clinical workflow.
Breast cancer is a malignant disease characterized by uncontrolled proliferation of epithelial cells in breast tissue and remains a major global health challenge [1–3]. Its etiology is multifactorial, involving genetic susceptibility, hormonal influences, and lifestyle and environmental exposures [4–6]. Because clinical outcomes are closely linked to stage at diagnosis, timely and accurate detection remains central to effective management, which has sustained interest in noninvasive imaging-based diagnostic pathways [7, 8].
Mammography is the established standard for population-level screening; however, its diagnostic sensitivity decreases in women with dense fibroglandular tissue and can be lower in younger populations [9–12]. This limitation motivates the use of complementary imaging modalities to reduce missed findings and improve diagnostic confidence [13–15]. Ultrasound is widely used as an adjunct because it is accessible, does not involve ionizing radiation, and supports dynamic assessment in real time [16]. It is particularly valuable for clarifying equivocal mammographic findings and for evaluation in dense breasts [17]. At the same time, ultrasound interpretation can be operator dependent, contributing to interobserver variability and inconsistent reporting across settings [18–20]. These challenges strengthen the case for analysis approaches that are objective, repeatable, and easier to standardize.
In parallel, artificial intelligence-driven image analysis has expanded rapidly across medical imaging, including oncologic applications [21, 22]. Recent work suggests that deep learning systems can support classification, detection, and segmentation tasks, with the potential to reduce variability in routine interpretation when appropriately validated [23–25]. In breast ultrasound specifically, the central question is not only whether automated models can achieve high accuracy, but also whether they do so in a way that is practical for real-world use.
Architecturally, Convolutional Neural Networks (CNNs) have been the dominant approach because they learn robust local features efficiently [26–28]. Vision Transformer (ViT) models have emerged as an alternative by modeling broader contextual relationships, which may be advantageous when diagnostically relevant cues extend beyond localized textures [29, 30]. At the same time, transformer-based approaches may require higher computational resources and can be more sensitive to dataset scale and heterogeneity, which are common constraints in medical imaging research [31].
Recent studies in breast ultrasound have increasingly explored hierarchical transformer designs and multimodal strategies to combine global context with localized feature extraction [32]. For example, multimodal transformer-based fusion has been evaluated for malignancy prediction in retrospective multicenter settings [33], and transformer-driven end-to-end frameworks have been reported for multicenter molecular subtype classification using multimodal ultrasound-derived information [34]. These advances indicate active progress, yet direct benchmarking studies that compare modern transformer models with widely used CNN baselines while also reporting computational efficiency remain relatively limited.
Motivated by this gap, the present study provides a focused comparative evaluation of two transformer-based models, Swin Transformer Base and DeiT Base, against two established CNN architectures, InceptionV3 and MobileNetV3 Large, for three-class breast ultrasound classification. Beyond predictive performance, we explicitly consider efficiency-related indicators to clarify the tradeoff between accuracy and computational demand, supporting more informed model selection for different deployment constraints.
We used the publicly available Breast Ultrasound Images (BUSI) dataset distributed via Kaggle [35]. BUSI comprises 780 ultrasound images labeled into three categories: benign, malignant, and normal. This three-class structure supports a direct evaluation of how well different architectures separate lesion types while also distinguishing normal tissue patterns.
Images were separated using a stratified split of 70 percent for training (n = 545), 15 percent for validation (n = 115), and 15 percent for testing (n = 120). Although K-fold cross-validation is a viable alternative, a fixed hold-out split was preferred here to ensure an identical and completely independent test environment for comparing diverse CNN and Transformer paradigms. Stratification was applied to preserve the original class proportions across subsets and to reduce the risk of majority class dominance during optimization, given the smaller size of the normal category (n = 133) relative to benign (n = 437). The validation subset was used for model selection and early stopping, while the test subset was held out and used only for the final comparison across models. A full breakdown is provided in Table 1.
Distribution of BUSI images across training, validation, and test subsets.
| Classes | Train (70%) | Validation (15%) | Test (15%) | Total (100%) |
|---|---|---|---|---|
| Benign | 305 | 65 | 67 | 437 |
| Malignant | 147 | 31 | 32 | 210 |
| Normal | 93 | 19 | 21 | 133 |
| Total | 545 | 115 | 120 | 780 |
BUSI: Breast Ultrasound Images.
