Affiliation:
1New Drug Screening and Pharmacodynamics Evaluation Center, National Key Laboratory for Multi-Target Natural Drugs, China Pharmaceutical University, Nanjing 210009, Jiangsu, China
†These authors contributed equally to this work.
ORCID: https://orcid.org/0000-0001-7998-5129
Affiliation:
2College of Life Science, Jilin Agriculture University, Changchun 130118, Jilin, China
†These authors contributed equally to this work.
Affiliation:
1New Drug Screening and Pharmacodynamics Evaluation Center, National Key Laboratory for Multi-Target Natural Drugs, China Pharmaceutical University, Nanjing 210009, Jiangsu, China
ORCID: https://orcid.org/0009-0000-8857-4399
Affiliation:
3Department of Pharmaceutical Engineering, School of Engineering, China Pharmaceutical University, Nanjing 210009, Jiangsu, China
ORCID: https://orcid.org/0009-0006-9137-8691
Affiliation:
4School of Pharmacy, Shanghai Jiao Tong University, Shanghai 200240, China
ORCID: https://orcid.org/0000-0001-5569-3535
Affiliation:
5School of Pharmacy, China Pharmaceutical University, Nanjing 210009, Jiangsu, China
ORCID: https://orcid.org/0009-0009-6067-7893
Affiliation:
1New Drug Screening and Pharmacodynamics Evaluation Center, National Key Laboratory for Multi-Target Natural Drugs, China Pharmaceutical University, Nanjing 210009, Jiangsu, China
ORCID: https://orcid.org/0009-0004-8134-7897
Affiliation:
6Department of Pharmacy, Qurtuba University of Information Science & Technology, Peshawar 25000, Pakistan
ORCID: https://orcid.org/0000-0002-4594-9821
Affiliation:
1New Drug Screening and Pharmacodynamics Evaluation Center, National Key Laboratory for Multi-Target Natural Drugs, China Pharmaceutical University, Nanjing 210009, Jiangsu, China
ORCID: https://orcid.org/0009-0006-9103-7969
Affiliation:
1New Drug Screening and Pharmacodynamics Evaluation Center, National Key Laboratory for Multi-Target Natural Drugs, China Pharmaceutical University, Nanjing 210009, Jiangsu, China
Email: yuanst1967@163.com
ORCID: https://orcid.org/0000-0003-2714-4197
Affiliation:
2College of Life Science, Jilin Agriculture University, Changchun 130118, Jilin, China
Email: cysun@jlau.edu.cn
ORCID: https://orcid.org/0009-0006-2733-3739
Explor Dig Dis. 2026;5:1005120 DOI: https://doi.org/10.37349/edd.2026.1005120
Received: January 01, 2026 Accepted: March 02, 2026 Published: April 13, 2026
Academic Editor: Jose C. Fernandez-Checa, Institute of Biomedical Research of Barcelona (IIBB), CSIC, Spain
The article belongs to the special issue Prevention, Screening and Diagnosis for Primary Liver Cancer
Hepatocellular carcinoma (HCC) functions as a major cancer-related death factor around the world. Research indicates that long non-coding RNAs (lncRNAs) play essential roles during HCC onset and development because they belong to the novel RNA subclass that extends beyond 200 nucleotides without protein-coding capability. LncRNAs regulate the expression of downstream target genes and cancer-related signaling pathways, thereby promoting the proliferation, migration, invasion, autophagy, and apoptosis of tumor cells. The study of lncRNA function has been substantially facilitated by the emergence of lncRNA-specific microarrays and the increased accessibility of next-generation sequencing technologies. The function of lncRNAs can be predicted using computational and molecular methodologies. LncRNAs have the potential to function as repressors, scaffolds, regulators of super-enhancers, or molecular decoys. Proliferation, invasion, survival, DNA damage response (DDR), and chromatin dynamics can all be influenced by lncRNAs. Additionally, they can affect stemness/differentiation. The recurrence of tumors may be facilitated by the aberrant expression of these transcripts, which may result in therapy resistance. LncRNAs have the potential to function as innovative prognostic or theranostic biomarkers in HCC and other malignancies. In addition, RNA-based therapeutics may be implemented to target lncRNAs as a novel treatment approach for primary or recurrent HCC. In this review, we investigate the functions of lncRNAs in the pathophysiology of HCC and suggest their potential for novel therapeutic application in the treatment of HCC.
While most of the human genome receives transcription signals, the total protein-coding genes represent only a small fraction, which amounts to less than 3% of all transcribed genes. Research on the roles of non-coding RNA (ncRNA) has fluctuated between classifying the molecules as unimportant transcripts and crucial epigenetic controllers for many decades. Previous studies on ncRNAs mainly focused on the functions of small RNA molecules, especially microRNAs (miRNAs) [1]. Scientific research has established long ncRNAs (lncRNAs) as molecules that manipulate both biological processes and disease pathogenesis [2]. Among various types of versatile transcripts, lncRNAs demonstrate vital functions that direct subcellular placement and manage transcriptional regulation, and mediate epigenetic modification. LncRNAs produce extensive phenotypic changes because of their genome-wide regulatory capacity [3]. The number of scientific lncRNA studies has experienced explosive growth since 2010, which represents more than 90% of total research about this entity. This is a fast-evolving field of research, with several prospects for groundbreaking discoveries across multiple disciplines. In order to place greater functional and pathological significance on lncRNAs, it is imperative to point out the existing knowledge foundations that provide a systematic correlation between ncRNAs and disease phenotypes and genetic variation. The ncRNA Phenotype Database (ncRPheno), ncRNA Variations (ncRNAVar), and other databases combine extensive experimental and computational data on the links between ncRNAs and human diseases on a global scale, both at the genetic and transcriptional levels, and provide a comprehensive global perspective on ncRNA dysregulation in human pathology. In the same way, ncRNAVar expands on this by compiling empirically validated relationships between ncRNAVars and disease features, containing more than 4,500 interactions in more than 700 diseases, including HCC. It is also worth noting that hepatocellular carcinoma (HCC) is one of the disease phenotypes with the greatest number of associated ncRNA variants in ncRNAVar, which highlights the clinical importance of ncRNA genetic variation in liver cancer [4].
HCC exhibits aggressive characteristics because it typically appears advanced when providing treatment becomes challenging. The diagnosis of HCC occurs mainly at an advanced stage because it leads to poor clinical outcomes rendering surgical resection and liver transplantation (LT) ineffective for treatment [5]. Standard HCC treatment includes surgical procedures along with LT and transarterial chemoembolization (TACE) as well as sorafenib which functions as an FDA-approved kinase inhibitor for advanced HCC cancer cases. Despite rigorous treatment options, the 5-year survival rate for HCC remains poor. Patients with advanced HCC survive for only a short period of less than two years when treated with systemic therapies, including sorafenib and immune checkpoint inhibitors. Recurrence of HCC occurs frequently following liver transplant procedures or resections, which negatively affects the overall poor treatment outcomes for patients with this cancer [6].
Research has uncovered how lncRNAs could play a part in HCC development. These molecules, longer than 200 nucleotides, do not produce proteins; instead, they manage gene expression through three regulatory systems, including chromatin remodeling and epigenetic regulation as well as transcription factor modulation [7]. The non-coding nature of lncRNAs has led researchers to discover their involvement in HCC tumorigenesis, together with metastasis and angiogenesis, alongside drug resistance mechanisms. Through this discovery, there are emerging opportunities in diagnosis and therapy development. In this study, we address what is known about lncRNA function and how it might be utilized to develop future therapeutic techniques in the treatment of HCC.
LncRNAs have extensive effects on transcriptional and post-transcriptional regulation, depending on their subcellular localization. Whereas nuclear lncRNAs mediate the communication between chromatin-modifying complexes and chromatin, and promote higher-order chromatin structure, cytoplasmic lncRNAs mediate the regulation of mRNA decay and protein translation or serve as miRNA bait. With the distinct expression and excellent tissue-specificity of lncRNAs, researchers have been examining the functional crosstalk between lncRNAs and epigenetic machinery in the promotion of HCC progression.
In case of their functionality, lncRNAs that are enriched at their own loci of transcription should be part of the process of chromatin modification. The functions of lncRNAs in regulating the transcriptional activity of local or distant genes can be achieved through various methods, including but not limited to the down-regulation of the activity of epigenetic regulators, direct contact between lncRNAs and the DNA, or enhancing enhancer-promoter interaction.
