Affiliation:
1Department of Diagnostic Imaging and Radiotherapy, Kulliyyah of Allied Health Sciences, International Islamic University Malaysia, Kampus Kuantan, Kuantan 25200, Pahang Darul Makmur, Malaysia
ORCID: https://orcid.org/0000-0002-2384-8771
Affiliation:
1Department of Diagnostic Imaging and Radiotherapy, Kulliyyah of Allied Health Sciences, International Islamic University Malaysia, Kampus Kuantan, Kuantan 25200, Pahang Darul Makmur, Malaysia
ORCID: https://orcid.org/0009-0002-8600-1102
Affiliation:
1Department of Diagnostic Imaging and Radiotherapy, Kulliyyah of Allied Health Sciences, International Islamic University Malaysia, Kampus Kuantan, Kuantan 25200, Pahang Darul Makmur, Malaysia
ORCID: https://orcid.org/0009-0001-7344-6308
Affiliation:
2Department of Information Systems, Kulliyyah of Information and Communication Technology, International Islamic University Malaysia, Gombak 53100, Selangor Darul Ehsan, Malaysia
ORCID: https://orcid.org/0000-0003-1397-8174
Affiliation:
3Department of Optometry and Visual Sciences, Kulliyyah of Allied Health Sciences, International Islamic University Malaysia, Kampus Kuantan, Kuantan 25200, Pahang Darul Makmur, Malaysia
Email: zulfaezal@iium.edu.my
ORCID: https://orcid.org/0000-0001-5496-0822
Affiliation:
4Department of Computer Science and Software Engineering, College of Information Technology, United Arab Emirates University, P.O. Box 15551, Al Ain, Abu Dhabi, United Arab Emirates
ORCID: https://orcid.org/0000-0001-6661-8469
Explor Neuroprot Ther. 2026;6:1004141 DOI: https://doi.org/10.37349/ent.2026.1004141
Received: December 05, 2025 Accepted: March 05, 2026 Published: March 23, 2026
Academic Editor: P. David Mozley, Cornell University Weill School of Medicine, USA
Background: This systematic review critically evaluates whether machine learning (ML) identifies biologically meaningful sex-related brain architecture or merely exploits methodological artifacts and allometric scaling. While ML models achieve high classification accuracies, it remains unclear if these reflect stable, mechanistically informative dimorphism or are driven by confounds such as total intracranial volume (TIV) and site-specific noise. We examine how imaging modalities, algorithms, and population strata influence both classification outcomes and biological interpretability.
Methods: Following Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines, we searched Web of Science, PubMed, and Scopus through January 2024. Included studies [healthy humans, 3T magnetic resonance imaging (MRI), ML-based sex classification] were assessed for risk of bias, focusing on data leakage, validation strategies, and confound management.
Results: Thirty-five studies (n > 110,000) were included. While reported accuracies reached 98.06% for T1-weighted MRI, 96.0% for diffusion MRI (dMRI), and 94.72% for functional MRI (fMRI), performance was highly dependent on population characterization and age. Deep learning consistently outperformed traditional ML (TML) but showed high sensitivity to methodological artifacts. Notably, studies failing to correct for TIV reported potentially inflated accuracies, suggesting that many models identify physical scale rather than intrinsic neuroanatomical dimorphism.
Discussion: High classification accuracies are often bolstered by methodological confounds and a lack of cross-site validation. There is a significant discrepancy between ML-driven predictive power and biological inference validity. Current pipelines do not yet allow for robust, generalizable inference about brain sex. To move beyond statistical separation toward mechanistic understanding, the field must prioritize TIV-corrected benchmarks and diverse non-WEIRD (Western, Educated, Industrialized, Rich, Democratic) datasets. We conclude that while ML is a powerful pattern detector, its results must be interpreted with caution regarding biological dimorphism.
Magnetic resonance imaging (MRI) has transformed in vivo brain research by offering high-resolution insights into neuroanatomy and function. Its ability to delineate intricate tissue structures has made it indispensable in studying sex-based variations in brain morphology. Early reports highlighted anatomical differences such as greater brain weight and circumference in males [1]. Subsequent research using statistical morphometry identified sex differences in grey matter, although with varying magnitudes and conflicting biological interpretations [2–5]. For example, male brains are typically 8–12% larger than female brains—a difference that emerges early and persists across the lifespan [6, 7].
Recent advances in machine learning (ML) have enabled multivariate modeling of complex neuroimaging data, frequently demonstrating classification accuracies exceeding 90% [8, 9]. Compared to conventional statistics, ML models can identify subtle, distributed neuroanatomical patterns undetectable by univariate approaches. However, a critical gap remains between predictive performance and inferential validity. While ML is a powerful pattern detector, it is often unclear whether these high accuracies reflect stable, mechanistically informative dimorphism or are driven by proxies such as global size (allometric scaling) and demographic structure. If the predictive signal is largely a product of uncorrected total intracranial volume (TIV) or site-specific artifacts, then current models are constrained in what they can legitimately claim about sex-related differences in brain architecture.