Figure 1 provides representative examples from each class to illustrate the visual variability encountered in BUSI. In general, benign lesions often present more regular margins and relatively homogeneous echotexture, whereas malignant lesions more frequently show irregular borders and heterogeneous internal appearance. Normal examples are included to reflect typical background parenchymal patterns. The figure also highlights common ultrasound characteristics such as speckle and limited contrast, which contribute to the difficulty of the classification task.
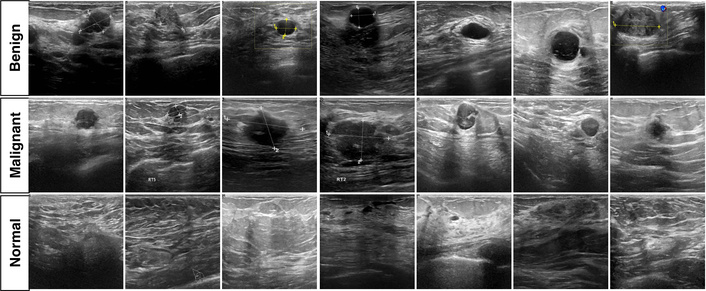
Representative BUSI examples for malignant, benign, and normal categories. BUSI: Breast Ultrasound Images.
To reduce overfitting and improve robustness, we applied on-the-fly augmentation during training [36, 37]. As a preprocessing step, segmentation mask files (mask.png) were excluded because the present work focuses on image-level classification. During training, each image underwent stochastic geometric and photometric transformations so that the model was exposed to new variations across epochs.
The augmentation pipeline included random resized cropping with an area range of 0.08 to 1.00 and an aspect ratio range of 0.75 to 1.33. Cropped regions were then resized to 224 × 224 using randomly sampled interpolation methods (bilinear, bicubic, or Lanczos). Horizontal flipping was applied with probability 0.50, and color jitter of 0.40 was used to perturb brightness, contrast, and saturation. Augmentations were applied only to the training subset, while validation and test images were processed without stochastic transformations to ensure a consistent evaluation setting.
We compared four architectures selected to capture two common design families and two practical extremes in computational footprint: two CNN models (InceptionV3, MobileNetV3 Large) and two ViT models (Swin Transformer Base, DeiT Base). This selection supports a direct assessment of performance alongside efficiency-related costs.
InceptionV3 [38] is a widely used CNN baseline built around inception modules that process multiple receptive field sizes in parallel. This multi-scale design can be advantageous when target patterns vary in size and appearance, which is typical in ultrasound imaging. InceptionV3 also incorporates design choices such as factorized convolutions and auxiliary heads that support stable optimization in deeper networks.
MobileNetV3 Large [39] represents an efficiency-oriented CNN family designed for constrained compute. It relies on depthwise separable convolutions to reduce operations, uses inverted residual blocks, and includes squeeze and excite style channel recalibration. Its architecture was informed by neural architecture search (NAS), making it a strong reference point when computational budget is central.
Swin Transformer Base [40] is a hierarchical ViT that computes self-attention within local windows and shifts window partitions across layers. This design enables cross-window information flow while keeping attention complexity manageable compared with global self-attention. The hierarchical representation also aligns more closely with the multistage feature hierarchy commonly used in CNNs.
DeiT Base [41] targets data efficiency for ViTs through a distillation-based training strategy that introduces a distillation token to transfer information from a teacher model. The motivation for including DeiT is to examine whether a data-efficient transformer training paradigm offers advantages in settings where labeled medical datasets are relatively limited.
The selection of InceptionV3, MobileNetV3 Large, Swin Transformer, and DeiT was strategically intended to evaluate distinct structural approaches to medical image analysis. Swin Transformer and DeiT were chosen to assess the efficacy of global self-attention mechanisms in capturing long-range dependencies, which are often missed by traditional CNNs but are crucial for identifying subtle architectural distortions in breast tissue. Conversely, InceptionV3 was selected for its proven multi-scale feature extraction capabilities, which align with the diverse size and morphology of breast lesions. MobileNetV3 Large was included as a benchmark for computational efficiency, representing a practical baseline for deployment on resource-constrained clinical hardware. By comparing these four paradigms, we aimed to identify the optimal balance between capturing complex pathological patterns and operational feasibility.