Abnormal histone modification patterns are also commonly witnessed in the course of HCC development due to epigenetic controller dysregulation. It is important to note that several transcriptionally repressive histone methyltransferases, such as Enhancer of zeste homolog 2 (EZH2), SET domain bifurcated histone lysine methyltransferase 1 (SETDB1), and G9a are often deregulated in HCC, leading to the epigenetic silencing of tumor suppressor genes [8]. To substantiate the argument that lncRNAs once again communicate with these epigenetic modifiers to regulate the expression of genes, a recent paper employed an array of genetic and chemical methods to highlight the roles of RNAs in regulating the activity of polycomb repressive complex 2 (PRC2) [9]. Taurine upregulated gene 1 (TUG1) promotes the binding of PRC2 to H3K27me3 (histone H3 lysine 27 trimethylation) silencing mark at the promoter of Kruppel-like factor 2 (KLF2) and inhibits KLF2 transcription in HCC [10]. In addition to repressive histone methyltransferases, multiple lncRNAs have also contributed to the movement of transcription activators to the particular loci. An example of this action is the interaction between HOTTIP (HOXA transcript at the distal tip) and the WDR5/MLL (WD repeat domain 5/mixed-lineage leukemia protein) complex, which relocates WDR5/MLL to the area surrounding the HOXA locus and results in transcriptional activation of HOXA genes, and consequently, hepatocarcinogenesis [11]. The cis-regulatory action of 91H on its neighboring gene, insulin-like growth factor 2 (IGF2), was also established by another study by recruiting RBBP5 (retinoblastoma binding protein 5) [12]. This recruiting mechanism is also noted with the help of lncAKHE, which works with transcription activator YEATS4 (YEAST domain containing 4) to increase NOTCH2 (notch receptor 2) signaling in HCC [13].
HCC has unique DNA methylation patterns that relate to HCC grades and patient survival. Some of the studies have postulated the functions of lncRNAs to alter the activity of the DNA methyltransferases (DNMTs) which leads to the change of DNA methylation. LncRNA34a has been reported to recruit DNMT3a to enhance the activity of DNA methylation of miR-34a promoter in HCC. This recruitment enhances transcriptional silencing of miR-34a by lnc-34a [14]. In addition to SWI/SNF (switch/sucrose non-fermentable) complexes, HAND2-AS1 (heart and neural rest derivatives 2-antisense1) recruits INO80 (inositol requiring 80) to the BMPR1A (bone morphogenetic protein receptor 1A) promoter region, thereby activating BMPR1A transcription and promoting the self-renewal of liver cancer stem cells (CSCs) [15]. Collectively, these results indicate the functional cross-talk between lncRNAs and the epigenetic modifiers in the regulation of the transcriptional output of the target genes.
Although lncRNAs engage with epigenetic regulators and provide them with the capacity to target gene loci, increasing evidence also indicates that lncRNAs are strongly linked to chromatin architecture. R-loops are one of the ways in which lncRNAs can directly interact with genomic DNA. As an example, the antisense lncRNA VIM-AS1 (vimentin antisense 1) forms an R-loop at the VIM promoter, resulting in the relaxation of chromatin that allows NF-κB binding and increases VIM transcription [16]. The other way through which direct lncRNA-chromatin interactions can occur is through the establishment of RNA-DNA triplexes, which may mediate transcriptional activation [17]. An earlier study reported an interesting model where an antisense lncRNA Khps1 (K-region homologous protein splicing-derived transcript 1) creates an RNA/DNA triplex at the sphingosine kinase 1 (SPHK1) promoter to induce an open chromatin structure and recruitment of CBP/p300, which in turn, induces transcriptional activation of SPHK1 [18]. For example, CCAT1 (colon cancer-associated transcript 1), which is transcribed through a super-enhancer at the upstream end of MYC, promotes transcription of MYC in human colorectal cancer (CRC) by promoting chromatin looping that results in the proximity of CCAT1 to the MYC locus by binding cytosine-thymine-cytosine-factor (CTCF) [19]. All these results underline the essential nature of lncRNAs in the regulation of chromatin structure to facilitate the expression of target genes. Nevertheless, several questions are yet to be answered about what motivates lncRNAs to alter chromatin architecture and whether this type of transcriptional control may contribute to the further progression of HCC.
LncRNAs are not limited to regulating protein localization on chromatin. Most lncRNAs are exported to the cytoplasm and regulate two processes that take place in the cytoplasm, which significantly affect the process of protein translation turnover and translation of the mRNA.
LncRNAs are capable of controlling mRNA dynamics at the post-transcriptional level by (1) miRNA sponge, (2) mRNA contact, and (3) RBP recruitment. The sequence-specific interaction makes lncRNAs fine-tune the gene expression with a preference toward the HCC tumorigenesis through regulating the mRNA stability and miRNA availability. miRNAs have also been shown to bind the target mRNAs, which are suppressed by the lncRNA-mRNA complex, reversing the suppressive action of miRNAs on mRNA targets [20]. As an example, the lncRNA DANCR (differentiation antagonizing non-protein coding RNA) has been shown to bind the miRNA binding site of CTNNB1 (catenin beta 1) on 3′ UTR that amplifies the number of tumor cells with stemness characteristics [21].
Evidence compilation has suggested that lncRNAs are able to mediate protein stability by regulating ubiquitin or proteasome machinery. In HCC, some lncRNAs stabilize oncoproteins by exerting their oncogenic activities [22]. As an example, lncCSMD1-1 binds to and stabilizes MYC protein by inhibiting its ubiquitin-proteasome-mediated degradation, thereby promoting hepatocarcinogenesis [22]. In the same way, lncRNA PSTAR (p53-stabilizing and activating RNA) suppresses the tumorigenicity of HCC through increasing the SUMOylation of heterogeneous nuclear ribonucleoprotein K (hnRNP-K), which also intensifies the interaction between hnRNP-K and p53, resulting in transactivation of p53 [23]. Interestingly, there are lncRNAs that regulate protein translation. GMAN is a gastric cancer metastasis-associated lncRNA that was discovered to bind with eukaryotic translation initiation factor 4B (eIF4B) and stimulate its phosphorylation through inhibition of dephosphorylation of another protein mediator phosphatase 2A subunit B (PPP2R2A). The above changes in turn enhance anti-apoptotic protein expression, thus leading to tumorigenesis of HCC [24].
Research has failed to explain either the biological functions or molecular mechanisms of most lncRNAs because scientists have only partly characterized a small fraction of these molecules. The existing evidence shows that these molecules operate as vital components for regulating particular cellular events with a focus on protein-coding gene expression across epigenetic, transcriptional, and post-transcriptional levels (Figure 1).
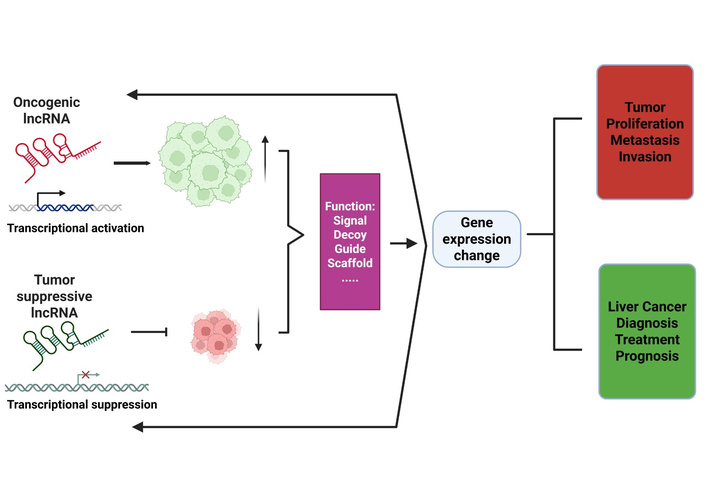
Summary of the molecular roles of lncRNA in liver cancer and its use in clinical therapies. Dysregulation of lncRNAs through epigenetic and genetic changes causes the lncRNAs to play the roles of either activators or suppressors in the process of initiating liver cancer. The dysregulated expression of lncRNAs can serve as a signal, guide, decoy, and scaffold and influence the expression of protein-coding genes, which consequently triggers complex interactions and regulations with either positive or negative effects on tumor proliferation and metastasis. The abnormal lncRNAs can be detected, targeted, and even used as indicators of the prognosis of liver cancer. LncRNA: long non-coding RNA.
