Affiliation:
1Polytechnic University of Coimbra, 3045-093 Coimbra, Portugal
2H&TRC - Health & Technology Research Center, Coimbra Health School, Polytechnic University of Coimbra, 3045-043 Coimbra, Portugal
ORCID: https://orcid.org/0009-0009-1910-310X
Affiliation:
1Polytechnic University of Coimbra, 3045-093 Coimbra, Portugal
2H&TRC - Health & Technology Research Center, Coimbra Health School, Polytechnic University of Coimbra, 3045-043 Coimbra, Portugal
3Research Center for Natural Resources, Environment and Society (CERNAS), Polytechnic University of Coimbra, 3045-601 Coimbra, Portugal
4MARE - Marine and Environmental Sciences Centre/ARNET - Aquatic Research Network, University of Coimbra, 3000-456 Coimbra, Portugal
Email: valado@estesc.ipc.pt
ORCID: https://orcid.org/0000-0002-0157-6648
Explor Neuroprot Ther. 2026;6:1004147 DOI: https://doi.org/10.37349/ent.2026.1004147
Received: September 04, 2025 Accepted: February 10, 2026 Published: April 15, 2026
Academic Editor: Vladimir Josef Balcar, The University of Sydney, Australia
Background: Multiple sclerosis (MS) is a chronic, autoimmune, inflammatory disease that affects the central nervous system. Although the true etiology of MS remains unknown, recent research suggests that it arises from a combination of genetic vulnerability and environmental factors. The human leukocyte antigen (HLA) region is a highly polymorphic locus on chromosome 6 encoding antigen-presenting molecules central to adaptive immunity. MS exhibits significant genetic and geographic heterogeneity, reflecting complex interactions between HLA polymorphisms and environmental influences. Risk and protective alleles differ across populations, reinforcing the importance of studying such variations to better understand the disease’s pathogenesis and guide therapeutic strategies.
Methods: This systematic review followed the “Preferred Reporting Items for Systematic reviews and Meta-Analyses” (PRISMA) guidelines, and a bibliographic search was conducted in the Medline (PubMed) and Web of Science databases using the keywords “Multiple Sclerosis”, “Genetic Polymorphisms”, “SNPs”, and “Human Leukocyte Antigen”.
Results: Twenty-one studies were included, comprising a total of over 50,000 participants across diverse populations. The reviewed studies demonstrate that the alleles DRB1*15:01, DQB1*06:02, DRB1*03:01, DRB1*04:01, DRB1*15:03, DPB1*03:01, as well as the haplotypes DRB1*15:01~DQB1*06:02 and DRB1*15:01~DQA1*01:02~DQB1*06:02, show high expression and are strongly associated with MS susceptibility. In contrast, the alleles A*02:01 and DRB1*01:01 have shown a protective role.
Discussion: The evidence confirms a central role of HLA class II alleles and conserved extended haplotypes, particularly DRB1*15:01-containing haplotypes, in MS susceptibility, while highlighting protective alleles and marked variability across ancestral backgrounds. These findings underscore the importance of high-resolution HLA typing, standardized haplotype definitions and inclusion of diverse populations to refine MS risk estimates.
Multiple sclerosis (MS) is a chronic, autoimmune, inflammatory disease that affects the central nervous system (CNS) [1]. It affects 2.9 million people worldwide, being more frequent in women (69%) than in men (31%), and is the most common cause of neurological disability in young adults [1, 2]. It is characterized by inflammation, demyelination of the myelin sheath, axonal loss, and brain atrophy, leading to motor, sensory, and cognitive impairments [1, 3]. Although the exact etiology of MS remains unknown, recent research suggests it results from a combination of genetic susceptibility and environmental factors [4]. Genetic, environmental, and infectious factors, along with smoking and low vitamin D levels, all contribute to the development of this complex and heterogeneous disease [4, 5].
The disease is known for its highly heterogeneous clinical onset and progression, which are mostly unpredictable [6]. The International Advisory Committee on Clinical Trials in Multiple Sclerosis categorized the clinical course descriptors, also known as Lublin-Reingold descriptors, into four main categories [7, 8]. The relapsing-remitting MS (RRMS) is characterized by distinct acute episodes (relapses) separated in time with dissemination in space (i.e. lesions in more than one region of the CNS), followed by recovery [9, 10]. However, partial or complete recovery may become less evident as relapses persist over time, often progressing to secondary progressive MS (SPMS) [7]. The SPMS describes the transition from a previously relapsing-remitting disease to a progressively disabling course, with or without relapses [8, 9]. The primary progressive MS (PPMS) is marked by continuous worsening of neurological disability from disease onset, without relapses or remission phases [7, 10]. The progressive relapsing MS (PRMS) is rarer and presents with steady deterioration from the beginning of the disease, occasionally accompanied by relapses [7].
Diagnosis typically occurs between the ages of 20 and 50, with 85% of cases initially presenting as RRMS and 15% as PPMS [7]. The diagnosis is primarily clinical, based on the assessment of symptoms, neurological examination, and additional investigations such as magnetic resonance imaging (MRI) and cerebrospinal fluid (CSF) analysis [4, 9]. The degree of patient disability should also be evaluated using a standardized scale. The Expanded Disability Status Scale (EDSS) assesses eight functional systems: visual, brainstem, pyramidal, cerebellar, sensory, bladder, bowel, and cerebral functions. The EDSS ranges from 0 (no disability) to 10 (death due to MS) [11].