In this review, “sex” refers strictly to biological attributes—such as chromosomal, hormonal, and anatomical characteristics assigned at birth—and does not encompass gender identity or sociocultural constructs. Our goal is to systematically evaluate whether ML-based neuroimaging pipelines currently allow for confound-controlled, cross-site generalizable inference about sex, or if they merely enable statistical separation at the population level. By interrogating what these high classification numbers truly mean, we aim to clarify the contribution of ML to our understanding of sex-related neuroanatomical organization, with broader implications for disorders exhibiting sex-specific prevalence or expression [10–15].
This systematic review was conducted in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement. The protocol for this review was not prospectively registered in a systematic review database (e.g., PROSPERO).
Studies were identified through an electronic search of the Web of Science, PubMed, and Scopus databases conducted on 19th January 2026. These databases were selected for their comprehensive coverage of biomedical and scientific literature, ensuring access to a wide array of relevant research at the intersection of neuroimaging, sex differences, and ML.
The search strategy employed keywords and indexed terms including “sex”, “gender”, “dimorphism”, “predict”, “classify”, “disparities”, “difference”, “discriminate”, “distinguish”, “brain”, “magnetic resonance imaging”, “MRI”, “machine learning”, “deep learning”, and “classifier”. These terms were combined using Boolean operators to refine the results. For example, the following string was adapted for each database: (sex OR gender) AND (dimorphism OR predict OR classify OR disparities OR difference OR discriminate OR distinguish) AND (brain) AND (magnetic resonance imaging OR MRI) AND (machine learning OR deep learning).
To ensure transparency and reproducibility, the exact Boolean search strings used for each database are provided below:
TS = ((“brain” OR “cerebral” OR “neuro” OR “cortex”) AND (“sex differences” OR “gender differences” OR “sexual dimorphism” OR “sex classification”) AND (“artificial intelligence” OR “ai” OR “machine learning” OR “deep learning”) AND (“magnetic resonance imaging” OR “MRI” OR “neuroimaging” OR “brain imaging”)).
(“brain” [tiab] OR “cerebral” [tiab] OR “neuro” [tiab] OR “cortex” [tiab]) AND (“sex differences” [tiab] OR “gender differences” [tiab] OR “sexual dimorphism” [tiab] OR “sex classification” [tiab]) AND (“artificial intelligence” [tiab] OR “ai” [tiab] OR “machine learning” [tiab] OR “deep learning” [tiab]) AND (“magnetic resonance imaging” [tiab] OR “MRI” [tiab] OR “neuroimaging” [tiab] OR “brain imaging” [tiab]).
(TITLE-ABS-KEY (brain OR cerebral OR neuro OR cortex)) AND (TITLE-ABS-KEY (“sex differences” OR “gender differences” OR “sexual dimorphism” OR “sex classification”)) AND (TITLE-ABS-KEY(“artificial intelligence” OR “ai” OR “machine learning” OR “deep learning”)) AND (TITLE-ABS-KEY(“magnetic resonance imaging” OR “MRI” OR “neuroimaging” OR “brain imaging”)).
The search encompassed all available literature up to 19 January 2026, with no date restrictions applied. This inclusive approach ensured that both foundational historical studies and recent developments were captured, enhancing the review’s robustness across a broad temporal range. Furthermore, the search date ensures the inclusion of the most recent advancements in the field as of early 2026.
To ensure a high level of technical consistency and diagnostic relevance, studies were selected based on the following criteria:
In vivo studies in healthy humans;
Use of 3T MRI;
ML-based sex classification;
Peer-reviewed, full-text English articles.
Post-mortem studies;
Clinical populations with neurological or psychiatric disorders;
Non-English articles;
Reviews, conference abstracts, and proceedings;
Studies using field strengths other than 3T (e.g., 1.5T or 7T) or those not utilizing ML for classification.
The emphasis on 3T MRI was chosen due to its prevalence in contemporary research, offering superior spatial resolution and signal-to-noise ratio (SNR) compared to lower field strengths. This standardization minimizes variance across scanners and enhances the reliability of the subtle brain measurements required for accurate sex classification.
The inclusion of both structural and functional imaging [T1-weighted, diffusion MRI (dMRI), and functional MRI (fMRI)] allows for a multidimensional assessment. While structural modalities provide insights into neuroanatomical morphology, fMRI captures dynamic activity, offering a holistic view of sex-based neurobiological variation [16–24].