The diversity of lncRNA structures results in a wide array of molecular functional roles in both normal and abnormal biology. A complex regulatory system that regulates the expression of lncRNAs has developed as a result of the magnitude of the non-coding transcriptome. The expression pattern of lncRNA molecules depends on developmental processes that make them both cell-type-specific and time-dependent. LncRNA expression in specific tissues helps preserve cell differentiation alongside cellular features. Many EZH2-associated lncRNAs have been demonstrated to exhibit tissue-specific binding capacity, implying a tissue-specific function for lncRNAs in chromatin remodelling and cellular imprinting [25]. Achieving global transcriptional control occurs through two methods: chromatin remodelling along with transcriptional repression/activation. Temporal control of lncRNA expression may influence differentiation and cell fate as global transcription regulators, either directly or epigenetically. LncRNAs utilize specific signals in their 3′ UTRs to regulate spatial positioning as well as temporal activities of their transcript location. Studies show this transcript placement pattern mainly occurs within nervous tissues because their cells have intricate structures [26]. The DNA-binding ability of lncRNAs allows them to recruit proteins, which initiate DNA looping that results in super-enhancer activation both within cis regulators and between trans regulators. The lncRNAs UCA1 and HCCL5 activate YAP gene super-enhancers through their association with epithelial-mesenchymal transition (EMT) transitions in ovarian cancer cells as well as HCC cells [27, 28]. The DNA-binding ability of lncRNAs enables them to attach to promoter areas where they potentially operate as either activating or repressive elements. LncRNAs can bring in transcription factors or DNMTs. The transcriptional machinery cannot make contact with DNA when lncRNAs bind to it in antisense orientation to block transcription. Molecular sponge functions of lncRNA transcripts enable them to trap both transcription factors and miRNAs. These transcripts function as endogenous competing endogenous RNAs (ceRNAs) that have the ability to either deplete proteins/enzymes or small RNA transcripts to block normal functionality. The transcripts can be used to transport their binding targets into subcellular areas where they prompt physiological responses. Messenger RNA stability changes through two distinct actions by lncRNAs: they either trap miRNAs or directly interact with them. When lncRNA binds RNA directly, it causes cells to move proteins or enhance stability while suppressing expression and changing processing outcomes. LncRNAs perform dual roles within the cell by orchestrating alternative splicing events in other transcripts and engaging in post-translational modifications (Figure 2). The intricate structures and substantial lengths of lncRNA transcripts render them ideal components for functioning as subunits of ribonucleoprotein (RNP) complexes. Within these complexes, lncRNAs provide structural support while also guiding them to their designated binding sites. The RNPs that lncRNAs create consist of polycomb repressive complex alongside histone deacetylases (HDACs) and histone acetyl transferases (HATs), as well as other components. The formation of heterochromatin and euchromatin, along with chromatin stability, receives essential contributions from lncRNAs. XIST is a paradigmatic chromatin-regulatory lncRNA that facilitates X-chromosome inactivation, and its female-specific expression is a paradigm example of sex-biased lncRNA expression [29]. Research shows that particular lncRNAs might create heritable effects through modifying epigenetic patterns [30].
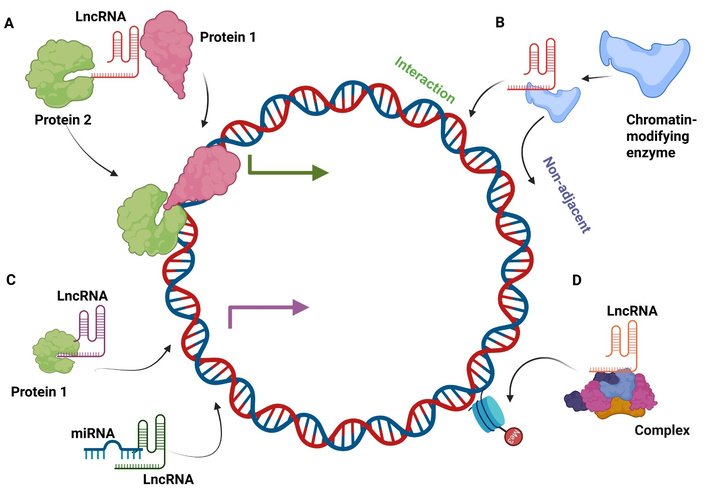
Mechanistic roles of lncRNAs. (A) Signal lncRNA. The primary role of a signal lncRNA is to act as a molecular signal to coordinate the expression of some genes to different stimuli, e.g., transcription factors or chromatin modifiers. (B) Guide lncRNA. Guide lncRNAs are associated with chromatin-modifying enzymes and are directed to the particular location on the genome in order to control gene expression. (C) Decoy lncRNA. Decoy lncRNAs attach to the miRNAs or the transcription factors in order to sequester the miRNA or transcription factor away from its target, affecting translation and transcription. (D) Scaffold lncRNA. LncRNAs are capable of serving as scaffolds to multi-component complexes, including ribonucleoprotein (RNP) complexes, to temporarily form and influence histone modifications. LncRNA: long non-coding RNA; miRNA: microRNA.
The capability of lncRNAs to exist in multiple forms throughout time produces significant cellular phenotypic control. Tumorigenic potential shows signs of being controlled by master regulatory lncRNA function in cancer contexts. Evidence shows that multiple intracellular or phenotypic processes fall under the regulatory potential of various lncRNAs. The investigation of mechanistic roles of lncRNAs remains limited in HCC disease context. This section delineates the established functions of lncRNAs across various malignancies to underscore the unexplored potential that further investigation of lncRNAs in the context of HCC may reveal.
Cancer cell migration and stromal invasion, while related to metastasis, significantly influence disease development and treatment. HCC is considered a whole-organ disease due to the highly invasive and migratory nature of tumor cells within the liver. These characteristics lead to extensive intrahepatic spread, making HCC a particularly aggressive form of cancer [31]. Surgical removal of highly infiltrative tumors proves complex and tends to result in tumor recurrence after surgery. Cancer cell invasion and migration potentials show an association with numerous lncRNAs, which demonstrate both promoting and inhibiting effects. The invasive and migratory behavior in cervical cancer cells increases when lncRNA RHPN1-AS1 is expressed because it regulates the miR-299-3p/FGF2 signaling pathway [32]. Papillary thyroid carcinoma susceptibility candidate 3 (PTCSC3) overexpression leads to inhibition of cervical cancer cell invasion and migration through capturing the miR-574-5p miRNA [33]. Breast cancer cells exhibit enhanced EMT and proliferation due to nuclear enriched abundant transcript 1 (NEAT1), but this noncoding RNA also enables endometrial cancer cells to migrate and invade through controlling miR-144-3p/EZH2 activity [34]. NEAT1 drives metastasis and invasion events in colon cancer cells by means of the miR-185-5p/IGF2 regulatory pathway [35]. Through different signaling pathways, these three lncRNAs, EZR-AS1, LINC00261, and LINC01082, exhibit suppressive effects on cell migration in colon cancer cells [36]. The MYLK-AS1 axis leads EGFR-AS1 to promote migration and invasion in HCC patients [37].
The rapid tumor cell multiplication leads to higher requirements for gas and nutrient exchange processes. Metastasis depends on tumor-associated blood vessels. Both angiogenesis promotion and endothelial cell differentiation capabilities are common in cancer cells [38]. HITT functions as a novel translation-level inhibitor of HIF-1α that is frequently downregulated in human cancers, thus promoting both angiogenesis and tumor growth through HIF-1α expression [39]. The repression of HIF-1α occurs when HITT functions together with EZH2 to establish epigenetic regulation [40]. Breast cancer metastasis and angiogenesis occur due to hypoxic conditions that induce transcription of the lncRNA RAB11B-AS1 [41]. Breast cancer angiogenesis occurs through IGF-1/ERK signaling regulation, which is controlled by the lncRNA NR2F1-AS1 [42]. The NF-κB-interacting lncRNA (NKILA) controls immune responses against cancer together with its ability to regulate blood vessel formation in breast cancer cells by modifying NF-κB/IL-6 signaling pathway activity [43]. The expression of mesoderm-specific transcript (MEST) becomes inhibited by LINC00284 through an NF-κB dependent pathway, which causes increased angiogenesis in ovarian cancer cells [44]. The DANCR facilitates tumor angiogenesis by regulating the miR-145/VEGF relationship in ovarian cancer [45]. The therapeutic outcomes for HCC patients can be improved through LINC01446-targeted therapy because it limits both tumor evolution and angiogenesis through the SRPK2/SRSF1/VEGFA165 pathway [46].
The behavior of cancer cells includes continuous, uncontrolled cellular reproduction. The key characteristic of many cancer cells involves their ability to bypass replicative senescence as they develop replicative immortality by stabilizing their chromosomes, most commonly through extended telomeres [47]. The RNA molecule that plays the primary role in telomere maintenance is telomeric repeat-containing RNA (TERRA) [48]. Elongated telomeres are linked with high TERRA expression in the human placenta [49]. Increased expression of TERRA has been observed in multiple human cancer cell lines [50]. SENEBLOC inhibits cellular senescence through the stabilization of p53-MDM2 interaction, and inhibition of p21 transcription, and also by an independent p53-independent mechanism, which includes HDAC5 [51]. Linc-ASEN disables cell aging through p21 reduction, which permits unlimited cell cycle progression [52].
Cancer cells develop extremely high mutation rates when they undergo multiple divisions triggered by genomic instability. Genomic instability in cells causes the removal of tumor suppressors and the duplication of oncogenes. The loss of the PTEN gene marks an example of this phenomenon in HCC. LncRNA transcripts have been identified as genomic instability regulators. TERRA is a telomere-regulated lncRNA, which helps in maintaining telomeres and stability at chromosome ends, hence maintaining genomic integrity [53]. The lncRNA MANCR (LINC00704) demonstrates an elevated expression within highly mitotic cells in triple-negative breast cancer (TNBC) [54]. ncRNA activated by DNA damage (NORAD) is a long noncoding RNA that functions as a scaffold for the building of topoisomerase complexes essential for preserving genomic stability [55]. NORAD also fosters chromosomal integrity by sequestering destabilizing PUMILIO (PUF family RNA-binding) proteins [56]. When NORAD expression decreases in cancer cells, it may create genomic instability, which produces more copy number variants (CNVs) along with elevated mutational loads. CCAT2 has been shown to induce chromosomal instability by activating the BOP1–AURKB signaling axis [57].