According to the literature, the core of the disease’s pathogenesis lies in the imbalance between autoimmunity and self-tolerance [12]. MS is defined by the presence and activity of autoreactive T cells that escape clonal deletion and are peripherally activated by the presentation of myelin antigens by antigen-presenting cells (APCs) [12–14]. This process may occur through molecular mimicry between a microbial antigen and the myelin autoantigen [12]. After crossing the blood-brain barrier (BBB), microglia present antigens to T cells through classical major histocompatibility complex (MHC) class I and II pathways [3]. In the MHC class II pathway, microglia and other APCs present peptides from extracellular proteins to CD4+ T cells [13]. These cells secrete specific cytokines and differentiate into various T helper (Th) cell subsets. Th1 cells release pro-inflammatory cytokines such as interleukin-2 (IL-2), tumor necrosis factor-alpha (TNF-α), and interferon-gamma (IFN-γ), while Th17 cells secrete IL-17, IL-21, and IL-22. In contrast, the MHC class I pathway presents peptides derived from endogenously synthesized proteins to CD8+ T cells, which can recognize and kill infected or abnormal cells [3]. Altogether, these processes lead to increased acute neuroinflammation in the CNS, resulting in demyelination, axonal damage, and progressive disease worsening [13].
The treatment of MS has evolved significantly over the past decades, focusing primarily on modulating inflammatory activity, preventing disability progression, and improving patients’ quality of life. Currently, MS management involves relapse treatment, disease-modifying therapy (DMT), and functional and symptomatic rehabilitation. DMTs represent the cornerstone of MS treatment, particularly in RRMS, as they primarily act by reducing inflammatory activity and, consequently, the frequency of relapses, although they do not effectively prevent the underlying neurodegeneration driving disease progression. This therapy acts through complementary immunomodulatory mechanisms; for example, IFN-β and glatiramer acetate dampen pro-inflammatory T-cell activity. In SPMS, treatment options remain limited; however, recent advances have shown that simvastatin may reduce brain atrophy, and ongoing clinical trials are evaluating biotin, laquinimod, and ocrelizumab for this disease form. In PPMS, therapeutic resources are also scarce, with the main goal being to slow functional decline, since inflammation is less prominent. For the treatment of acute relapses, methylprednisolone remains the drug of choice. In cases of progressive MS that is refractory to conventional DMTs, autologous hematopoietic stem cell transplantation has emerged as an effective alternative [1, 4, 15].
Genome wide association studies (GWAS) test common genetic variants across the entire genome, typically in large cohorts of individuals with and without the trait, to identify associations between specific genetic markers [e.g., single nucleotide polymorphisms (SNPs)] and that trait [16]. Until 2007, when the first GWAS [17] was published, the only confirmed genes associated with MS were human leukocyte antigen (HLA)-DR15, -DQw6, and the class I HLA alleles A3 and B7 [16]. Since then, the International Multiple Sclerosis Genetics Consortium (IMSGC) has conducted several GWAS with progressively larger cohorts; in 2019, the study included 47,429 MS patients and 68,374 controls [18]. GWAS have successfully identified multiple genetic loci associated with MS by scanning the genotypes of large population samples, and have detected more than 200 SNPs associated with an increased risk of developing the disease [19]. However, due to linkage disequilibrium, SNPs located near risk variants also tend to be inherited together, amplifying the genetic association with MS [16].
In the 1970s, the first genetic factor associated with MS was identified: the MHC region, known as the HLA [16]. It was recognized as a key genetic determinant in autoimmune diseases, including MS, accounting for approximately 10.5% of the genetic variance underlying MS susceptibility [20, 21]. The HLA locus comprises a highly complex network of 224 functional genes, densely clustered and interconnected, both anatomically and biochemically [13]. The molecules encoded in this region are highly polymorphic cell surface glycoproteins, essential for the adaptive immune response and for antigen presentation to T cell recognition [22]. HLA genes are located across a 3.6 Mb segment on the short arm of chromosome 6 (6p21) [16].
There are three class I HLA proteins: HLA-A, -B, and -C, encoded by their respective genes. These proteins consist of three domains (α1, α2, and α3) and form membrane-bound heterodimers together with the invariant β2-microglobulin. Antigenic peptides bind to the α1 and α2 domains and are recognized by CD8+ T cell receptors (TCRs). The class I HLA structure allows for the binding of peptides nine to ten amino acids long, and these molecules are expressed on all nucleated cells and platelets, with the exception of intact neurons, the cornea, sperm cells, and some trophoblastic cells in immunoprivileged sites [16].
A second set of HLA genes encodes class II HLA molecules: HLA-DR, -DQ, and -DP. These are composed of two different protein chains, α and β, which pair to form the functional class II molecules. The α1 and β1 domains of these chains bind peptides that are recognized by CD4+ TCRs. While class II HLA molecules typically bind peptides of nine to ten residues, they can present peptides up to 25 amino acids, as their peptide-binding groove is open at both ends, unlike class I molecules. Class II HLA expression is typically restricted to APCs, but under inflammatory conditions, it can also occur in other cell types, such as thyroid epithelial cells, astrocytes, and oligodendrocytes [16].
HLA-DQ and HLA-DP molecules are composed of polymorphic α and β chains, encoded by HLA-DQA1, -DPA1, -DQB1, and -DPB1 genes. HLA-DR molecules consist of a non-polymorphic α-chain (DRα), encoded by HLA-DRA, and a highly polymorphic β-chain (DRβ), which may be encoded by four different genes: HLA-DRB1, -DRB3, -DRB4, and -DRB5 [16]. Many polymorphisms in the DRB chain have been associated with increased susceptibility to specific diseases [13].
Class II HLA expression is essential for maintaining resting immune cell populations and for regulating immune tolerance [13, 23]. Furthermore, HLA class II molecules are strongly associated with autoimmunity due to TCR remodeling influenced by genetic and environmental factors. Their structure facilitates the spontaneous presentation of self-antigens, triggering a signaling cascade that can lead to persistent autoreactivity [13, 23]. Genetic mutations in HLA genes or the presence of certain specific alleles can significantly influence diseases such as fibromyalgia, Alzheimer’s disease, infectious diseases like COVID-19, and autoimmune diseases such as MS [13].