The exclusion of non-English publications and secondary literature was a strategic decision to ensure uniformity in data interpretation and to maintain a focus on original, peer-reviewed findings. These constraints were designed to balance the breadth of the review with the need for high-quality, reproducible data.
Two reviewers independently screened all titles and abstracts identified through the search. The same reviewers independently assessed full-text articles for eligibility. Disagreements were resolved through consensus, or when necessary, by consultation with a senior author.
To evaluate the internal validity and methodological rigor of the included studies, a structured narrative quality assessment was performed. In the absence of a singular, universally accepted risk-of-bias tool specifically for ML-neuroimaging, we adapted criteria from the PROBAST (Prediction model Risk Of Bias Assessment Tool) and QUADAS-2 frameworks, tailored to address three critical domains:
A primary focus of our assessment was the identification of data leakage, a significant source of inflated accuracy in ML. We scrutinized studies for:
Feature selection leakage: Ensuring feature selection was performed strictly within cross-validation loops rather than on the entire dataset.
Subject overlap: Verifying that samples used for training were strictly isolated from testing, particularly in longitudinal or multi-modal studies where the same participant might appear across different data points.
Validation strategy: We prioritized studies using k-fold cross-validation or, ideally, hold-out independent test sets. Studies utilizing simple “train-test splits” on small samples were flagged for high risk of overfitting.
We assessed the “power” of the findings based on sample size and diversity:
Sample size: Small-scale studies (n < 100) were categorized as having a higher risk of capturing dataset-specific noise rather than generalizable sex differences.
Population bias: We evaluated the reliance on high-income, “WEIRD” datasets [e.g., Human Connectome Project (HCP), UK Biobank (UKB)]. While these datasets offer high technical quality, their demographic homogeneity limits the external validity of the resulting models.
The assessment monitored the handling of technical and biological confounds:
TIV and global scaling: Studies were evaluated for whether they corrected for TIV. As noted in our results, the absence of this correction in several high-accuracy studies represents a significant source of classification bias.
Scanner effects: We noted whether studies employed multi-site data or harmonization techniques (e.g., ComBat). Models trained and tested on a single scanner were considered to have a moderate risk of bias due to potential “site-signature” learning.
Across the 35 included studies, the risk of bias was found to be moderate to high. While the transition toward deep learning (DL) has improved the handling of complex data, consistent gaps remain—specifically regarding the reporting of TIV-normalization and the use of external, cross-site validation sets. These gaps suggest that the “ceiling” of sex-classification accuracy may be partially bolstered by methodological artifacts.
The study selection and screening process is illustrated in the PRISMA flow diagram (Figure 1). The initial search across three electronic databases yielded 112 records. Following the removal of 42 duplicate entries, 70 unique records remained for title and abstract screening. Of these, 14 irrelevant studies were excluded, and 56 articles underwent full-text assessment for eligibility. After a detailed review, 21 articles were excluded for not meeting the pre-defined inclusion criteria (e.g., use of 1.5T MRI, focus on clinical populations, or lack of ML-based classification).
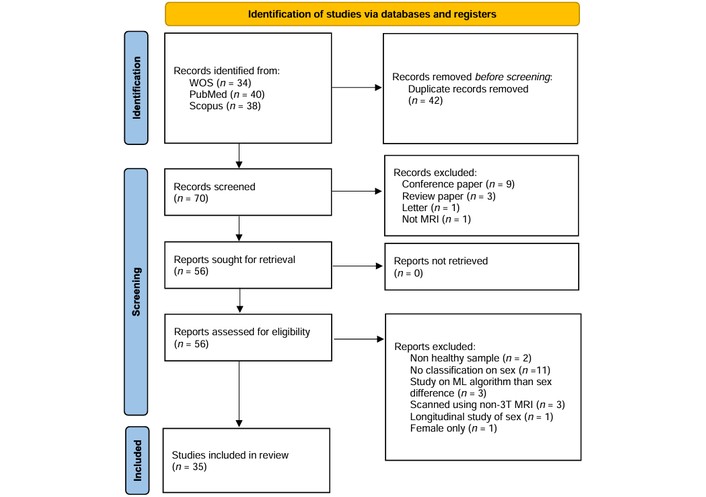
PRISMA 2020 flow diagram for new systematic reviews. Adapted from Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021;372:n71 (doi: 10.1136/bmj.n71). © Author(s) (or their employer(s)) 2021. CC BY. PRISMA: Preferred Reporting Items for Systematic Reviews and Meta-Analyses.