Among the fundamental characteristics of cancer cells are sustained proliferation, together with growth suppressor evasion. Evidence shows that the lncRNA CASC11 (cancer susceptibility candidate 11) promotes bladder cancer cell proliferation by binding to miR-150 [58]. Studies have revealed that the lncRNA CCAT1, which was discovered in CRC, actively stimulates bladder cancer cell proliferation as well as enhances cell migration and invasion [59]. The lncRNA GClnc1 activates proto-oncogene MYC to promote both proliferation and invasion in bladder cancer cells [60]. The antisense lncRNA DLG1-AS1 enables cervical cancer cell proliferation by reducing miR-107, which allows ZHX1 levels to increase [61]. The research community has extensively studied the lncRNA transcript NEAT1 because it activates breast cancer proliferation and EMT pathways [62]. The lncRNA SNHG6 affects breast cancer cell proliferation and invasion through its impact on miR-26a/VASP signaling [63]. The breast cancer proliferation and migration rates increase due to lncRNA PSMG3-AS1-mediated titration of miR-143-3p [64]. Evidence shows that elevated TUG1 levels in HCC tissue enhance cell proliferation and inhibit apoptosis, which indicates its possible use as both a diagnostic tool and therapy target for this disease [10].
Cancer cells employ two distinct immune evasion strategies: intrinsic mechanisms that suppress immune recognition and extrinsic mechanisms that reprogram normal immune cells to support tumor survival. The tumor creates inflammatory situations that promote tumor growth. Research shows that lncRNAs take part in programming cancer cells for immune-related processes. SATB2-AS1 is a tumor-suppressive lncRNA in colorectal cancer; the reduced expression of it is linked to an aggressive form of the disease, whereas its increased expression suppresses the progression of CRC by inhibiting the transcription of Snail through SATB2 and epithelial-mesenchymal transition [65]. LINK-A stands as a novel lncRNA that suppresses tumor antigenicity and immune-related tumor suppression by activating TRIM71 signaling pathways in TNBC cells [66]. NKILA has been shown to promote activation-induced cell death in cytotoxic T cells, resulting in immune evasion in breast and lung cancer [67]. During breast cancer development, SNHG1 regulates regulatory T cell (Treg) differentiation, which affects immune escape through the miR-448/IDO pathway [68]. Some tumors use recruitment tactics to bring tumor-associated macrophages (TAMs) to help tumors grow and survive. Breast cancer cells receive survival-promoting glycolysis stimulation through the HISLA lncRNA, which TAM cells release within extracellular vesicles [69]. The lncRNA DGCR5 functions as a tumor suppressor for HCC while influencing immune-related biological processes by employing three mechanisms that include sponge activity against miR-346 and the regulation of KLF14 and Wnt signaling pathway inhibition [70].
Multiple cancers develop their metastatic ability in specific groups of perivascular cells. Metastatic cells must transfer into lymphatic system before exiting to establish new tumors at sites removed from the primary tumor. The evidence suggests that lncRNAs provide metastatic abilities to these specialized cells. Research shows that the novel lncRNA XLOC_006390 promotes cervical cancer tumorigenesis and metastasis by acting as a competing endogenous RNA against miR-331-3p and miR-338-3p [71]. Under hypoxic conditions, the lncRNA BX111 gets activated to promote pancreatic cancer metastasis through EMT activation of ZEB1 transcription [72]. The lncRNA PTAR facilitates ovarian cancer metastasis through inhibition of miR-101-3p, which releases ZEB1 expression to promote EMT [73]. The functions of lncRNAs are not usually explicitly defined. Metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) has been characterized as imparting metastatic capability, especially in lung-to-brain metastasis via the regulation of EMT [74]. MALAT1 promotes metastatic progression through the suppression of pyroptosis and T cell-mediated destruction of incipient metastatic cells in breast cancer [75]. LINC01133 prevents breast cancer metastasis by regulating SOX4 expression through EZH2 activity [76]. LINC0273 promotes breast cancer metastasis by stabilizing hnRNPL, hence activating AGR2 transcription [77].
Replicative immortality represents a distinctive cancer cellular feature because it enables cells to escape from normal programmed death mechanisms. The regulation of apoptosis by lncRNAs includes both positive and negative effects, which have been documented in this cellular hallmark. In the same manner, lncRNA ENST00000422059 was demonstrated to stimulate the growth of breast cancer cells and suppress their apoptosis via miR-145-5p/KLF5 axis [78]. Similarly, another lncRNA PART1 controls apoptosis in prostate cancer cells through the regulation of Toll-like receptor signaling pathways [79]. LncRNA-ATB has been found to induce apoptosis in non-small cell lung cancer via miR-200a/β-catenin [80]. The absence of the lncRNA MEG3 in prostate cancer cells causes lower apoptosis levels because QKI-5 expression is not restrained by miR-9-5p [81]. Research indicates that LncKLHDC7B links to TNBC resistance against cell death along with unfavorable clinical expectations [82]. CCAT1 enhances the progression of tumor by sponging miR-181a-5p and modulating the autophagy involving ATG7 in HCC [83].
Since HCC has become more resistant to traditional treatment methods such as sorafenib, alternative treatment modalities, such as those that take advantage of a regulated cell death mechanism such as ferroptosis are urgently needed [84]. This is especially considering that inhibition of the retinoblastoma protein has been identified to increase the activity of sorafenib by aggravating ferroptosis in vitro and in vivo [85]. Ferroptosis, a specialized type of programmed cell death that involves accumulation of lipid peroxides and reactive oxygen species, which is iron-dependent, is a promising option to avoid apoptosis resistance that is common in most cancer types. In fact, it has been shown that ferroptosis can be used to suppress HCC progression and increase its sensitivity to sorafenib, and ferroptosis inhibition predictably accelerates the progression of HCC and promotes resistance to the drug [86]. Therefore, the complexity of regulatory mechanisms of ferroptosis, especially lncRNAs, is an essential barrier to the creation of effective treatment methods.
The therapeutic target of aberrant metabolism in cancer cells attracts researchers due to its long-standing attractiveness. Cancer cells exhibit the Warburg effect while producing energy from aerobic glycolysis for their cellular needs [87]. The research shows that LINC00504 stimulates aerobic glycolysis in ovarian cancer cells by depleting miR-1244 [88]. The lncRNA MALAT1 creates simultaneous glycolytic increases and suppresses gluconeogenesis in HCC by enhancing mTOR-mediated translation of transcription factor 7-like 2 (TCF7L2) [89]. EPB41L4A-AS1 functions as a Warburg effect repressor through its binding with HDAC2 and NPM1, which occurs within multiple cancer sites [90]. The lncRNA SNHG14 (small nucleolar RNA host gene 14) acts as an important promoter for cancer progression in HCC. The H3K27 acetylation of PABPC1 regulated by SNHG14 leads to increased cell proliferation as well as migration and angiogenesis in HCC tissues and cells [91]. The research shows that SNHG14 affects metabolic processes in HCC cells by controlling the activity of genes connected to glycolysis and other metabolic pathways. The therapeutic evaluation of SNHG14 or its subsequent targets represents a possible therapeutic approach to treat HCC.
Research indicates that CSCs function as the main agents behind tumor formation while simultaneously causing treatment resistance. CSCs present three essential capabilities, including self-renewal, multipotency, and survival capacity. CSC replication and differentiation produce a hierarchical cell arrangement within tumors, which enhances their aggressive nature and makes them hard to overcome. LncRNAs sustain both stemness properties and cellular fate programs with tumor microenvironment (TME) niches that support CSCs. MALAT1 has also been reported to mediate miR-375/YAP1 axis to promote cancer stem cell properties in HCC [92]. Research indicates that stemness-related transcription factor Oct4 operates in a feedback loop by triggering the expression of MALAT1 and NEAT1 [93]. The YAP transcription factor functions as an oncogene that enables multiple cancer programs including stemness [94]. The lncRNA B4GALT1-AS1 functions to bring YAP into the nucleus so it can boost its transcriptional output, which increases cancer stemness in colon cancer cells [95]. It has been reported that SNHG20 enhances the development of HCC and is linked to an unfavorable prognosis, partially by regulating the EZH2/E-cadherin axis [96].