MS exhibits remarkable global heterogeneity, both in its prevalence and in the genetic factors that influence disease susceptibility [4, 5]. Its geographic distribution reflects a complex interaction between genetic polymorphisms, particularly in the HLA region, and environmental factors [20, 21]. Studies have shown that the contribution of HLA alleles to MS risk varies significantly across different populations. For example, alleles such as HLA-DRB1*15 and HLA-DQB1*06:02 increase disease risk in multiple ethnic groups, although with varying magnitudes [6]. Similarly, alleles considered protective in some populations may not exert the same effect in others, highlighting the importance of genetic variation in understanding the disease. The investigation and identification of polymorphisms associated with increased MS risk may help uncover the biological mechanisms underlying disease progression, as well as predict treatment response [4, 6]. In this context, the present systematic review aims to examine HLA-associated polymorphisms with high expression in MS across different populations, deepening our understanding of the genetic influence on the disease’s heterogeneity.
This systematic review was written in accordance with the Preferred Reporting Items for Systematic reviews and Meta-Analyses (PRISMA) guidelines [24].
To guide the relevant research for this systematic review, the following keywords were defined: “Multiple Sclerosis”, “Genetic Polymorphisms”, “SNPs”, and “Human Leukocyte Antigen”, using the PubMed and Web of Science databases. Additionally, the terms “Europe”, “America”, “Africa”, and “Population” were used to maximize the inclusion of studies involving diverse population groups. The details of the search are described in Table 1.
Search details for each database.
| Database | Search details |
|---|---|
| PubMed | ((Multiple Sclerosis) AND ((Genetic Polymorphisms) OR (SNPs) OR (Human Leukocyte Antigens))) Filters: Free full text, Full text, Adaptive Clinical Trial, Case Reports, Classical Article, Clinical Study, Clinical Trial, Comparative Study, Controlled Clinical Trial, Meta-Analysis, Randomized Controlled Trial, Humans, Exclude preprints, from 2019 - 2025((Multiple Sclerosis) AND (HLA) AND ((Europe) OR (America) OR (Africa) OR (Population))) Filters: Free full text, Full text, Humans, from 2019 - 2025 |
| Web of Science | (ALL=(Multiple Sclerosis) AND (ALL=(Polymorphisms) OR ALL=(SNP) OR ALL=(HLA))) and Open Access and 2020 or 2021 or 2022 or 2023 or 2024 or 2025 or 2019 (Publication Years) and Article (Document Types)((ALL=(Multiple Sclerosis) AND ALL=(HLA)) AND (ALL=(Europe) OR ALL=(America) OR ALL=(Africa) OR ALL=(Population))) and Open Access and 2025 or 2024 or 2023 or 2022 or 2021 or 2020 or 2019 (Publication Years) and Article (Document Types) |
SNPs: single nucleotide polymorphisms; HLA: human leukocyte antigen.
Inclusion criteria were established, such as original research studies, randomized clinical trials, and meta-analyses involving human subjects, written in Portuguese or English, with full and free access, and published between 2019 and 2025.
For the exclusion criteria, articles with unrelated titles, abstracts, or content, as well as reviews, systematic reviews, and duplicate articles from different databases were excluded from this selection.
All articles retrieved from the search were organized using reference management software (Mendeley Desktop 1.19.8). After the removal of duplicates, the articles were screened by title and abstract based on the established criteria. The screened articles were then analyzed in full text, and only those with applicable information were selected. The PRISMA flowchart outlines the selection process (Figure 1).
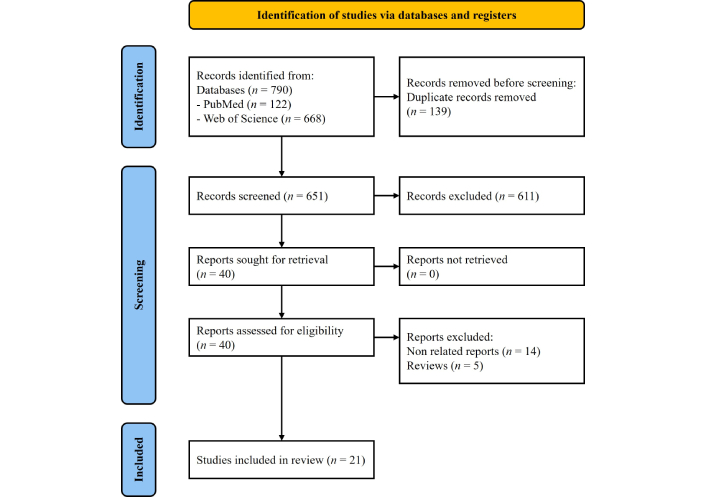
PRISMA flowchart with the description of the selected articles. PRISMA: Preferred Reporting Items for Systematic reviews and Meta-Analyses. Adapted from https://www.prisma-statement.org/prisma-2020-flow-diagram. Accessed May 5, 2025. Distributed under a Creative Commons Attribution (CC BY 4.0) license.
All studies included in this article were categorized as non-randomized; therefore, the risk of bias was assessed using the ROBINS-I tool, considering bias due to confounding (D1), selection of participants (D2), classification of interventions (D3), deviations from intended interventions (D4), missing data (D5), measurement of outcomes (D6), and selection of the reported results (D7) [25, 26]. Of the 21 articles evaluated, the study “Distribution of Major HLA-A, -B, -DR, and -DQ Loci Potentially Associated with Multiple Sclerosis in a Healthy Population from Southern Morocco” was judged at serious risk of bias in outcome measurement because, in the context of this review, it did not directly assess MS cases or HLA-MS associations, but only reported HLA frequencies in healthy individuals, providing indirect evidence for the outcome of interest. The results of the risk of bias assessment are summarized in Figure 2 and explained in Figure S1.
According to the PRISMA guidelines, a total of 790 articles were retrieved, of which 139 duplicates were removed. The remaining 651 articles were screened based on title, abstract, and eligibility criteria, resulting in 40 articles selected for full-text assessment. A total of 40 full-text articles were evaluated for eligibility, and based on this assessment, 21 articles were included in this systematic review. The genetic markers and typing methods used by the included studies are summarized in Table S1. All data and characteristics of the selected studies are presented in the following Table 2.