Ultimately, 35 articles [4, 9, 16–48] were deemed eligible and included in the final analysis. Data regarding sample population, age, morphometric features, ML algorithms, and sex-classification accuracy were extracted from these studies and are summarized in Table S1. Our quality assessment revealed a critical methodological landscape: while the field has transitioned toward powerful DL architectures, there remain significant gaps in inference validity. Notably, only a minority of high-accuracy studies explicitly corrected for TIV or employed external, cross-scanner validation sets. This suggests that the reported “performance ceiling” of sex classification may be partially bolstered by allometric scaling and site-specific artifacts.
This systematic review investigated the efficacy of various ML algorithms in classifying sex based on gray matter (GM) morphometric features. Overall, the classification models exhibited high accuracy in distinguishing between sexes, ranging from 56.0% to 98.06%. The highest performing method (98.06%) employed a specialized DL approach termed Multi-Layer 3D Convolution Extreme Learning Machine (MCN-ELM) [25], while the lowest accuracy (56.0%) was observed using a support vector machine (SVM) [31].
The selection of morphological features is a critical determinant of model performance. Particular emphasis was placed on GM segmentation data, assessed through various measurement approaches:
Grey matter volume: Five studies focused on GM volume [29, 34, 39, 42, 46]. Zhang et al. [42] (2020) achieved the highest accuracy at 94.3%, followed by Sanchis-Segura et al. [34] (2022), who reached 90.0% using SVM. Wang et al. [39] (2012) and Matte Bon et al. [29] (2024) reported accuracies of 86.1% and 82.3%, respectively. Conversely, Yang et al. [46] (2021) reported the lowest accuracy in this category at 66.0%. Notably, four of these studies utilized SVM; however, Matte Bon et al. [29] (2024) employed gradient-boosted trees, achieving 82.3% on limbic volume compared to 77.6% for non-limbic volume.
Global intensity and segmentation: Seven investigations incorporated global GM data or intensity features [17, 18, 20, 23, 25, 30, 40]. Following the 98.06% accuracy of MCN-ELM [25], Bi et al. [17] (2023) achieved 97.3% using a Convolutional Neural Network (CNN). Other notable results included 88.0% by Feis et al. [23], 84.0% using linear regression [40], and an area under the curve (AUC) of 0.85 using a CNN [30].
Cortical architecture: Six studies examined intricacies such as 3D morphology, thickness, surface area, and folding indices (gyrification, sulcal depth, and fractal dimension) [16, 18, 24, 28, 37, 48]. Luo et al. [28] (2019) reached 96.77% accuracy using hierarchical sparse representation-based classification (HSRC). Ge et al. [24] (2021) found that among five cortical features, cortical thickness yielded the highest accuracy (80.0%), followed by the gyrification index (72.0%) and sulcal depth (70.0%). Similarly, Metoki et al. [48] (2024) reported 76.0% accuracy for cortical thickness.
Modified indices: Studies employing modified indices (asymmetry, z-scores, or networks) generally reported lower performance [4, 9, 31]. For example, Dumitru [4] (2023) utilized brain asymmetry indices [subtraction index (SI), distance index (DI), and laterality index (LI)], yielding accuracies between 62.0% and 64.0%. However, Ebel et al. [9] (2023) achieved 95.78% using z-score normalization of GM voxels with logistic regression (LR).
The influence of global scaling: We observed a clear performance discrepancy linked to TIV management. Studies utilizing uncorrected GM volume, such as Sanchis-Segura et al. [34] (2022), reached 90.0% accuracy. However, the same study demonstrated that these differences attenuate significantly when TIV is controlled for, suggesting that uncorrected models may simply be identifying “head size”.
Feature sensitivity: Specialized DL approaches, such as the MCN-ELM [25], achieved the review’s highest accuracy (98.06%) by learning complex non-linear patterns. In contrast, models using brain asymmetry indices reported significantly lower accuracies (62.0–64.0%) [4], highlighting that predictive power is highly dependent on the “proxy potential” of the selected features.
Seven studies explored combining multiple morphological features, consistently revealing higher accuracies than individual features alone [18, 23, 24, 39, 40, 42, 47]. For instance, Brennan et al. [18] (2021) observed an increase in accuracy to 86.33% (from a baseline of 69.0–76.0%) when combining cortical area, thickness, volume, and intensity metrics.
Identifying discriminative brain regions further illuminates the landscape of sexual dimorphism. Matte Bon et al. [29] (2024) found that limbic volume (82.3%) was more predictive than whole-brain volume (79.7%). Conversely, Ebel et al. [9] (2023) reported that the whole brain exhibited the highest predictive power (AUC = 0.98), followed by the cerebellum (0.95) and thalamus (0.88). Luo et al. [28] (2019) noted that frontal regions were the most discriminative (87.87%), whereas the temporal gyrus was the least (62.89%).