Standard cancer treatments through radiation therapy or chemotherapy drugs struggle against cancer cells because these cells keep their DNA repair systems in overdrive. The transcriptional programs linked to DDR are controlled by specific lncRNAs. Homologous recombination and other DDR pathways are actively expressed in multiple myeloma (MM) cells when NEAT1 becomes activated [97]. The structural role of NEAT1 in forming paraspeckles plays a potential part in both DDR regulation and activity [98]. The DNA repair mechanism of TNBC depends on LINP1, which functions as a structural part of the IGFBP-3/NONO/SFPQ complex [99]. Alternative NHEJ in MM occurs through MALAT1, which binds PARP1 and LIG3 as well [100]. The sorafenib resistance mechanism in HCC depends on MALAT1, which regulates miR-140-5p/Aurora-A signaling, thus providing potential outlooks for patient prognosis and therapy development when using sorafenib [101]. The research about lncRNAs and drug resistance has been the focus of attention in recent years. Many lncRNAs have been demonstrated to be linked with drug resistance [102]. LncRNAs are important in chemotherapy resistance by controlling major cellular processes like drug efflux, DNA repair, apoptosis, and cell survival [103]. As illustrations, lncRNAs such as the H19 and MALAT1 have been reported to increase drug efflux-related resistance by increasing transporters including the MDR1/ABCB1 (P-glycoprotein) and MRP1, leading to decreased intracellular accumulation of drugs and decreased chemosensitivity [102]. Furthermore, the HCC-related lncRNA HANR has been reported to cause the resistance of HCC cells to doxorubicin through its interaction with GSKIP and subsequent alteration of GSK3β phosphorylation, which is a factor that leads to chemoresistance [104].
Although ICIs have a promising future, a significant percentage of patients show primary or acquired resistance, which restricts their effectiveness, and this indicates the necessity of further investigation of the mechanisms of resistance [105]. LncRNAs have the capability to control different cellular activities that are essential in immune evasion and tumor growth such as the regulation of the expression of immune checkpoint proteins and the complex signal transduction networks that regulate immune cell activities [106]. Moreover, some lncRNAs including lncRNA-MIAT (myocardial infarction associated transcript) and lncRNA-MIR155HG have been directly associated with the expression of immune checkpoints (PD-1, PD-L1, and CTLA-4), hence enabling immune evasion in HCC [103]. This is a regulatory capacity that is also applicable to tumor-infiltrating lymphocytes, and promotes T-cell exhaustion and dysfunction, which in turn impairs the anti-tumor immune response and promotes an immunosuppressive microenvironment favorable to ICI resistance [107].
The TME depends on lncRNAs to progress disease and invade tissues by using these molecules to change tumor-stroma relations and develop new blood vessels and hide from immunity and dismantle extracellular matrix structures [108]. Proliferating fibroblast cells in CRC tumors transfer the H19 ncRNA through exosome transmissions to generate cells with increased resistance to chemotherapy and stem-like properties [109]. Stromal lncRNA lnc-CAF, which is upregulated in the formation of CAF, enhances the growth of OSCC by maintaining IL-33 signaling [110]. Hypoxia-induced HIF-1alpha transcriptionally increases lncRNA HAS2-AS1 in OSCC that facilitates EMT and invasiveness through the stabilization of HAS2 [111]. The TGF-β signaling regulator lncRNA-ATB is believed to increase the risk of HCC metastasis and represents a promising therapeutic candidate to prevent disease spread [112]. To be precise, the MALAT1 overexpression in the HCC cells facilitates the angiogenic activity and creates an immunosuppressive environment. This is achieved when MALAT1 interacts with miR-140 to inhibit the activity of miR-140 and, therefore, enhances the production of VEGF-A, which assists the progression of HCC by promoting angiogenesis and the polarization of macrophages towards the M2 immunosuppressive lineage [113]. In addition to macrophages, lncRNAs have important functions in regulating T-cell functions in HCC, which affects the development of the disease and immune evasion. Indicatively, lncRNA epidermal growth factor receptor (lnc-EGFR) is profusely expressed in Tregs in the HCC. Lnc-EGFR binds to EGFR, prevents its ubiquitination through c-CBL, and enhances downstream signaling through AP-1/NFAT1 to induce differentiation of Treg cells and immune evasion [114]. The MIAT is another lncRNA whose expression is elevated in multiple cell types that are related to the disease, tumor cells, FoxP3+ Tregs, PD-1+ CD8+ T cells, and GZMK+ CD8+ T cells. Moreover, the expression of MIAT is related to the response of patients to sorafenib. The expression of this lncRNA is also correlated significantly with the expression of PD-L1, which is an immune evasion protein expressed by cancer cells [115]. The immune environment in HCC is a complicated interaction of different types of cells, which are coordinated and controlled by lncRNAs. In a study, Jiang and Li [116] have shown that this is indicated by the polarization of macrophages, which is a vital cellular constituent in the environment of HCC interplay.
Conversely, the non-coding developmental regulatory lncRNA, fetal-lethal ncRNA [FOXF1 adjacent non-coding developmental regulatory RNA (FENDRR)], is a miR-423-5p sponge that inhibits the immune-suppressive activities of Tregs. An overexpressed FENDRR competitively binds miR-423-5p and increases growth arrest and DNA-damage-inducible beta protein (GADD45B), which have negative relationships with the number of Treg-cells and leads to decreasing immunosuppressive cytokines TGF-β1 and IL-10, resulting in tumor-cell apoptosis [117]. Besides, lncRNAs Tims and lncNNT-AS1 are linked with less tumor CD4 and CD8 T cell infiltration, which affects the clinical outcomes and immunotherapy response [118].
In HCC, lncRNAs have emerged as key regulators of tumor development, progression, and metastasis (Table 1). The HCC-related lncRNA HULC modifies autophagy by maintaining Sirt1 stability, which affects chemosensitivity. A promising therapeutic strategy exists in targeting HULC or any resulting effects from its downstream mechanisms to improve HCC treatment results [119]. Similarly, the HCC cell proliferative capabilities decrease through a CUL4A-mediated ubiquitination pathway as lncRNA AIRN promotes cell death by controlling STAT1 ubiquitination [120].
Key lncRNAs, pathways, and functions in HCC.
| LncRNAs | Pathway/Process | Function | Clinical relevance | Reference |
|---|---|---|---|---|
| LINC00958 | Lipid metabolism | Promotes lipogenesis and tumor progression | Upregulated in HCC, associated with tumor progression, metastasis, and poor prognosis | [121] |
| PAPAS | miR-188-5p axis | Promotes cell proliferation/tumor progression | Involved in tumour progression; possible function in transcriptional dysregulation in cancer | [122] |
| TUG1 | miR-137/AKT2 axis | Promotes cell proliferation and invasion | High expression correlates with advanced stage, poor prognosis, and therapy resistance | [123] |
| LINC01503 | MAPK/ERK pathway | Activates MAPK/ERK signaling, promoting tumor growth and metastasis | Associated with aggressive tumor behavior and unfavorable prognosis | [124] |
| HULC | NF-κB pathway | Modulates inflammation and immune responses, contributing to HCC progression | Poor prognosis, early diagnosis | [125] |
| TINCR | miR-375/ATG7 axis | Regulates cell proliferation and invasion | Overexpression is associated with tumour advancement and reduced overall survival | [126] |
| lnc-PIK3R1 | PI3K/AKT pathway | Inhibits tumor progression by modulating the PI3K/AKT pathway | Dysregulation associated with HCC development and pathway-driven oncogenesis | [127] |
| LncRNA NORAD | miR-211-5p/FOXD1/VEGF-A axis | Promotes proliferation, migration, and angiogenesis | Overexpression correlates with advanced stage and poor prognosis | [128] |
| CASC2 | miR-155/SOCS1 axis | Regulates cell proliferation and invasion | Reduced expression in HCC is linked to a poorer prognosis | [129] |
| USP27X-AS1 | USP7-mediated AKT stabilisation | Promotes hepatocellular carcinoma progression | Overexpression predicts aggressive phenotype and poor clinical outcome | [130] |
| MALAT1 | MAPK/ERK, apoptosis, EMT | EMT, apoptosis, metastasis | Post-transplant recurrence, survival | [113] |
| NEAT1 | IL-6/STAT3 signaling | Invasion, ferroptosis, metastasis, proliferation, drug resistance | Drug resistance, poor outcome | [131] |
LncRNAs: long non-coding RNAs; HCC: hepatocellular carcinoma; PAPAS: promoter and pre-rRNA antisense; TUG1: taurine upregulated gene 1; AKT2: AKT serine/threonine kinase 2; MAPK/ERK: mitogen-activated protein kinase/extracellular signal-regulated kinase; HULC: highly upregulated in liver cancer; TINCR: terminal differentiation-induced non-coding RNA; ATG7: autophagy related 7; PI3K/AKT: phosphoinositide 3-kinase/protein kinase B; NORAD: non-coding RNA activated by DNA damage; CASC2: cancer susceptibility candidate 2; MALAT1: metastasis-associated lung adenocarcinoma transcript 1; EMT: epithelial-mesenchymal transition; NEAT1: nuclear enriched abundant transcript 1.
HCC metastasis depends heavily on EMT as lncRNAs function remarkably in controlling this developmental stage. LncRNA HOTAIR drives EMT by bringing the PRC2 complex, which results in decreased E-cadherin while enhancing vimentin expression [132]. In HCC cells, inhibition of apoptosis together with autophagy occurs because MALAT1 targets the miR-146a/PI3K/AKT/mTOR pathway. The HCC cells survive and proliferate more aggressively because of this mechanism. The therapeutic approach to suppress MALAT1 expression shows promise for enhancing HCC patient treatment success rates and survival outcomes [133].