Characteristics and main results of the included studies.
| Reference | Author/Year | Database | Study objective | Summary of results |
|---|---|---|---|---|
| [27] | Asouri et al./2020 | Web of Science | Genotyping of 14 SNPs in HLA-DRA and 14 SNPs in IL2RA by NGS in 102 Iranian MS patients | No significant association found between the SNPs and MS |
| [28] | Chi et al./2019 | PubMed | Genetic mapping and ancestry analysis of HLA alleles in African American, Hispanic, and Asian populations | Differences in MS risk were detected depending on HLA allele ancestry. European alleles DRB1*15:01, HLA-B*07:02, and HLA-A*03:01 conferred higher MS risk than their African counterparts |
| [29] | Fguirouche et al./2025 | Web of Science | Genotyping of HLA-A, -B, -DR, -DQ in healthy individuals from southern Morocco | Predisposing alleles identified: DRB1*03, *13, *15. No differences found between HLA class I alleles and MS susceptibility. Further studies in MS patients are needed |
| [30] | Khdair et al./2025 | Web of Science | Genotyping of HLA-DRB1 and HLA-DQB1 in MS patients from Jordan | Significant associations found between HLA alleles and MS susceptibility: DRB1*03:01 and DRB1*04:01 |
| [31] | Beecham et al./2022 | PubMed | Analysis of genetic risk modified by ancestry in HLA haplotypes in a multiethnic population | SNPs rs2844503, rs3021302, and rs760145 showed effects modified by global/local ancestry. HLA-A*02:01 had a stronger protective effect when of European origin, and HLA-B*53:01 conferred protection when inherited from African origin, especially among Hispanics |
| [32] | Goodin et al./2021 | PubMed | Analysis of full HLA haplotypes in African Americans with MS | The extended haplotype DRB1*15:01~DQB1*06:02 was most strongly associated with MS. The DRB1*15:03~DQB1*06:02 haplotype had a less consistent association depending on the specific haplotype |
| [33] | Osoegawa et al./2021 | PubMed | NGS genotyping of 11 HLA genes in families with MS | Association found between the haplotype DRB5*01:01~DRB1*15:01 and the allele DPB1*104:01 with MS. Alleles such as DRB1*01:01 and DQB1*03:01 showed a protective effect |
| [34] | Akel et al./2022 | PubMed | High-resolution HLA class II sequencing in Swedish MS patients | 69 distinct genotypes detected; extended haplotypes such as DRB5*01:01~DRB1*15:01-DQB1*06:02 were more frequent; protective and risk effects depended on specific combinations |
| [35] | Briggs and Sept/2021 | Web of Science | Identification of genetic association patterns in MS patient data | Combinations of variants such as HLA-DRB1*15:01, SNP rs56678847, and rs6880809 conferred a 20.2-fold increased risk of developing MS. Computational methods identified complex susceptibility patterns |
| [36] | Hedström et al./2021 | PubMed | Investigation of gene-environment interaction (HLA and factors such as smoking, EBV, obesity) | DRB1*15:01 allele increased the risk of developing MS additively, and the absence of A*02:01 increased the risk fivefold |
| [40] | Boullerne et al./2024 | PubMed | Validation of tagging SNPs for HLA risk alleles in MS using the 1000 genomes panel | SNPs with high performance for inference of DRB1*15:01, DQB1*06:02, and A*02:01 were identified across multiple populations |
| [37] | Barnes et al./2021 | PubMed | Analysis of common genetic variants in MS in Orkney and Shetland populations | SNP rs9271069 was more frequent in the islands than on the mainland and partly explained the excess MS cases |
| [38] | Gontika et al./2020 | Web of Science | Genotyping HLA-DRB1 in POMS and AOMS patients | HLA-DRB1*03 allele was significantly higher in the POMS group than in the AOMS group. In the POMS group, DRB1*11, *15, *03, *04, and *16 were identified as predisposing |
| [39] | Derdelinckx et al./2020 | PubMed | Analysis of myelin antigen reactivity in MS patients with different HLA class II genotypes | No correlation found between HLA class II genotype and myelin peptide reactivity |
| [41] | Mack et al./2019 | PubMed | Genotyping of HLA class I and II by NGS and analysis of KIR loci and their ligands | Association of DRB1*15:01 with MS was confirmed. Protective haplotypes identified: C*03:04~B*40:01; A*02:01 |
| [42] | Creary et al./2019 | Web of Science | High-resolution sequencing of DRB1*15:01 and DRB1*04:01 haplotypes in North Americans | Confirmation of DRB1*15:01~DRB5*01:01 haplotype in MS susceptibility; DRB1*04:01 had haplotype-dependent effects |
| [43] | da Silva Bernardes et al./2019 | PubMed | Genotyping of HLA-DR15 in familial MS cases in Brazil | Extended HLA-DR15 haplotype was found in 44% of familial cases and in none of the healthy controls |
| [44] | Asouri et al./2020 | Web of Science | Genotyping of 36 SNPs located in HLA-DRA, IL2RA, and HMGB1 genes using PCR and NGS | SNPs rs4935356, rs3177928, and rs7197 were considered predisposed to MS |
| [45] | Watanabe et al./2021 | Web of Science | Genotype-phenotype correlation with HLA-DRB1/DPB1 alleles in MS | DRB1*15:01 was associated with susceptibility and worse prognosis in MS; DRB1*04:05 with earlier onset |
| [46] | Ogawa et al./2019 | PubMed | NGS genotyping of 16 HLA genes (classical and non-classical) in Japanese MS patients | DRB1*15:01, DRB1*04:05, B*15:01, and B*39:01 were independently associated with MS |
| [47] | Burnard et al./2022 | Web of Science | Application of penalized regression (elastic net) to identify HLA and NK SNP associations with MS | Variants overlooked by traditional GWAS were identified, such as SNP rs2844482 |
SNPs: single nucleotide polymorphisms; HLA: human leukocyte antigen; NGS: next-generation sequencing; MS: multiple sclerosis; EBV: Epstein-Barr virus; POMS: pediatric-onset MS (early-onset, pediatric and adolescent, MS accounts for approximately 3–5% of all MS cases); AOMS: adult-onset MS; PCR: polymerase chain reaction; NK: natural killer; GWAS: genome wide association studies.