Diffusion-weighted imaging (DWI) examines water diffusion patterns to assess white matter microstructural integrity and brain connectivity. Five of the included studies utilized diffusion-based features for sex classification [19, 23, 43, 46, 47].
Fractional anisotropy (FA) and DL: Xin et al. [43] (2019) achieved a peak accuracy of 93.3% using FA maps paired with a 3D CNN; notably, their comparative SVM model reached only 78.2% on the same data. Chen et al. [19] (2024) reported a similar accuracy of 91.0% using FA with a 2D CNN, which unexpectedly outperformed both 3D CNN and Vision Transformer (ViT) architectures. This indicates that DL excels at extracting subtle connectivity signatures that traditional ML (TML) misses.
Performance variance: Yeung et al. [47] (2023) obtained comparatively lower accuracies—79.74% for FA and 78.15% for mean diffusivity (MD)—using the BrainNet CNN framework. Similarly, Yang et al. [46] (2021) reported an accuracy of only 55.0% using an SVM on diffusion tensor data.
Multimodal enhancement: A consistent finding across diffusion studies was the benefit of multimodal integration. Feis et al. [23] (2013) achieved 83.0% accuracy using FA data alone, which increased significantly to 96.0% when combined with T1- and T2-weighted GM segmentation features. This trend was mirrored by Yang et al. [46] (2021), whose accuracy improved from 55.0% to 72.0% when diffusion features were fused with T1-weighted data.
fMRI captures dynamic brain activity by measuring blood-oxygen-level-dependent (BOLD) signals. This review evaluated both task-based fMRI, which maps activity during specific cognitive challenges, and resting-state fMRI (rs-fMRI), which assesses intrinsic functional connectivity.
Task-based fMRI: Two studies utilized task-based paradigms, reporting moderate classification performance. Leming and Suckling (2021) achieved an AUC of 0.783 using an emotion-processing task, while Xu et al. [44] (2020) reported an accuracy of 75.86% using a language-based paradigm.
rs-fMRI: Resting-state studies generally yielded higher accuracies, particularly when employing DL architectures. Ryali et al. [33] (2024) achieved a peak accuracy of 94.72% using a spatiotemporal deep neural network (stDNN). High performance was also reported by Sen and Parhi [36] (2021) at 94.0%, utilizing a random forest (RF) classifier on dynamic functional connectivity measures (tensor PARAFAC). Fan et al. [22] (2020) reported 93.05% accuracy by implementing a hybrid CNN-LSTM (long short-term memory) architecture to capture temporal dependencies.
Standard performance and baselines: Most rs-fMRI studies reported accuracies between 80.0% and 90.0% [21, 32, 40, 48]. Conversely, the lowest performances were observed in studies utilizing SVM: Patel et al. [32] (2024) reported 67.53%, and Satterthwaite et al. [35] (2015) reported 71.0%.
A key finding across the literature is the significant improvement in classification accuracy achieved through the integration of multiple MRI sequences. Five studies demonstrated that fusing morphological and functional data—including T1-weighted, T2-weighted, diffusion-weighted, and functional (resting-state and task-evoked) MRI—consistently outperformed single-modality models.
Incremental gains: The earliest evidence for this trend came from Wang et al. [39] (2012), who observed that combining GM volume with rs-fMRI increased accuracy to 89.1%, surpassing the 86.1% achieved using GM volume alone.
Structural synergy: Feis et al. [23] (2013) achieved a near-perfect accuracy of 96.0% by integrating features from three structural sequences: T1-weighted (88.0%), T2-weighted (85.0%), and diffusion-weighted MRI (83.0%).
Holistic multimodal integration: More recent investigations utilizing both structural and functional domains have reported the highest accuracies in the review. Weber et al. [40] (2022) achieved 93.3%, while Zhang et al. [42] (2020) reached a peak of 96.6% by synthesizing features from GM, white matter, rs-fMRI, and task-evoked fMRI.
A comparative analysis of the included studies reveals distinct patterns between TML and DL architectures. While TML models—including SVM, RFs, and gradient boosted trees—demonstrated high performance, DL models consistently established the upper bounds of classification accuracy.
TML: Among studies utilizing TML, the peak accuracy reached was 96.77%, achieved by Luo et al. [28] (2019) through their HSRC approach. SVM was the most prevalent algorithm, utilized in 17 studies. Zhang et al. [42] (2020) reported the highest SVM-based accuracy (96.6%) by integrating structural and functional features. However, TML performance varied significantly, with some SVM models reporting accuracies as low as 55.0% [31, 46], highlighting a high sensitivity to feature selection and preprocessing.
DL: DL techniques, particularly CNNs, pushed accuracy thresholds further. The highest overall accuracy in this review (98.06%) was achieved using a specialized MCN-ELM [25].