All experiments were implemented in PyTorch using the timm library. Training was performed on an Ubuntu Linux workstation equipped with an NVIDIA GeForce RTX 5090 GPU with 32 GB VRAM. A global seed of 42 was used to improve run-to-run consistency. Input resolution was standardized to 224 × 224 pixels. Label smoothing was set to 0.10 to reduce overconfident predictions during optimization.
Optimization used stochastic gradient descent (SGD) with momentum 0.90 and weight decay 2.0 × 10–5. Models were trained for up to 300 epochs with a batch size of 16 under a cosine learning rate schedule with an initial learning rate of 0.10. A linear warmup of 5 epochs was applied starting from 1.0 × 10–5. Early stopping with patience 10 was used based on validation performance, and the best-performing checkpoint on the validation subset was selected for final testing.
The comparative results are summarized in Table 2, which reports predictive performance together with model size and computational complexity. We further examine error patterns for the best-performing model using the confusion matrix in Figure 2 and provide qualitative visual explanations using Gradient Weighted Class Activation Mapping (Grad-CAM) in Figures 3 and 4.
Comparative performance and complexity of the evaluated models.
| Models | Accuracy | Precision | Recall | F1-score | Params (M) | GFLOPs |
|---|---|---|---|---|---|---|
| DeiT Base | 0.8750 | 0.8805 | 0.8437 | 0.8555 | 85.80M | 33.6955 |
| InceptionV3 | 0.8583 | 0.8468 | 0.8337 | 0.84 | 21.79M | 5.6719 |
| MobilenetV3 Large | 0.8583 | 0.8476 | 0.8392 | 0.8408 | 4.21M | 0.4307 |
| Swin Transformer Base | 0.9167 | 0.9409 | 0.8686 | 0.8981 | 86.75M | 30.3375 |
GFLOPs: Giga Floating-Point Operations.
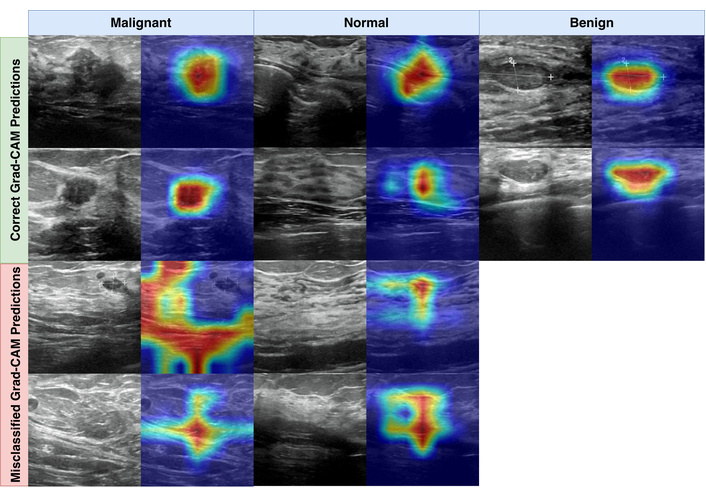
Grad-CAM examples for Swin Transformer Base, showing representative correct and misclassified predictions.
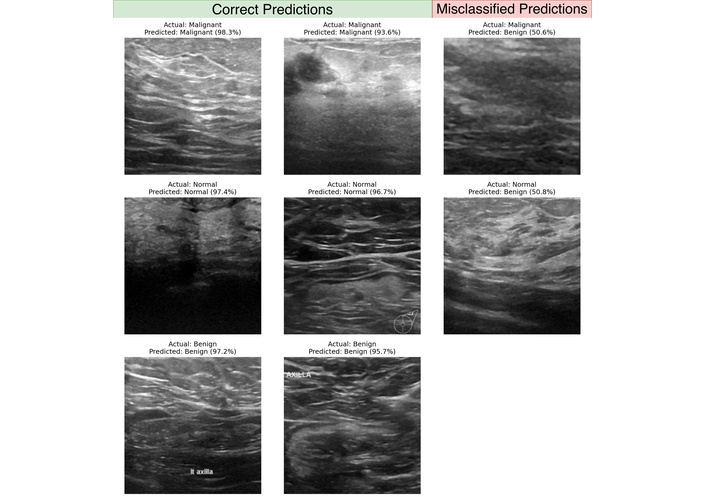
Representative correct and incorrect Swin Transformer Base predictions with prediction confidence values.