CSCs function as the source of new tumors and recurring cancer manifestations in HCC patients. MALAT1, lncRNA, enhances the ability of self-renewal and maintenance of cancer stem cell properties in HCC by sponging miR-375, and regulating YAP1 [92]. H19 lncRNA serves as a cancer-causing gene to advance both liver cancer types, along with HCC and cholangiocarcinoma, through its effects on cell proliferation, apoptosis, and invasion and metastasis behaviors [134]. These findings emphasize the role of lncRNAs in sustaining the CSC population in HCC.
The Warburg effect, also referred to as aerobic glycolysis, represents a defining characteristic of cancer metabolic functions. SNHG1 lncRNA has also been demonstrated to induce glycolysis in the miR-326/PKM2 axis in HCC [135]. On a mechanistic level, CCAT1 reacts with PTBP1 and stabilizes it by preventing its ubiquitin-mediated degradation, thus, increasing glycolysis and advancing gastric cancer development [136]. These studies highlight the role of lncRNAs in the metabolic reprogramming of HCC cells.
Hypoxia represents a widespread condition within HCC TMEs, so lncRNAs assume vital functions in hypoxic adaptation. An axis that includes HIF1A together with lncRNA RAET1K and miR-100-5p influences hypoxia-mediated glycolytic activity in HCC cells, which could affect HCC disease progression [137]. Additionally, the stability of BSG mRNA increases due to lncRNA BSG-AS1, which drives HCC cells to proliferate and migrate, while BSG-AS1 overexpression indicates unfavorable outcomes [138]. The study shows that lncRNAs play an essential part in the development of HCC caused by hypoxic conditions.
Therapy resistance is a major challenge in HCC treatment, and lncRNAs have been implicated in modulating resistance to various therapies. LncRNA H19 plays a role in sorafenib resistance in HCC by upregulating miR-675, offering new potential therapeutic strategies for sorafenib-resistant liver cancer patients [139]. Similarly, the cancer drug resistance mechanisms of lncRNA PVT1 involve regulatory functions that control transcription as well as splicing and epigenetic processes [140]. These studies highlight the role of lncRNAs in therapy resistance and the need to target them for overcoming resistance.
LncRNAs have emerged as promising diagnostic and prognostic biomarkers in HCC (Figure 3). Research shows that lncRNA HULC displays high expression levels in both HCC tissues and sera, while its expression levels match with tumor size, metastasis, and poor patient outcomes [141]. Similarly, the consolidated prognostic marker built from 9 m6A regulator-related lncRNAs shows promise as a diagnostic tool and prognostic indicator for HCC [142]. The findings demonstrate the practical applications of using lncRNAs as biomarkers in HCC.
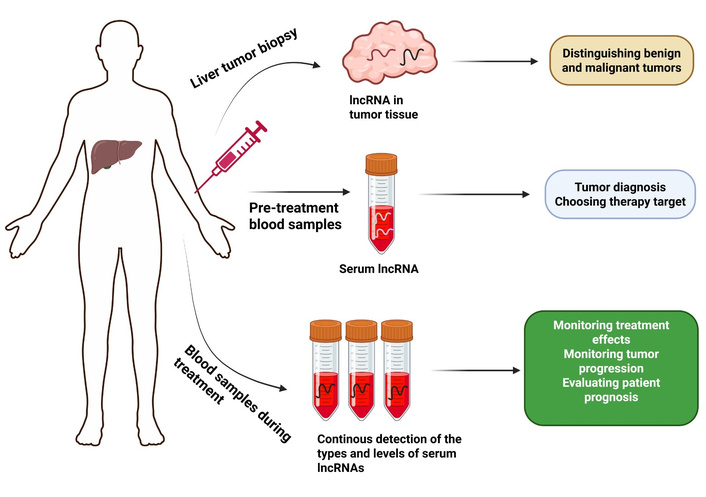
Potential clinical applications of lncRNAs as biomarkers for HCC diagnosis and treatment. The use of lncRNAs serves as a marker to distinguish tumors of benign characteristics from those that are malignant. The diagnostic use of serum lncRNAs as potential biomarkers for HCC continues to advance, so they can serve alongside clinical references for treatment decisions. The effectiveness of treatment for HCC patients, together with disease progression prediction and prognostic assessment, can be evaluated through ongoing blood monitoring and lncRNA type and level assessments during treatment. LncRNAs: long non-coding RNAs; HCC: hepatocellular carcinoma.
Given their critical role in HCC pathophysiology, lncRNAs represent attractive therapeutic targets. Science has investigated multiple approaches to focus on lncRNAs through RNA interference (RNAi), antisense oligonucleotides (ASOs) and CRISPR/Cas systems [143]. Research has shown that blocking lncRNA HOTAIR by RNAi treatment both reduces HCC cell proliferation and cell metastasis abilities [144]. In addition, nanoparticle-based delivery systems have been created to increase the stability and bioavailability of lncRNA-targeting therapies [145]. These studies highlight the potential of lncRNA-based therapies in HCC treatment.
In order to further focus on the HCC-specific mechanisms, it is imperative to combine the evidence in the in vivo models that represent the complicated molecular and pathological environment of liver cancer. Recent rat HCC research has shown that therapeutic intervention with sorafenib, Theranekron, deinoxanthin, and desloratadine attenuates major pathways associated with carcinogenesis, apoptosis, oxidative stress, and inflammatory signaling that are processes known to be regulated by HCC-related lncRNAs [146]. Combined Theranekron dextromethylamide and sorafenib treatment of experimental HCC models had a strong inhibitory effect on tumor progression, causing apoptosis, enhancing antioxidant capacity, and restoring normal biochemical profiles, including cell-cycle arrest and mitochondrial dysfunction pathways, which are commonly regulated by lncRNAs such as MALAT1 and NEAT1 [147]. Further in vivo research also showed that the treatments regulate clinically relevant biomarkers like alpha-fetoprotein (AFP), heat shock protein 90 alpha (HSP90α), and indoleamine 2,3-dioxygenase 1 (IDO1), which supports their translational applicability to the development of HCC and therapy response [148]. Also, the protective action of deinoxanthin on sorafenib-induced nephrotoxicity and the anti-tumor activity of desloratadine through inhibition of TLR4/MYD88/NF-κB signaling pathway further indicate that inflammatory and immune-related pathways are necessary in HCC and are becoming a more frequently recognized subject of lncRNA-mediated control [149, 150]. Together, the three in vivo studies provide mechanistic support of the pathways typically regulated by oncogenic lncRNAs and the reinforced biological relevance of lncRNA-based regulatory networks in HCC.
Recent findings have radically changed the old perception according to which lncRNAs do not have the ability to code any protein and have shown that a significant fraction of lncRNAs contain non-canonical open reading frames that encode functional microproteins or small peptides. It is emerging that these microproteins coded by lncRNAs are key regulatory factors in tumorigenesis because they can directly regulate important oncogenic signaling pathways. In HCC, Zhao et al. [151] systematically discovered hundreds of translatable lncRNAs and established that the lncRNA ASH1L-AS1 encodes a microprotein, APPLE, which consists of 90 amino acids and is strongly expressed in HCC tissues and is linked to poor prognosis. Mechanistically, APPLE enhances HCC development by maintaining MAPK/ERK signaling through direct interaction with ERK1/2 and preventing PP1/PP2A-mediated ERK dephosphorylation, which is a non-canonical pathway activation mechanism that does not rely on classical RAS mutations [151]. These results demonstrate that lncRNAs may have oncogenic actions that are mediated by both RNA-mediated regulatory processes and through the products of their translation, microproteins, and so the translational potential of lncRNAs should be included in future analysis of HCC biology and therapeutic targeting.
It is increasingly evident that lncRNAs play key post-transcriptional regulatory roles, including but not limited to alternative splicing, and thus play a role in cancer pathogenesis and progression, such as in HCC. Malakar et al. [152] have shown that the lncRNA MALAT1 enhances the development of hepatocellular carcinoma by increasing the activity of the splicing factor, SRSF1, which leads to oncogenic alternative splicing and tumor growth. Mechanistically, MALAT1 has an oncogenic effect through the upregulation of SRSF1 and alteration of alternative splicing of cancer-related transcripts, which is accompanied by the activation of Wnt signaling and the output of the mTOR pathway, which triggers the development and progression of HCC [152]. These results have suggested alternative splicing as a major post-transcriptional mechanism by which lncRNAs promote hepatocarcinogenesis, and the crucial role of integrating splicing-based lncRNA mechanisms into models of HCC pathogenesis.