In head-to-head evaluations, the choice of algorithm often dictated performance. Sanchis-Segura et al. [34] (2022) compared five algorithms [linear discriminant analysis (LDA), LR, MARS, RF, and SVM] in a sex- and age-matched cohort (n = 876), finding that SVM outperformed the others with an accuracy of 90.0% when using uncorrected GM volume.
However, the assumption that DL always outperforms TML was challenged by Ebel et al. [9] (2023). In their study using the Study of Health in Pomerania (SHIP) dataset, LR (95.78%) outperformed a CNN [9]. This was attributed to the use of z-score normalization on GM voxels, suggesting that rigorous feature engineering in TML can occasionally match or exceed the automated feature extraction of DL models. This dichotomy underscores that while DL generally offers higher ceilings, TML remains highly effective when paired with optimized preprocessing.
Given that brain maturation and aging involve distinct neurobiological processes, this review assessed how ML accuracy varies across the lifespan.
Children and adolescents (under 23 years): Eight studies focused on younger populations [17, 18, 30, 32, 35, 37, 38, 46]. The highest accuracy reported was 97.3% in a large-scale study of approximately 11,000 children (aged 9–12 years) from the Adolescent Brain Cognitive Development (ABCD) study [17]. This suggests that sex-specific neuroanatomical patterns are already highly distinguishable by late childhood.
Young adults (20–37 years): This demographic was the most frequently studied, with 17 investigations primarily utilizing the HCP dataset [9, 16, 19, 21, 22, 24, 25, 28, 29, 33, 34, 36, 40, 41, 43, 45, 46]. The review’s peak accuracy of 98.06% was achieved within this group using a DL architecture, suggesting that the early adult brain may represent the “pinnacle” of sexual dimorphism in neuroimaging features [25].
Lifespan and older adults (14–90 years): Six studies examined broader age ranges to capture aging effects [9, 21, 26, 41, 42, 47]. The highest accuracy in these heterogeneous samples was 95.78%, achieved through the SHIP cohort [9]. The slightly lower ceiling in lifespan studies compared to young adult cohorts may reflect the increased inter-individual variability and cortical atrophy associated with aging, which can obscure sex-specific features.
To reach further stratification power, the included studies were characterized by developmental stage and morphometric constraints. A clear performance gradient emerged when stratifying by age: studies focusing on the adolescent stratum (9–12 years) and young adult stratum (20–35 years) reported the highest mean accuracies (97.3% and 98.06%, respectively). In contrast, studies spanning the full adult lifespan (14–90 years) showed increased variance and a slightly lower performance ceiling (95.78%), likely due to age-related cortical atrophy. Furthermore, when characterizing groups by TIV, studies that did not stratify for head size reported consistently higher accuracies. This suggests that the “power” of sex classification is significantly mediated by the homogeneity of the age and size strata being analyzed.
This systematic review demonstrates that while ML can classify biological sex from neuroimaging data with high precision, these results require a fundamental shift in interpretation. By synthesizing evidence across diverse age groups and imaging modalities, we find that while human brains can be categorized with up to 98.06% accuracy, these results are heavily influenced by model architecture, feature selection, and, crucially, methodological rigor. While high accuracies are often used to reinforce the existence of sex-related neuroanatomical patterns [28, 29], they must be interrogated to determine if they represent biologically meaningful dimorphism or merely enable statistical separation via proxies.
The most consequential finding regarding the validity of these models is the role of confounding factors, primarily TIV. Because male brains are, on average, 8–12% larger than female brains [10], models trained on uncorrected volumetric data may achieve “near-perfect” accuracies by exploiting global scaling cues rather than intrinsic multivariate morphology. Our review found that reporting on TIV correction was alarmingly limited [9, 34]. As Sanchis-Segura et al. [34] demonstrated, sex differences appear “large” when uncorrected but become significantly attenuated once TIV is controlled. If a model “cheats” by using head size as a proxy for sex, it is not identifying neurobiological sex differences, but rather allometric scaling. This distinction is vital: if the predictive signal is size-driven, the field is not yet in a position to use these models as evidence for stable, mechanistically informative dimorphism.
The transition toward DL has pushed classification boundaries toward a 98% ceiling [25]. Architectures such as 3D CNNs excel by automatically learning complex, non-linear spatiotemporal patterns. However, DL models showed higher sensitivity to methodological artifacts. While TML, specifically SVMs, remains the most utilized due to transparency [18, 23], DL is the primary driver of state-of-the-art precision. Yet, without robust explainability, it remains unclear if these “black box” models are identifying biological truth or “predictive noise”.