Swin Transformer Base achieved the highest overall performance, reaching an accuracy of 0.9167 and an F1-score of 0.8981. This gain came with a substantial computational footprint, including 86.75M parameters and 30.3375 Giga Floating-Point Operations (GFLOPs). In contrast, MobileNetV3 Large reached an accuracy of 0.8583 with 4.21M parameters and 0.4307 GFLOPs. Relative to Swin, MobileNetV3 Large uses approximately 20.61 times fewer parameters and approximately 70.44 times fewer GFLOPs, making the efficiency contrast more pronounced than the accuracy gap.
DeiT Base ranked second by accuracy (0.8750) and achieved an F1-score of 0.8555. Its parameter count (85.80M) is comparable to Swin, while its GFLOPs are slightly higher (33.6955). The CNN baselines placed behind the transformer models in accuracy, with InceptionV3 and MobileNetV3 Large both reaching 0.8583. However, their resource profiles differed markedly. InceptionV3 required 21.79M parameters and 5.6719 GFLOPs, whereas MobileNetV3 Large matched the accuracy with approximately 5.18 times fewer parameters and approximately 13.17 times fewer GFLOPs. These results highlight a consistent pattern: higher accuracy was associated with higher computational demand, while the most compact model provided the strongest efficiency profile.
From an application perspective, the practical choice depends on the deployment context. If the primary aim is maximizing performance under ample compute, a model such as Swin Transformer Base may be preferred. When compute budgets are limited or latency constraints are strict, MobileNetV3 Large offers a favorable accuracy to cost balance. These workflow-oriented interpretations should be treated as provisional until supported by direct measurements of inference latency, memory use, and throughput on the target clinical hardware.
To complement the quantitative metrics, we inspected model behavior using Grad-CAM. Grad-CAM produces heatmaps that indicate which regions contributed most strongly to the predicted class. Heatmaps were generated using the PyTorch Grad-CAM library. For transformer models, maps were extracted from the final normalization layer of the last transformer block, and for CNN models from the final convolutional layer. Saliency maps were normalized with min-max scaling to support consistent visual comparison.
Figure 3 shows representative examples of correct and incorrect predictions. In correctly classified malignant and benign cases, the heatmaps frequently concentrate around lesion regions and their immediate boundaries. For correct normal predictions, attention tends to be distributed across broader tissue patterns rather than focusing on a single focal structure. In misclassified examples, heatmaps can become diffuse or shift toward dense parenchymal regions, which is consistent with common ultrasound challenges such as speckle, low contrast boundaries, and tissue patterns that visually resemble benign margins. These visualizations are post hoc and should be interpreted as supportive qualitative evidence rather than definitive localization of pathology.
Figure 4 provides additional examples with associated prediction confidence values. In the true prediction panel, correct predictions across classes often exceeded confidence 0.9300. In the misclassified panel, errors aligned with the main confusion matrix patterns, including malignant and normal cases predicted as benign, with example confidence values of 0.5060 and 0.5080. These borderline confidence levels suggest that at least a subset of errors may occur in visually ambiguous cases where the model does not strongly separate competing classes.
The results in Table 2 indicate a consistent tradeoff between predictive performance and computational demand when comparing ViT models with CNN baselines for breast ultrasound classification. Swin Transformer Base achieved the strongest overall performance, with an accuracy of 0.9167, supported by a high precision of 0.9409 and an F1-score of 0.8981. This pattern is compatible with the notion that hierarchical attention, implemented through shifted windows, can capture both local and broader contextual cues that are relevant in ultrasound images. At the same time, the computational cost is substantial, with 86.75M parameters and 30.3375 GFLOPs.