Beylerli et al. [153] emphasized the fact that tumor cells secrete lncRNAs into human biological fluids, creating stable circulating lncRNAs, insensitive to RNA degradation. These lncRNAs have been detected in cancer patients as being expressed aberrantly [153]. In this way, lncRNAs are becoming an effective alternative to the conventional biomarkers (AFP, etc.) to diagnose HCC and provide a prognosis. The effectiveness of AFP as an early predictor of HCC has been a controversial subject since it raises the question of its sensitivity and specificity [154]. LncRNAs provide greater sensitivity and specificity, which may overcome limitations of conventional markers, e.g., AFP [155]. The resultant effect is an increasing need to identify new diagnostic markers that can potentially substitute AFP, and lncRNAs may be one of the possible alternatives. Thus, the expression level of these lncRNAs may be an important indicator of progression and prognosis of the diseases.
As the trend moves towards minimally invasive and non-invasive methods of diagnostic technology, circulating lncRNAs in serum are currently under intense research. As an example, overall survival, progression-free survival, tumor size, TNM stage, levels of C-reactive proteins, T stage, and portal vein thrombosis were all correlated with high serum levels of lncRNA-ATB, which suggests that they might be useful as serum biomarkers in patients with HCC [156].
Prominently, the lncRNAs like MVIH, X91348, and HOTTIP have demonstrated potential as prognostic markers in HCC. MVIH has been identified as a high level of microvascular invasion which is known to be an independent risk factor of recurrence-free survival and overall survival in HCC patients [157]. Similarly, lncRNA X91348 is significantly downregulated in HCC relative to healthy subjects, and reduced X91348 expression is associated with unfavorable overall survival [158].
In addition, a number of studies revealed lncRNAs, including UCA1 and WRAP53, as the potential biomarkers in the diagnosis of HCC when combined with AFP [159]. LINC00152, RP11-160H22.5, and XLOC014172 were also found to be new biomarkers of HCC by another group. A combination of these lncRNAs with the traditional marker AFP was observed to increase the accuracy of diagnosis of HCC, which suggests that they can be used to enhance the diagnosis of HCC [160].
In the environment of chemotherapy resistance, some lncRNAs, including CAHM, were recognized as main predictive biomarkers. By applying machine learning algorithms, CAHM was described as a central lncRNA, whose expression is upregulated in sorafenib-resistant cell lines, which makes it a promising biomarker for chemotherapy resistance in the future [161].
In short, lncRNAs have become the potential biomarkers of HCC, and the importance of lncRNAs in diagnostic and prognostic precision medicine is clear. Despite the fact that PIVKA-II is a well-known biomarker of HCC, and lncRNAs have been independently identified as promising non-invasive biomarkers, there is little direct comparison between PIVKA-II and lncRNA biomarkers [162]. Large multicentric studies are required to confirm the clinical importance of lncRNAs. Also, it is essential to standardize approaches to detect lncRNAs that are circulating to achieve the same results. Nevertheless, the prospects of studying lncRNAs as biomarkers in HCC indicate that such studies can improve the diagnosis and prognosis of this malignancy, as well as targetable treatment.
LT is a highly demanded intervention to treat patients with HCC and cirrhosis, yet the emergence of HCC after the surgery is a critical issue that affects patient survival. LncRNAs have emerged as important regulators of diverse cellular functions such as tumorigenesis, immune regulation, and tissue repair, which are important in determining the success of LT in HCC patients. The role of lncRNAs in the development of HCC has been emphasized in a number of studies, with molecules such as HOTAIR and MALAT1 contributing to the proliferation, migration, and invasion of tumor cells [163]. The abnormal expression of the following lncRNAs can be a contributor to the aggressive progression of HCC, thus enhancing the chances of the tumor relapse after transplantation. Moreover, NEAT1 and other lncRNAs have been associated with sensitizing immune cell activation and the immunological reaction, which is essential in graft rejection or tolerance [164]. Unregulated lncRNAs in patients with HCC can upset the immune homeostasis, which can cause transplant rejection or post-surgical complications. Other than immune modulation, lncRNAs are also essential to liver regeneration, which is necessary to promote graft survival following LT. Research indicates that lncRNAs such as lncRNA-HEIH have a role in the proliferation of hepatocytes and changes in their expression may result in the failure of liver recovery after transplantation [165]. As illustrated by Lai et al. [166], MALAT1 is a lncRNA and is overexpressed in HCC and is linked to a high probability of tumor recurrence after LT. This research indicates that high MALAT1 expression is an independent prognostic factor for low disease-free survival, especially in patients who surpass the Milan criteria. In addition, the in vitro silencing of MALAT1 decreased cell viability, motility, and invasiveness and increased apoptosis, indicating that it could serve as a therapeutic target in the treatment of HCC patients to improve their LT [166]. Preclinical models showed promising outcomes of lncRNA-targeting in reducing metastasis and improving survival, and thus, lncRNA-targeted therapy may be considered to add to the overall success of LT in HCC patients.
Many researchers analyze and categorize hidden disease markers that are hard to identify. Diagnostic, prognostic, and theragnostic serve as the three primary types of biomarkers, which do not necessarily require exclusive distinctions from each other. Biomarkers with diagnostic functions help distinguish between normal tissues and cancer tissues. The expression levels of prognostic indicators demonstrate an independent relationship with disease prognosis in addition to disease progression. The detectable expression alterations of theragnostic biomarkers show forecasting capability regarding therapeutic response. Certain cancer-related lncRNAs are detectable in serum, even in circulating exosomes, and thus lncRNAs can be used as noninvasive biomarkers [167]. The diagnostic value of MEG3 tumor suppressor has been reported in the serum of CRC patients [168]. Studies reveal that CASC9 demonstrates elevated expression levels in HCC patients versus normal healthy subjects, along with prognostic potential for tumor metastasis outcomes [169]. Multiple examples of lncRNAs are activated due to radiation therapy [170]. The cellular induction of LINP1 occurs after radiotherapy in cervical cancer cells to enable DNA repair [171]. HOTAIR, BRM, and ICR serum lncRNA have demonstrated the potential of being prognostic biomarkers of HCC. High serum concentrations of these lncRNAs were linked to undesirable clinicopathological characteristics and unfavorable prognosis, and a combination of their evaluation could help to identify HCC at an early stage. Diagnostic performance was also compared in the mentioned study in combination with AFP and the panel of AFP and HOTAIR, BRM and ICR exhibited the best diagnostic accuracy [172]. The molecule MALAT1 presents an opportunity for both prognostic assessment and therapeutic interventions among HCC patients receiving sorafenib treatment [101]. The expression levels of lncRNA-GAS5 together with the promoter region rs145204276 polymorphism serve as prognostic biomarkers that predict postoperative pain experienced by patients who undergo hepatectomy for HCC [173].
So far, the overall prognosis of HCC is poor, and at least partially it is explained by the absence of a therapeutic target. The targeted drug most frequently used in the treatment of HCC is sorafenib, which targets RTKs, but resistance to sorafenib is commonly observed with the treatment of HCC [174]. The important functions of lncRNAs in HCC render them attractive drug targets for new treatment regimens. Moreover, the lncRNA-targeting strategies possess certain benefits in comparison with protein-targeting strategies base-pairing principle is far simpler to use than designing a certain protein-binding inhibitor. ASOs and RNAi are the primary forms of these lncRNA-targeting and have both been demonstrated to exhibit positive anti-cancer effects against HCC through lncRNA targeting. For example, RNAi-mediated knockdown of CASC9 has been shown to suppress HCC growth [175]. In a nutshell, ASOs are short single-stranded DNAs that bind to a target lncRNA and make a DNA-RNA complex, which can be cleaved by RNaseH. RNAi, on the other hand, is short, double-stranded RNAs, which must be loaded into AGO2 protein to form an RNA-induced silencing complex (RISC) and subsequently, bind to target lncRNA to produce RNA-RNA complex and silence lncRNAs [176]. In addition to the knockdown of oncogenic lncRNAs, the delivery of tumor suppressive lncRNAs can also be used as an option. As an example, lncRNA PRAL can be a tumor suppressor by stabilizing p53, and its delivery by adenovirus vector has been demonstrated to significantly prevent HCC growth in mice bearing tumor significantly, which indicates its possible clinical use in the treatment of HCC [175]. It is important to mention that the latest pipeline of HCC-targeted ASOs and RNAi is extensive, and both of them have already been used to treat HBV. In short, ASOs and siRNAs are conjugated using chemical groups (e.g., N-acetylgalactosamine, GalNAc) or packaged in delivery vehicles (e.g., lipid nanoparticles); thus, an optimized pharmacokinetic profile is obtained [177]. The commercial experiences of HBV treatment with ASOs and RNAi give a significant foundation for lncRNA interference and the treatment of HCC.
LncRNA-targeted therapy also offers novel concepts for treating the disease. Unlike traditional treatments such as chemotherapy, which affect both cancer cells and normal ones, targeted therapies are designed to target and interfere with the very specific disease-causing parts, all without causing as much damage to normal cells as possible. Clinical trials have begun with research in H19-based treatment methods. A plasmid expressing diphtheria toxin can suppress tumor growth, and this has been confirmed in bladder cancer under the dual control of H19 and IGF2-P4 and is likely to be involved in HCC that expresses high levels of H19 [178]. Another possible therapeutic intervention is the interference of related regulatory pathways by exogenous lncRNA. In earlier research, scientists loaded MS2 virus-like particles (VLPs) with GE11 polypeptide and were able to form a MEG3 carrier, which was directed to EGFR-positive hepatocarcinoma cells using the clathrin-mediated endocytosis [179].