Integrating multiple MRI sequences consistently outperformed single-modality models [40]. Fusing structural (T1w, T1-weighted image), diffusion (dMRI), and functional (fMRI) data allows for a holistic view of sex-based variation. However, the increased accuracy of multimodal models (up to 96.6%) may also amplify the aggregation of confounding signals—such as motion artifacts in fMRI paired with size-bias in T1w data—unless rigorous cross-modal confound control is applied.
A critical methodological tension exists between traditional statistical-driven and ML-driven applications. While statistical approaches prioritize the identification of significant regional differences (inference), ML prioritizes the maximization of classification accuracy (prediction). Our analysis highlights that these two approaches do not always yield the same prioritization of brain features [9, 34]. For instance, an ML model might prioritize a combination of subtle, non-significant features—such as cortical folding patterns or functional connectivity weights—to reach high accuracy. This discrepancy suggests that ML may “prioritize” features that are statistically weak on their own but computationally powerful in aggregate, potentially exploiting “predictive noise” that lacks true inferential validity.
High accuracy does not equate to biological generalizability. A recurring limitation is the absence of cross-site or cross-scanner validation. Models trained on single-site datasets (e.g., HCP, UKB) may inadvertently learn “site-specific signatures”—such as scanner gradients or head-coil configurations—rather than universal biological signals. Furthermore, the reliance on WEIRD (Western, Educated, Industrialized, Rich, Democratic) populations introduces demographic biases. An accuracy of 98% may reflect overfitting to site-specific noise and demographic homogeneity, failing when applied to more diverse groups.
Crucially, these findings do not imply deterministic brain dimorphism. High classification accuracy is a testament to the power of multivariate statistics to find patterns in overlapping data, not an endorsement of binary biological essentialism. Individual brains are more accurately described as a “mosaic” [6] of features. Transparent reporting and cautious interpretation are essential to prevent the misuse of high-accuracy ML results in contexts that demand binary certainty, where nature provides a spectrum of variability.
The high classification accuracies identified in this review—reaching up to 98.06%—necessitate a move beyond viewing sex-classification as a purely technical challenge. Our findings reveal specific methodological trends that carry significant ethical weight.
Our review found that T1-weighted imaging and DL architectures (e.g., MCN-ELM) produce the highest accuracies. However, these results must be interpreted with caution. There is a profound ethical risk that these “near-perfect” statistical separations are used to support essentialist claims—the idea that male and female brains are “opposite” or “dimorphic”. As noted in our discussion of the “brain mosaic”, high accuracy in a multivariate model does not negate the extensive overlap in individual brain features. Misrepresenting these statistical patterns as deterministic labels could be misused to justify gender-based discrimination in education, employment, or forensic contexts.
Our findings show a heavy reliance on datasets like the HCP, UKB, and ABCD. Because these cohorts predominantly sample WEIRD populations, there is a direct ethical risk of creating inequitable algorithms. If a model achieves 98% accuracy on a high-income, Western cohort but is then deployed in a global health context without cross-cultural validation, it may produce biased or invalid results for underrepresented groups. The “robustness” reported in current literature is, therefore, demographically “fragile”, potentially reinforcing systemic healthcare disparities.
A critical ethical concern arising from our analysis is the lack of TIV correction in several high-accuracy studies. When models “cheat” by using global head size as a proxy for sex, they are not identifying neurobiological sex differences; they are identifying physical scale. Attributing this to “brain sex” is ethically problematic, as it presents a technical confound as an innate biological truth, potentially leading to flawed theories about cognitive or behavioral differences.
To mitigate these risks, we recommend that future neuroimaging-ML research adopt the following mandatory reporting standards:
Mandatory TIV disclosure: Researchers must explicitly report whether and how they corrected for TIV to ensure accuracy reflects morphology, not just size.
Granular demographic reporting: Manuscripts should include a detailed “diversity statement” regarding the training data, specifying the ethnicity, socioeconomic status, and geographic origin of participants.
Cross-dataset validation: High accuracy claims should be treated as preliminary until validated on an external, demographically distinct dataset to test for algorithmic fairness.
Interpretive caution: Results should be framed in terms of “statistical classification” rather than “biological essence”, explicitly acknowledging the role of neuroplasticity and environment in shaping the observed features.
A primary limitation of this systematic review is the methodological heterogeneity regarding confound management, particularly TIV. Since male brains are, on average, 8–12% larger than female brains, models that do not rigorously normalize for TIV—a gap identified in several high-accuracy studies—may be capturing allometric scaling rather than intrinsic neuroanatomical dimorphism. This “size-confounding” significantly inflates accuracy and obscures true biological signals.
Furthermore, the field’s heavy reliance on a few large, shared public datasets (e.g., HCP, UKB) introduces a profound population bias. These datasets represent WEIRD cohorts, which lack the diversity necessary to claim universal biological validity. The absence of cross-site validation in many studies suggests that models may be overfitting to site-specific noise—such as scanner manufacturer gradients or specific preprocessing idiosyncrasies—rather than generalizable neurobiology. This “site-signature” effect means that a 98% accuracy on one dataset might drop to chance levels when applied to another [44].