In contrast, MobileNetV3 Large delivered an accuracy of 0.8583 at a markedly lower computational footprint, using 4.21M parameters and 0.4307 GFLOPs, representing approximately 20.61 times fewer parameters and approximately 70.44 times fewer GFLOPs than Swin Transformer Base. This gap suggests that a large fraction of the performance obtained by the strongest model can be retained with far lower computational requirements, which may be important when inference latency, memory limits, or energy constraints are central. These deployment-oriented interpretations should remain cautious until confirmed with direct measurements of inference time, throughput, and memory use on the target hardware.
DeiT Base ranked second with an accuracy of 0.8750 and an F1 score of 0.8555, while maintaining a parameter count comparable to Swin Transformer Base (85.80M). The two CNN baselines, InceptionV3 and MobileNetV3 Large, reached the same accuracy value (0.8583) but differed substantially in cost. InceptionV3 required 21.79M parameters and 5.6719 GFLOPs, whereas MobileNetV3 Large achieved the same accuracy with approximately 5.18 times fewer parameters and approximately 13.17 times fewer GFLOPs. This contrast reinforces that architecture choice can materially change operational burden even when headline accuracy is similar.
Limitations should be considered when interpreting these findings. First, the study relies on a single public dataset of 780 images, which may not capture the full variability seen across scanners, acquisition protocols, and patient populations. Second, the class distribution is imbalanced, with fewer normal cases (n = 133) than benign cases (n = 437), which can influence model behavior across classes. Third, the dataset redistribution does not consistently provide patient identifiers, so the split is performed at the image level, and patient-level overlap across subsets cannot be fully excluded. Future work should prioritize external validation on multi-center cohorts, patient-level splitting when identifiers are available, and reporting that supports clinical translation, such as confidence intervals, calibration assessment, and a direct comparison with radiologist performance or established commercial computer-aided diagnosis systems. If the intended use case prioritizes specific error types, such as reducing false negatives for malignant lesions, cost-sensitive objectives, or class-balanced training strategies can be evaluated in a targeted way.
To summarize the overall impact and findings of this comparative evaluation. This study compared two ViT models (Swin Transformer Base and DeiT Base) and two CNN models (InceptionV3 and MobileNetV3 Large) for three-class breast ultrasound classification, emphasizing both performance and computational efficiency. Swin Transformer Base achieved the highest performance, reaching an accuracy of 0.9167 and an F1 score 0.8981, but with high computational demand. MobileNetV3 Large achieved an accuracy of 0.8583 with a substantially lower footprint, supporting its suitability when efficiency constraints are dominant. The results suggest that architecture selection should be aligned with the practical requirements of the intended workflow, and that stronger claims about deployment readiness should be supported by external validation and hardware-specific inference benchmarking.
BUSI: Breast Ultrasound Images
CNNs: Convolutional Neural Networks
GFLOPs: Giga Floating-Point Operations
ViT: Vision Transformer
Experimental computations were performed using the computing resources of Igdir University’s Artificial Intelligence and Big Data Application and Research Center. Artificial intelligence tools were used solely to assist with language-related tasks (e.g., translation, grammar/spelling correction, and improving readability). No AI tools were used for study design, methodology, data collection, data analysis, interpretation of results, or generation of scientific content. The authors take full responsibility for the content of the manuscript.
SN: Conceptualization, Methodology, Investigation, Formal analysis. YC: Conceptualization, Methodology, Visualization, Writing—original draft. IP: Supervision, Project administration, Funding acquisition, Writing—review & editing. All authors read and approved the submitted version.
The authors declare that they have no conflicts of interest.
Not applicable.
Not applicable.
Not applicable.
The datasets analyzed for this study can be found on the Kaggle platform: https://www.kaggle.com/datasets/sabahesaraki/breast-ultrasound-images-dataset.
Financial support is received from the Health Institutes of Türkiye (TÜSEB) under the “2023-C1-YZ” call, project number [33934]. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Yigitcan Cakmak, Ishak Pacal