Although the association between HLA antigens and MS has been known for over four decades, the confirmed link remains limited to a small number of genes and alleles, including HLA-DRB1 and the DRB1*15:01 allele [27]. This allele alone accounts for approximately 10% of the genetic variance in MS susceptibility, and its association has been consistently replicated across nearly all populations [27–29]. Its influence extends beyond disease onset, as the structural conformation of DRB1*15:01 enhances binding affinity for myelin basic protein (MBP) epitopes, thereby facilitating autoantigen presentation and chronic immune activation [27, 28].
The molecular mechanisms associated with the risk of developing MS through HLA-DRB1*15:01 include structural and expression alterations. The risk is linked to the presence of an alanine residue at position 71 of the DRβ chain (DRβ71), which creates a wide and hydrophobic P4 pocket in the peptide-binding groove, favoring the spontaneous presentation of autoantigens such as MBP by MHC II+ cells. Alleles like DRB1*15:06 and DRB1*13:09 also share this structural similarity, though they occur less frequently in the general population [30].
The HLA-DRB1*03:01, HLA-DRB5*01:01, and HLA-DQB1*06:02 alleles have also shown evidence of being associated with MS risk in various populations, with the latter in strong linkage disequilibrium with DRB1*15:01 [31–34].
Some alleles may play a protective role against MS due to altered molecular patterns. For example, DRB1*01:01 appears protective due to its reduced binding affinity for MBP peptides and non-classical fragments, resulting in lower binding rates and more efficient kinetic discrimination between endogenous autoantigens and exogenous peptides. Another example is HLA-A*02:01, whose protective role has been shown to be independent [29, 32, 35]. Moreover, the combination of DRB1*15:01 and the absence of A*02:01 increases MS risk fivefold [36].
Osoegawa et al. [33] conducted a study involving 477 families, each with one MS patient and two unaffected parents. Their results showed that the HLA-DRB5*01:01:01~HLA-DRB1*15:01:01:01 haplotype block was significantly associated with MS susceptibility [odds ratio (OR) = 3.02; confidence interval (CI) = 2.55–3.58; p < 2.22 × 10–16].
Besides having bidirectional effects on MS risk, HLA alleles and haplotypes demonstrate significant genetic heterogeneity, population effects, epistasis, and sexual dimorphism, reflecting the biological complexity of these associations [35]. Some MS susceptibility variants show effects modified by global and local ancestry, suggesting that ancestral differences may partly stem from correlated socioeconomic or environmental factors [31]. Beecham et al. [31] supported this premise, observing in the Hispanic population that two SNPs—rs2844503 and rs3021302—exhibited differing effects depending on Native American ancestry proportions. SNP rs2844503 significantly increased MS risk in individuals with high Native American ancestry, whereas rs3021302 showed the opposite pattern: conferring higher risk in those with low ancestry and protective effects in those with high ancestry. In the African American population, SNP rs760145 was protective against MS in individuals with lower African ancestry. Additionally, the HLA-A*02:01 allele showed a more pronounced protective effect when of European origin, whereas HLA-B*53:01 conferred protection when of African origin, especially among Hispanics [31].
Due to MS’s broad heterogeneity and complexity, understanding genetic polymorphisms across populations is crucial to clarifying how specific HLA variants influence susceptibility or protection.
Chi et al. [28], Beecham et al. [31], and Briggs and Sept [35] conducted studies involving European Caucasian individuals, all of which demonstrated that the presence of the HLA-DRB1*15:01 allele was significantly associated with MS susceptibility (Table 3). Additional risk alleles such as HLA-DQB1*06:02 and HLA-DPB1*03:01 were also identified (Table 3) [31].
Polymorphisms with high expression in MS reported in more than one study and with statistical significance.
| Polymorphism | Author/Year | Number of participants | Pathological stage | Region/Population | OR | p-value |
|---|---|---|---|---|---|---|
| DRB1*15:01 | Briggs and Sept/2021 | 386 | - | Europe | 1.89 | 3.8 × 10–3 |
| Chi et al./2019 | 3,777 | - | Europe | 2.45 | 2.45 × 10–11 | |
| Beecham et al./2022 | 2,652 | - | Europe | 1.83 | 9.63 × 10–4 | |
| Hedström et al./2021 | 13,950 | - | Sweden | - | - | |
| Gontika et al./2020 | 440 | RRMS-152, SPMS-130, PPMS-12 | Greece | POMS: 1.94 | 2.5 × 10–2 | |
| Fguirouche et al./2025 | 685 | - | Morocco | - | - | |
| Creary et al./2019 | 2,828 | - | Euro-Americans | 3.20 | < 2.2 × 10–16 | |
| Beecham et al./2022 | 2,435 | - | African Americans | 1.91 | 3.48 × 10–5 | |
| Chi et al./2019 | 3,692 | - | African Americans | 2.00 | 1.26 × 10–8 | |
| da Silva Bernardes et al./2019 | 153 | RRMS-24, SPMS-7, PPMS-6 | Brazil | - | - | |
| Chi et al./2019 | 4,915 | - | Asian Americans | 1.88 | 7.26 × 10–3 | |
| Ogawa et al./2019 | 474 | - | Japan | 3.44 | 2.1 × 10–5 | |
| Watanabe et al./2021 | 528 | RRMS-438, SPMS-70, PPMS-19 | Japan | 2.50 | < 1 × 10–3 | |
| DRB1*03:01 | Gontika et al./2020 | 440 | RRMS-152, SPMS-130, PPMS-12 | Greece | POMS: 2.44 | 4 × 10–3 |
| Chi et al./2019 | 3,692 | - | African Americans | 1.45 | 2.61 × 10–5 | |
| Fguirouche et al./2025 | 685 | - | Morocco | - | - | |
| Khdair et al./2025 | 140 | - | Jordan | 2.51 | 1.0 × 10–2 | |
| DRB1*04:01 | Fguirouche et al./2025 | 685 | - | Morocco | - | - |
| Khdair et al./2025 | 140 | - | Jordan | 18.28 | 4.0 × 10–3 | |
| DRB1*15:03 | Boullerne et al./2024 | 2,504 | - | Africa | - | - |
| Chi et al./2019 | 3,692 | - | African Americans | 1.37 | 8.11 × 10–5 | |
| DQB1*06:02 | Beecham et al./2022 | 2,652 | - | Europe | 2.70 | 2.66 × 10–24 |
| Akel et al./2022 | 100 | RRMS-92, SPMS-8 | Sweden | 1.64 | 2.52 × 10–5 | |
| da Silva Bernardes et al./2019 | 153 | RRMS-24, SPMS-7, PPMS-6 | Brazil | - | - | |
| Ogawa et al./2019 | 474 | - | Japan | 3.45 | 3.0 × 10–5 | |
| DPB1*03:01 | Beecham et al./2022 | 2,652 | - | Europe | 1.53 | 5.36 × 10–5 |
| Watanabe et al./2021 | 528 | RRMS-438, SPMS-70, and PPMS-19 | Japan | 2.33 | < 1 × 10–3 | |
| DRB1*15:01~DQA1*01:02~DQB1*06:02 | Derdelinckx et al./2020 | 30 | RRMS | Belgium | - | - |
| da Silva Bernardes et al./2019 | 153 | RRMS-24, SPMS-7, and PPMS-6 | Brazil | - | - | |
| DRB1*15:01~DQB1*06:02 | Mack et al./2019 | 831 | - | Euro-Americans | 3.98 | 2.22 × 10–16 |
| Goodin et al./2021 | 2,460 | - | African Americans | 3.30 | 1.0 × 10–4 | |
| Goodin et al./2021 | 29,636 | - | Europe | 3.25 | < 10–168 |
MS: multiple sclerosis; OR: odds ratio; RRMS: relapsing-remitting MS; SPMS: secondary progressive MS; PPMS: primary progressive MS; POMS: pediatric-onset MS.