The absence of prospective protocol registration in a repository such as PROSPERO is a notable limitation. Although the review followed a rigorous, predefined internal protocol verified by multiple independent authors to minimize selective reporting bias, the lack of public registration may impact the transparency of the study selection process. Additionally, while our search was expanded to include Web of Science, PubMed, and Scopus to capture the interdisciplinary intersection of neuroscience and engineering, the exclusion of other technical databases (e.g., IEEE Xplore) remains a potential constraint on the absolute comprehensiveness of the literature surveyed. Finally, the restriction to English-language publications may exclude culturally diverse findings that could further challenge the “binary” narrative of brain sex. These combined artifacts suggest that current literature may provide an inflated sense of certainty regarding the categorical distinction of the human brain.
This systematic review reveals that while ML can achieve peak accuracies of 98.06% in sex classification, these results must be viewed with significant skepticism. The high performance is likely bolstered by methodological artifacts, including inadequate TIV correction and the learning of dataset-specific scanner signatures. Rather than confirming a deterministic binary, these models identify population-level statistical trends that are heavily influenced by global head size and demographic homogeneity.
We conclude that the human brain is better described as a mosaic of overlapping features rather than two distinct categories. For ML to move from a “pattern detector” to a valid neuroscientific tool, future research must prioritize mandatory TIV disclosure, cross-site validation, and the use of demographically diverse datasets. Without these rigorous standards, high classification accuracy remains a statistical achievement of limited biological and clinical utility.
ABCD: Adolescent Brain Cognitive Development
AUC: area under the curve
CNN: Convolutional Neural Network
DI: distance index
DL: deep learning
dMRI: diffusion magnetic resonance imaging
DWI: diffusion-weighted imaging
FA: fractional anisotropy
fMRI: functional magnetic resonance imaging
GM: gray matter
HCP: Human Connectome Project
HSRC: hierarchical sparse representation-based classification
LDA: linear discriminant analysis
LI: laterality index
LR: logistic regression
LSTM: long short-term memory
MCN-ELM: Multi-Layer 3D Convolution Extreme Learning Machine
MD: mean diffusivity
ML: machine learning
MRI: magnetic resonance imaging
PRISMA: Preferred Reporting Items for Systematic Reviews and Meta-Analyses
RF: random forest
rs-fMRI: resting-state functional magnetic resonance imaging
SHIP: Study of Health in Pomerania
SI: subtraction index
SNR: signal-to-noise ratio
stDNN: spatiotemporal deep neural network
SVM: support vector machine
T1w: T1-weighted image
TIV: total intracranial volume
TML: traditional machine learning
UKB: UK Biobank
WEIRD: Western, Educated, Industrialized, Rich, Democratic
The supplementary table for this article is available at: https://www.explorationpub.com/uploads/Article/file/1004141_sup_1.pdf.
During the preparation of this work, the authors used Gemini (Google) to improve the manuscript's linguistic clarity, readability, and grammatical accuracy. This service was utilized solely as a language-polishing tool following the completion of the initial draft. The authors formally confirm that all substantive intellectual content, data analysis, interpretation of results, and final conclusions are entirely their own. The AI tool did not generate new ideas, perform statistical evaluations, or influence the scientific direction of the study. After using this service, the authors reviewed and edited the content as needed and take full responsibility for the entire content of the publication.
AHS: Investigation, Formal analysis, Data curation, Methodology, Writing—original draft. IJ: Supervision, Conceptualization, Methodology, Funding acquisition, Writing—review & editing. ZAAM: Supervision, Conceptualization, Methodology, Funding acquisition, Writing—review & editing. MIMT: Supervision, Conceptualization, Methodology, Funding acquisition, Writing—review & editing. MZCA: Supervision, Conceptualization, Methodology, Funding acquisition, Writing—review & editing. ST: Visualization, Validation. All authors read and approved the submitted version.
The authors declare that they have no conflicts of interest.
Not applicable.
Not applicable.
Not applicable.
The datasets supporting the findings of this study are available from the corresponding author upon reasonable request.
This research is part of the Fundamental Research Grant Scheme (FRGS/1/2024/WAS12/UIAM/01/3) under the Ministry of Higher Education Malaysia. The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
© The Author(s) 2026.
Open Exploration maintains a neutral stance on jurisdictional claims in published institutional affiliations and maps. All opinions expressed in this article are the personal views of the author(s) and do not represent the stance of the editorial team or the publisher.
Copyright: © The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
View: 1524
Download: 25
Times Cited: 0