In the recent past, immunotherapy has been a hot topic in the cancer therapy field. With significant roles in immunoregulation, lncRNAs have the potential for immense application in immunotherapy [180]. Indicatively, lncEGFR has the capacity to specifically bind EGFR and stabilize EGFR expression in the inhibition of its interaction with c-CBL and subsequent ubiquitination, which leads to differentiation of Tregs, inhibition of cytotoxic T cells, and disease progression in HCC [114]. Thus, lncRNA targeting and regulation have great potential to help immunotherapy and should be investigated in the future.
Within recent years, an increasing amount of interest has focused on the development of lncRNA-targeting nanoparticle-based therapies for HCC. The HCC nanoparticle therapyʼs most popular lncRNA is MEG3, which is a tumor suppressor and regulates p53 target gene expression. Nevertheless, lncRNA MEG3 exhibits a relatively low or no expression in human HCC. Thus, MEG3 has been used to inhibit the growth, migration, and invasion of HCC cells. Ren et al. [181] examined the importance of using polymeric nanoparticles in the delivery of select lncRNAs to HCC cells. They used a polymer termed pullulan, which has an efficient cell-targeting capacity (targeting liver cells). The purpose of this approach was to provide plasmids with the lncRNA MEG3 and the p53 gene. The delivery of these plasmids was done by co-delivery through pullulan-based ethanolamine-modified poly(glycidyl methacrylate) (PuPGEA). This approach was successful in improving the lncMEG3 and p53 levels in the HCC cells, following which the HCC cell proliferation, migration, invasion, and tumor growth were suppressed further. The paper has shown that these polymeric nanoparticles can be used as dual-targeted theranostic therapy of the HCC [181].
Wider knowledge about lncRNA processes will develop new therapeutic intervention targets. The standard chemotherapeutics often generate insufficient treatment results while patients develop resistance to these treatments. Early-stage HCC diagnosis allows curative treatments to be effective, but advanced HCC receives limited therapeutic options. The use of therapeutic methods against ncRNAs or through ncRNA molecules shows promise to improve HCC treatment.
Among the available variety of methodologies to detect lncRNAs, quantitative polymerase chain reaction (qPCR), RNA sequencing, and microarrays are commonly used, and each of them has specific strengths and limitations in terms of work with clinical samples such as blood or tumor tissue. Comprehensive studies of ncRNA transcripts are enabled by next-generation sequencing and microarrays, but NGS has higher sensitivity and does not require prior knowledge of target sequences [182]. qRT-PCR, on the other hand, is characterized by its sensitivity, reproducibility, and quantification capabilities of trace ncRNAs, which are especially useful in confirming the results of the larger screening techniques such as microarray or RNA sequencing [183]. Moreover, lncRNAs and other ncRNAs, including miRNAs and circular RNAs (circRNAs), are characterized by great stability and degradation resistance, which means that they can be detected by regular laboratory methods [184]. Nevertheless, these approaches to cell-free RNA (cfRNA) in liquid biopsies, particularly for long RNA, have drawbacks such as low reproducibility, technological and biological variability, and the absence of standard procedures [185]. In order to overcome these challenges, more advanced sequencing methods, including those that can forego poly-selection in library preparation or employ single-molecule long-read sequencing, are being developed to enhance comprehensive disease profiling and accurate identification of lncRNAs and their isoforms in complex biological samples [186].
Multiple obstacles persist in the execution of lncRNA applications for HCC research. The introduction of lncRNA therapeutics represents a bright future in precision medicine, which provides new ways of treating a wide range of pathologies, regulating gene expression/epigenetics, and organizing chromatin [187]. Although their potential is enormous, the ethical aspects of the development and implementation of such RNA-based treatments, in light of the unprecedented success of mRNA vaccines, require careful consideration. This is accompanied by a strict analysis of the factors, including risk-benefit analysis, patient autonomy, and fair access to such advanced treatments [188]. Moreover, the targeted and ubiquitous regulatory functions of lncRNAs present complicated concerns about off-target effects and long-term changes to physiological functions, thus necessitating the necessity to provide extensive safety evaluations [189].
Scientists have not clarified the molecular workings of lncRNAs alongside their context-specific functions and lack efficient delivery methods for therapeutic targeting of lncRNAs. Anticipated treatment of HCC through personalized lncRNA-based therapies faces obstacles due to its heterogeneous nature. Future studies should concentrate on untangling the molecular connections of lncRNAs in HCC, finding consistent biomarkers for treatment response, and investigating creative delivery methods for lncRNA-targeting medications.
LncRNAs demonstrate high versatility for creating numerous transcripts that have functions in both healthy processes and disease states. LncRNAs present themselves in multiple configurations with diverse sets of functional properties. A cell expresses regulatory mechanisms that dictate gene expression patterns through both spatial and time-dependent controls. The low frequency of occurrence of lncRNAs in cellular matter does not reduce their ability to significantly affect cell programs and trait manifestation. LncRNAs participate in all core cancer hallmarks by potentially creating targetable universal activities across cancer types. Emerging as important controllers of HCC pathophysiology, lncRNAs have been implicated in cell proliferation, apoptosis, EMT, cancer stemness, and treatment resistance. Their aberrant behavior in HCC draws attention to their possible use as prognostic and diagnostic indicators. Moreover, lncRNA targeting is a possible therapeutic approach for HCC management. Their mechanisms still need to be thoroughly understood by more study as well as the creation of successful treatment strategies. Even though lncRNAs have shown promise in cancer diagnostics of HCC, puzzles remain to be solved. The research on lncRNA remains in its early development, and its mechanism is yet to be clearly understood. Moreover, the targets of lncRNAs are too many, and the regulatory network of a particular lncRNA is hard to describe completely. Moreover, explaining the association of lncRNAs with the progression of HCC is especially essential to us as we will be able to fully comprehend the disease process, find appropriate treatment targets, and develop more effective and better treatment plans.
AFP: alpha-fetoprotein
ASOs: antisense oligonucleotides
BMPR1A: bone morphogenetic protein receptor 1A
CASC11: cancer susceptibility candidate 11
CCAT1: colon cancer-associated transcript 1
CRC: colorectal cancer
CSCs: cancer stem cells
DANCR: differentiation antagonizing non-protein coding RNA
DDR: DNA damage response
DNMTs: DNA methyltransferases
EMT: epithelial-mesenchymal transition
EZH2: Enhancer of zeste homolog 2
FENDRR: FOXF1 adjacent non-coding developmental regulatory RNA
HCC: hepatocellular carcinoma
HDACs: histone deacetylases
hnRNP-K: heterogeneous nuclear ribonucleoprotein K
HOTAIR: HOX transcript antisense intergenic RNA
HOTTIP: HOXA transcript at the distal tip
IGF2: insulin-like growth factor 2
Khps1: K-region homologous protein splicing-derived transcript 1
KLF2: Kruppel-like factor 2
LncRNAs: long non-coding RNAs
LT: liver transplantation
MALAT1: metastasis-associated lung adenocarcinoma transcript 1
MEST: Mesoderm-Specific Transcript
MIAT: myocardial infarction associated transcript
miRNAs: microRNAs
ncRNA: non-coding RNA
ncRNAVar: non-coding RNA Variations
ncRPheno: non-coding RNA Phenotype Database
NEAT1: nuclear enriched abundant transcript 1
NKILA: NF-κB interacting long non-coding RNA
NORAD: non-coding RNA activated by DNA damage
PART1: prostate androgen-regulated transcript 1
PRC2: polycomb repressive complex 2
PSTAR: p53-stabilizing and activating RNA
RNAi: RNA interference
RNP: ribonucleoprotein
SNHG14: small nucleolar RNA host gene 14
SPHK1: sphingosine kinase 1
TAMs: tumor-associated macrophages
TERRA: telomeric repeat-containing RNA
TME: tumor microenvironment
TNBC: triple-negative breast cancer
Treg: regulatory T cell
TUG1: taurine upregulated gene 1
WDR5/MLL: WD Repeat Domain 5/Mixed-Lineage Leukemia protein
Xist: X-inactive specific transcript
YEATS4: YEAST domain containing 4
M Sulaiman: Writing—original draft, Conceptualization. YN: Writing—original draft, Conceptualization. ZC: Resources, Data curation. UEH: Formal analysis. FR: Funding acquisition. MMY: Visualization, Validation. ZZ: Visualization, Validation. M Sher: Conceptualization. MY: Funding acquisition. SY: Resources, Data curation, Supervision. CS: Resources, Data curation. All authors read and approved the submitted version.
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Not applicable.
Not applicable.
Not applicable.
Not applicable.
Not applicable.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
View: 1492
Download: 29
Times Cited: 0
Balasubramaniyan Vairappan ... Biju Pottakkat