Sweden, which has one of the world’s highest MS prevalence rates, showed enrichment of three genotypes containing the high-risk haplotype HLA-DRB5*01:01:01~DRB1*15:01:01~DQA1*01:02:01~DQB1*06:02:01; alleles such as DQA1*01:01:02 (OR = 9.21; p = 2.18 × 10–4), DQA1*01:02:01 (OR = 1.85; p = 8.15 × 10–6), and DQB1*05:02:01 (OR = 4.36; p = 1.71 × 10–5) were significantly overrepresented in patients [34]. In a case-control study by Hedström et al. [36], the DRB1*15:01 allele additively increased the risk of developing MS. Compared to individuals without this allele, the adjusted OR was 3.7 (95% CI: 3.3–4.1) for heterozygotes and 7.8 (95% CI: 6.4–9.5) for homozygotes [36].
The Orkney Islands in Northern Scotland have the highest reported MS prevalence worldwide (402/100,000). In this population, analysis of 127 SNPs identified rs9271069, a tag for HLA-DRB1*15:01, as the strongest risk variant (OR = 2.77), with higher frequency on the islands than on the mainland; however, given the small populations of Orkney (22,000) and Shetland (23,000), these allele-frequency differences explain only about two excess MS cases per archipelago [37].
Gontika et al. [38] examined HLA-DRB1 alleles in pediatric-onset MS (POMS) and adult-onset MS (AOMS), finding DRB1*15 and DRB1*03 (Table 3) as the most frequent genotypes in POMS and a significantly lower frequency of DRB1*11 in patients (OR = 0.09; p = 3.3 × 10–2), suggesting a protective effect; in AOMS, DRB1*15 was even more frequent (31.9%) but did not reach statistical significance.
In Belgium, Derdelinckx et al. [39] reported that 40% of patients, versus about 10% of controls, carried the HLA-DR15 haplotype (DRB1*15:01~DQA1*01:02~DQB1*06:02), but CD4+ T-cell responses to myelin peptides did not differ between carriers and non-carriers, suggesting that this haplotype contributes to susceptibility but does not directly shape peripheral myelin-specific reactivity.
The DRB1*15:01 allele, associated with an increased MS risk (OR up to 3) in Caucasian populations, is relatively rare in African populations, where the DRB1*15:03 allele predominates. This allele is also associated with MS in admixed populations in Latin America and the Middle East [40].
In a cohort of healthy individuals studied by Fguirouche et al. [29], no statistically significant differences were observed between class I HLA alleles and MS susceptibility. Regarding class II HLA, the alleles considered predisposing to MS had the following frequencies: DRB1*03 at 19.2%, DRB1*15 at 13.3%, DRB1*13 at 15.8%, and DRB1*04 at 12.7% [29].
Mack et al. [41] and Creary et al. [42] conducted studies in American cohorts of European ancestry, and both analyses showed that HLA-DRB1*15:01 conferred a very high disease risk (Table 3). Additionally, the haplotypes DRB1*15:01~DQB1*06:02 (OR = 3.98; p = 2.22 × 10–16), DRB1*03:01~DQB1*02:01 (OR = 1.63; p = 1.41 × 10–3), C*07:02~B*07:02 (OR = 1.99; p = 8.8 × 10–7), and the A*30:02 allele (OR = 3.32; p = 1.37 × 10–2) were also associated with disease susceptibility [41, 42].
In an African American cohort, Beecham et al. [31] identified the HLA-DRB1*15:01 allele (Table 3) and the HLA-A*02:01 allele (additive protection; OR = 0.68; p = 5.07 × 10–5). Conserved extended haplotype analyses showed a strong association for the European DRB1*15:01~DQB1*06:02 haplotype (Table 3) and variable effects for the African DRB1*15:03~DQB1*06:02 [32].
In Brazil, the extended DR15 haplotype was frequent and correlated with European ancestry; DQB1*06:02 and DRB1*15:01 were enriched among cases [43].
Iranian studies led to mixed SNP associations [27, 44]. In Jordan, DRB1*03:01 and DRB1*04:01 increased risk (Table 3), whereas DRB1*04:03 and DQB1*06:03 were protective [30].
In Japan, the most significant association was with the HLA-DRB1*15:01 allele (Table 3). In addition, the alleles HLA-DQB1*06:02 (Table 3), HLA-B*15:01 (OR = 2.95; p = 2.2 × 10–4), DRB1*04:05 (OR = 2.00; p < 1 × 10–3), and DPB1*03:01 (Table 3) were associated with disease susceptibility. On the other hand, the alleles DRB1*09:01 (OR = 0.47; p < 1 × 10–3), DRB1*01:01 (OR = 0.38; p < 1 × 10–3), DRB1*13:02 (OR = 0.22; p < 1 × 10–3), and DPB1*04:01 (OR = 0.35; p < 1 × 10–3) were identified as protective [45, 46].
In an Australian and New Zealand cohort of European ancestry, Burnard et al. [47] used elastic net penalized regression to identify MS-associated SNPs that standard GWAS might miss. The strongest signal was rs9271366, a tag for the HLA-DRB1*15 haplotype, with additional robust hits in tight linkage disequilibrium (e.g., rs2395182, rs3117098, rs3129941, rs6903608, rs9267992) and independent effects for rs2299851 and rs3819721 [47].
This systematic review integrated evidence on HLA polymorphisms associated with MS across diverse populations, allowing a broader interpretation of patterns that extend beyond isolated allele–disease associations. Although the included studies reported considerable heterogeneity in genetic signals, a consistent finding emerged: HLA-DRB1*15:01 remains the most reproducible and influential genetic risk factor for MS. It was the main risk allele in European Caucasians [28, 33, 35], African Americans [31, 32], Americans of European ancestry [41, 42] and in cohorts from Sweden [34, 36], Scotland [37], Greece [38], Belgium [39], Morocco [29], Brazil [43], Japan [45, 46], Australia and New Zealand [47]. Evidence in Colombian, Caribbean, Middle Eastern, and North African populations further supports the global consistency of this association [6]. Additionally, in individuals with MS, class II HLA genes may be abnormally expressed in non-immune cells such as oligodendrocyte lineage cells. Falcão et al. [48] demonstrated that subsets of oligodendrocyte precursor cells can express MHC class II-related molecules and acquire immune-like functions, including myelin uptake and the capacity to activate CD4+ T cells, both in experimental demyelination and in human MS tissue.
DQB1*06:02, frequently in strong linkage disequilibrium with DRB1*15:01, showed risk associations in several cohorts, including Europeans [31], Swedes [34], Brazilians [43], and Japanese [46]. The DRB1*15:01~DQB1*06:02 haplotype was associated with MS risk in Europeans [32], Americans of European ancestry [42], and African Americans [32]. Interestingly, individuals carrying DQB1*06:02 without DRB1*15:01 have been reported in Norway, suggesting population-specific haplotype structures that may uncouple these typically linked alleles [6].
Population-restricted alleles were also evident. DRB1*15:03 appeared predominantly in Africans [40] and African Americans [28], confirming earlier studies showing that this allele confers risk primarily in individuals of African ancestry [31, 40].
Other alleles, such as DRB1*03:01, showed context-dependent risk, appearing predisposing in Morocco [29], Jordan [30], in African Americans [28], and in the Greek POMS cohort [38], in line with previous findings in Sardinian, Australian, Afro-American, and Arab cohorts [29, 30]. Similarly, DPB1*03:01 showed risk associations in Europeans [31] and Japanese [45], with Ogawa et al. [46] reporting the same association in a Japanese cohort. The extended DR15 haplotype was also confirmed as predisposing in the Belgian [39] and Brazilian cohorts [43].
Protective alleles were less frequent but more consistent across populations. HLA-A*02:01, one of the most extensively validated protective markers, reduced MS risk in Europeans [31], Americans of European ancestry [41], African Americans [31] and was replicated in Italy and Norway [29], which significantly reduced risk even in the presence of HLA-DRB1*15:01 [41]. DRB1*01:01, a well-established protective allele, also demonstrated protection in Morocco [29], consistent with its reduced ability to effectively bind and present myelin epitopes [6].
Although some polymorphisms are common among individuals of European and American origin, the analysis of admixed populations, such as African American and Hispanic groups, also brought new insights by showing that ancestry influences the expression and effect of HLA alleles. DRB1*15:03~DQB1*06:02 showed a variable effect depending on the haplotype context, ranging from risk association to protective effect. Susceptibility variants to MS were also identified that show an effect modified by global ancestry, indicating ancestral differences that may be partly due to correlated socioeconomic or environmental factors, and others with an effect modified by local ancestry, indicating true genetic differences in the degree of risk/protection in ancestral contexts. The Hispanic population provided a remarkable example of genetic effect modification by ancestry. SNPs such as rs2844503 and rs3021302 demonstrated opposite effects depending on the proportion of Native American ancestry [31]. This risk modulation highlights the need to consider heterogeneous genetic contexts in population analysis and reinforces the value of stratified approaches.
Contrasting results were found for DRB1*04:01, which was predisposing in Moroccan [29] and Jordanian [30] cohorts, but protective in Americans of European ancestry [42]. These opposing effects were also reported by De Silvestri et al. [6], where DRB1*04 was protective in the group with a high percentage of Caucasians and predisposing in an ethnically mixed group, including Middle Eastern and North African individuals, although without statistical significance [6].
The interaction between protective and risk alleles illustrates the role of epistasis within the HLA region. In particular, the protective effect of HLA-A*02:01, which mitigates the risk conferred by HLA-DRB1*15:01, exemplifies how combined effects of class I and class II loci can modulate overall susceptibility. This gene-gene interaction highlights the complexity of HLA effects in MS and shows that individual alleles should not be interpreted in isolation. The SNP rs3135388 presented perfect linkage disequilibrium with both DRB1*15:01 and DQB1*06:02 [40], and SNP rs9271366 showed perfect linkage disequilibrium in Europeans [40] and in populations from Australia and New Zealand [47]. These data suggest that both SNPs can be used as reliable genetic markers for the risk alleles associated with MS in these populations, facilitating their detection in genetic studies.
Environmental interactions also emerged as relevant modulators. Hedström et al. [36] estimated that approximately 20% of all MS cases in Sweden are attributable to smoking exposure, while this number rises to over 40% in individuals positive for DRB1*15 and negative for A*02. These data reinforce that although genetic predisposition plays a central role, environmental factors also have a significant impact on MS risk, highlighting the multifactorial nature of the disease [36].
Clinical relevance also emerged in the context of treatment response. Class II alleles such as DRB1*04:01 and DRB1*04:08 consistently increased the risk of neutralizing antibodies (NAbs) against IFN-β, reducing its therapeutic efficacy [49, 50]. While DRB1*15:01 has been associated with NAbs in specific formulations [51], it does not exert a universal effect across studies, suggesting preparation-specific rather than allele-specific susceptibilities [49, 51].
Although Fguirouche et al. [29] did not identify statistically significant associations between class I HLA loci and MS susceptibility in the Moroccan cohort, previous serology-based studies have suggested a predisposing role for alleles such as HLA-A3, -B7, and -B27. This apparent discrepancy likely reflects differences in typing resolution between serological and molecular methods, limited sample size, and the influence of linkage disequilibrium with class II risk haplotypes, rather than excluding a contribution of class I alleles [29]. Similarly, Asouri et al. [27] found no significant HLA-DRA associations in Iran, which the authors attributed to limited sample size, population-specific factors, and the inherently multifactorial nature of MS.
Despite the consistent findings, this review presents some limitations that should be considered. First, methodological heterogeneity among the included studies such as: different diagnostic criteria for MS, variability in genotyping methods, and lack of uniformity in haplotype analysis, may limit direct comparability between results. Additionally, many studies focus on European or European-derived populations, while African, Asian, and Latin American populations remain underrepresented, which limits statistical power, ancestry-specific analyses, and the generalizability of findings. Another critical point is the limitation in the analysis of linkage disequilibrium and local ancestry in some works, which may obscure relevant genetic effects in admixed populations. The lack of longitudinal studies also restricts the understanding of the role of HLA not only in susceptibility but also in disease progression and treatment response. Nevertheless, the consistent detection of similar HLA-associated susceptibility signals across diverse populations supports the robustness of the overall conclusions, despite the observed heterogeneity.
Although these findings are not yet directly applicable in clinical practice, the reproducible association of specific HLA polymorphisms with MS susceptibility may contribute to refining genetic risk stratification and supporting more personalized monitoring and therapeutic strategies.
In overview, the available evidence confirms a central role of HLA class II alleles and conserved extended haplotypes, particularly those including HLA-DRB1*15, in MS susceptibility, while also supporting protective effects for specific alleles. These associations are shaped by ancestry and admixture, and are influenced by haplotypic structure, epistasis within the MHC, and interactions with environmental exposures such as smoking. Altogether, these findings highlight the need for high-resolution genotyping, standardized haplotype definitions, and inclusion of diverse populations to obtain more accurate and general estimates of HLA-related MS risk.
Conclusion: The HLA region, especially class II alleles, is relevant to the genetic susceptibility to MS. The alleles DRB1*15:01, DQB1*06:02, DRB1*15:03, DRB1*03:01, DRB1*04:01, DPB1*03:01, as well as the haplotypes DRB1*15:01~DQB1*06:02 and DRB1*15:01~DQA1*01:02~DQB1*06:02, stand out for their high expression and recurrence in the studied cohorts, highlighting a strong association with MS susceptibility. In contrast, the alleles A*02:01 and DRB1*01:01 emerge as protective markers, suggesting an immunomodulatory role that may attenuate the risk of developing MS.
The recognition of HLA variants associated with MS susceptibility has relevant implications for clinical practice, especially in the context of personalized medicine. Early identification of individuals carrying high-risk alleles may allow for closer monitoring of patients in risk groups or with initial clinical manifestations. Moreover, knowledge of the genetic profile may be integrated into polygenic risk prediction models, contributing to the development of prevention strategies, differential diagnosis, and therapeutic selection.
In conclusion, future research must prioritize the use of high-resolution genotyping methodologies [such as next-generation sequencing (NGS)], incorporate global and local ancestry analyses, and adopt more ethnically diverse sampling, especially in African and Latin American populations. Multicentric and collaborative studies will be essential to overcome sample size and methodological barriers, aiming at the construction of more robust genetic-clinical models applicable in different population contexts.
AOMS: adult-onset multiple sclerosis
APCs: antigen-presenting cells
CI: confidence interval
CNS: central nervous system
DMT: disease-modifying therapy
EDSS: Expanded Disability Status Scale
GWAS: genome wide association studies
HLA: human leukocyte antigen
IFN-γ: interferon-gamma
IL-2: interleukin-2
MBP: myelin basic protein
MHC: major histocompatibility complex
MS: multiple sclerosis
NAbs: neutralizing antibodies
OR: odds ratio
POMS: pediatric-onset multiple sclerosis
PPMS: primary progressive multiple sclerosis
PRISMA: Preferred Reporting Items for Systematic reviews and Meta-Analyses
RRMS: relapsing-remitting multiple sclerosis
SNPs: single nucleotide polymorphisms
SPMS: secondary progressive multiple sclerosis
TCRs: T cell receptors
Th: T helper
The supplementary material for this article is available at: https://www.explorationpub.com/uploads/Article/file/1004147_sup_1.pdf.
LP: Conceptualization, Investigation, Writing—original draft, Writing—review & editing. AV: Conceptualization, Investigation, Writing—review & editing, Validation, Supervision. Both authors read and approved the submitted version.
The authors declare that they have no conflicts of interest.
Not applicable.
Not applicable.
Not applicable.
The primary data for this systematic review were sourced online from databases listed in the methods. References articles are accessible on PubMed and Web of Science. The raw data supporting the conclusions of this manuscript will be made available by the authors, without undue reservation, to any qualified researcher.
The author AV thanks the Portuguese Foundation for Science and Technology (FCT) for the financial support to the Research Centre for Natural Resources, Environment and Society — CERNAS (UIDB/00681/2025). The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
View: 939
Download: 16
Times Cited: 0
